Massively Parallel Molecular Dynamics Using Adaptive Weighted Ensemble

Massively Parallel Molecular Dynamics Using Adaptive Weighted Ensemble Badi’ Abdul-Wahid PI: Jesús A. Izaguirre CCL Workshop 2013
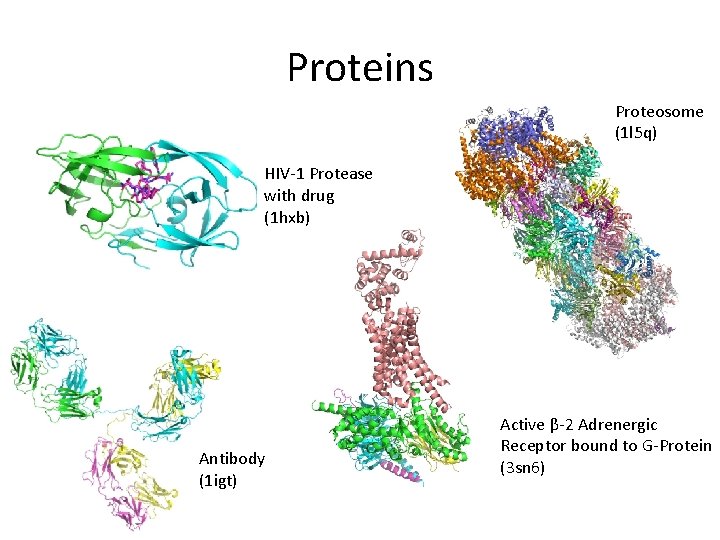
Proteins Proteosome (1 l 5 q) HIV-1 Protease with drug (1 hxb) Antibody (1 igt) Active β-2 Adrenergic Receptor bound to G-Protein (3 sn 6)
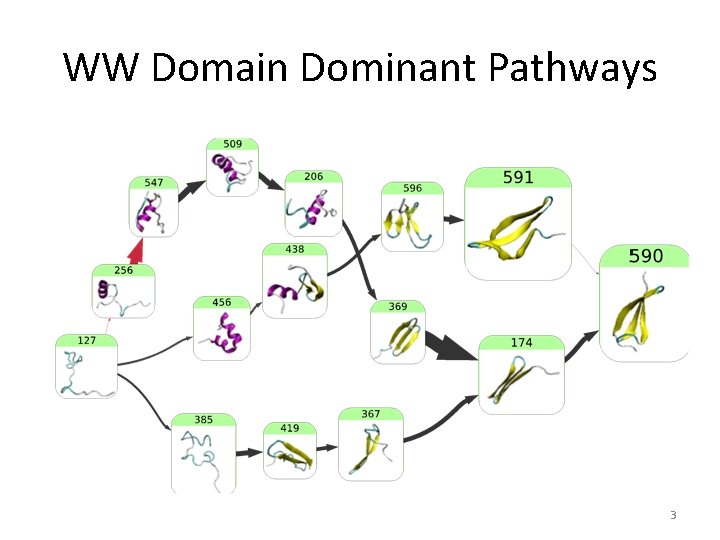
WW Domain Dominant Pathways 3
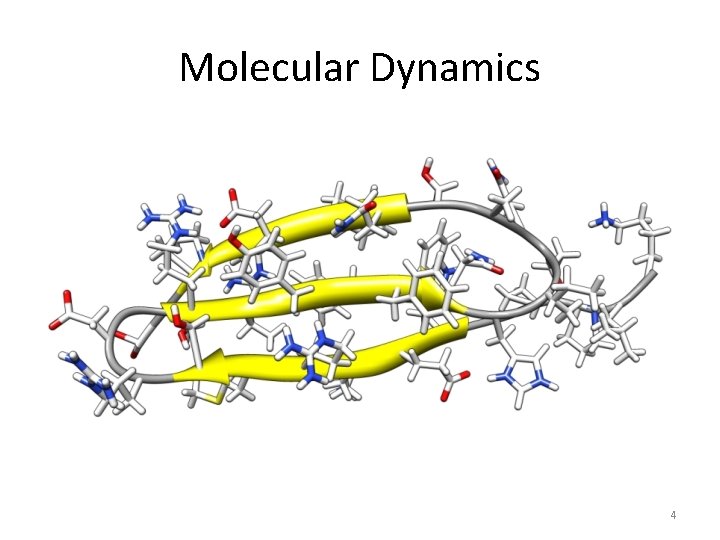
Molecular Dynamics 4

Many Complex Pathways Lane et al. 2013 Curr. Opin. Struct. Biol. 23 5
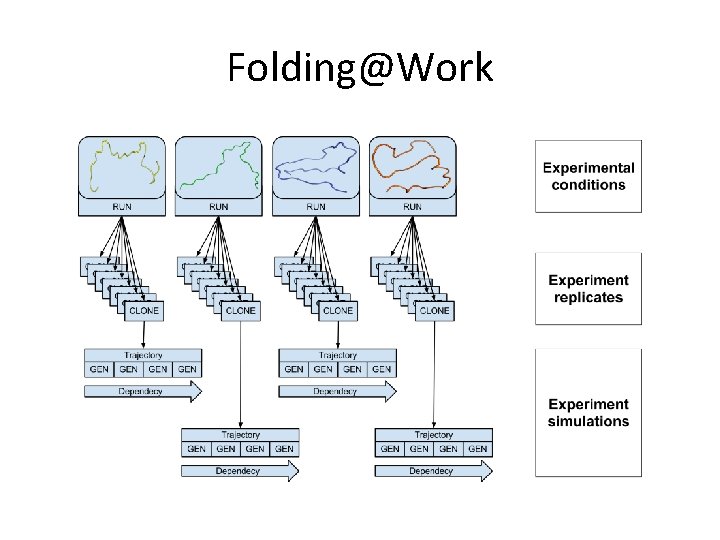
Folding@Work

Folding@Work Scaling
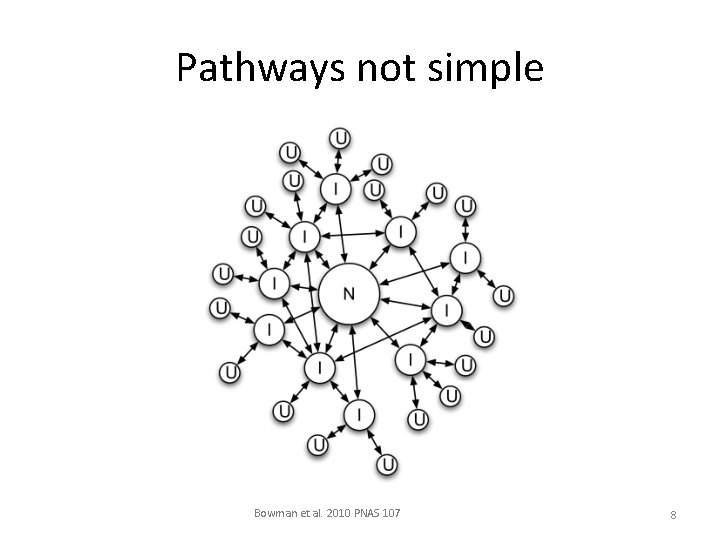
Pathways not simple Bowman et al. 2010 PNAS 107 8
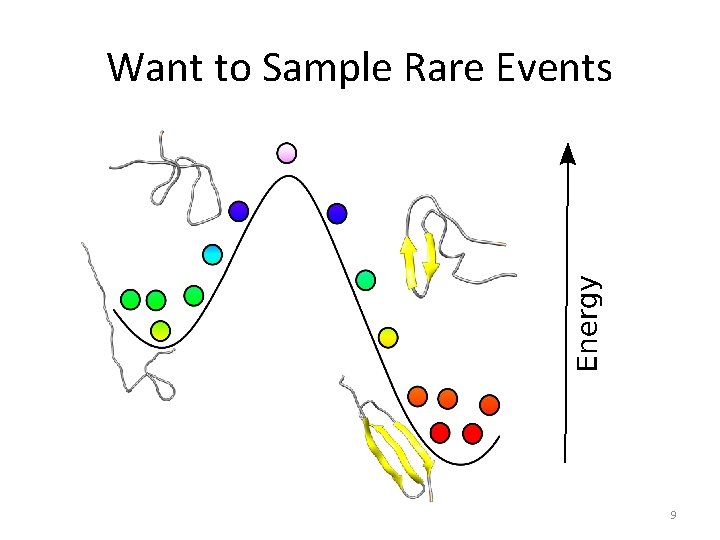
Want to Sample Rare Events 9
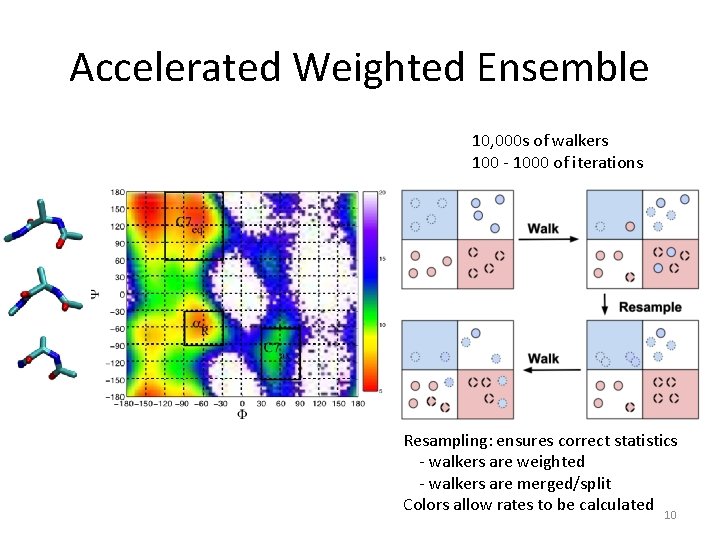
Accelerated Weighted Ensemble 10, 000 s of walkers 100 - 1000 of iterations Resampling: ensures correct statistics - walkers are weighted - walkers are merged/split Colors allow rates to be calculated 10
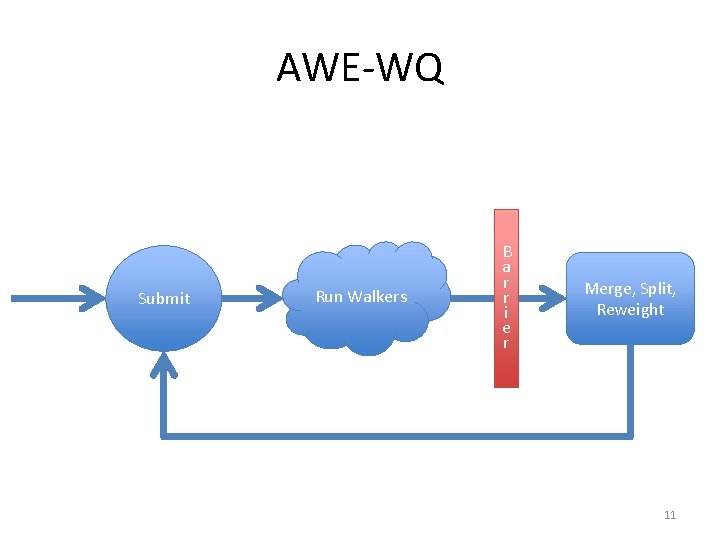
AWE-WQ Submit Run Walkers B a r r i e r Merge, Split, Reweight 11

How does WQ enable AWE?
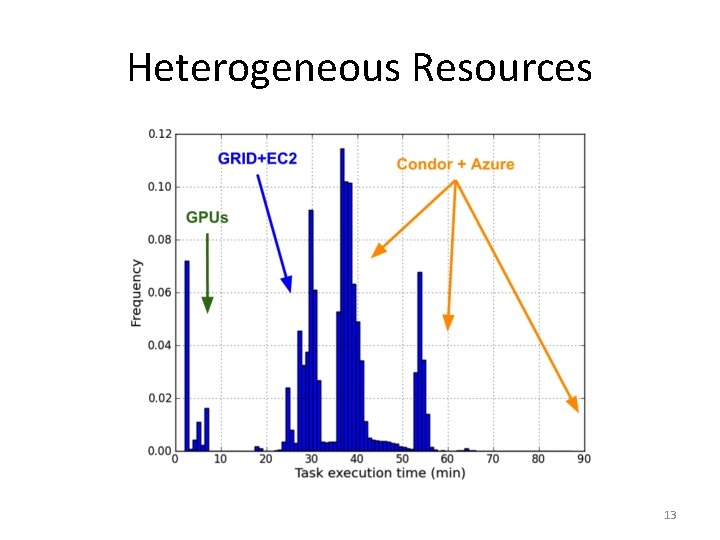
Heterogeneous Resources 13
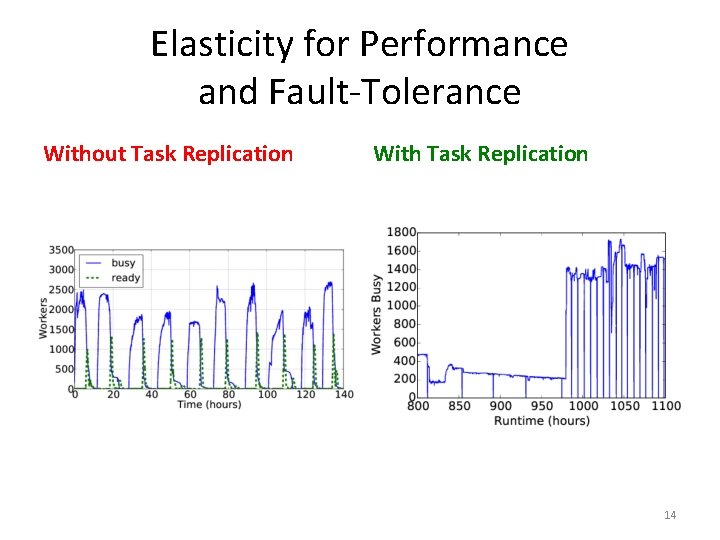
Elasticity for Performance and Fault-Tolerance Without Task Replication With Task Replication 14

Some Numbers • • • 3 million+ tasks executed 600+ years of CPU time 8 months wall time Aggregate 1 μs/day achievable 1. 5+ ms simulation time 2500+ sustained workers
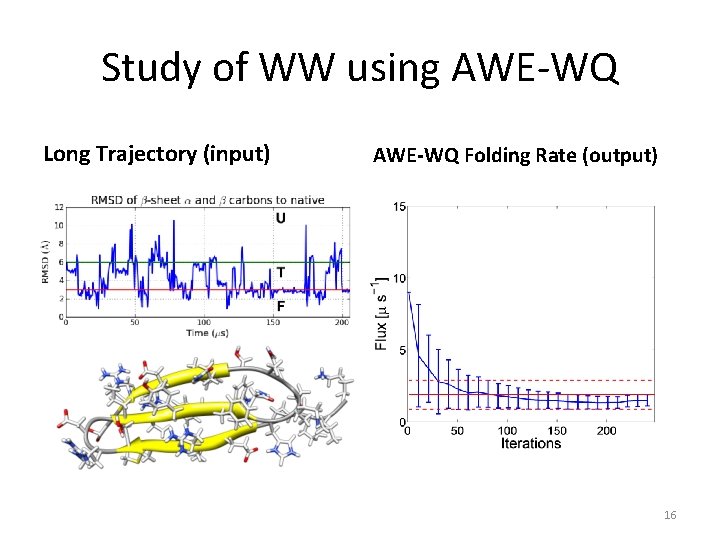
Study of WW using AWE-WQ Long Trajectory (input) AWE-WQ Folding Rate (output) 16

Future Work • Support for explicit solvent simulations • Improved cell discovery and partitioning • Incorporate improvements to Work Queue – GPU scheduling – very tricky! – Scheduling of multicore programs – Hierarchical Work Queue

Acknowledgements • Lab – – – – Prof. Jesús A. Izaguirre Dr. Chris Sweet Haoyun Feng Kevin Kastner Yong Hwan Kim Ronald Nowling James Sweet • Collaborators: – – – Prof. Douglas Thain Prof. Eric Darve (Stanford) Dr. Ronan Costaouec (Stanford) Dinesh Rajan Li Yu • Funding: – NSF CCF-1018570, NIH 1 R 01 GM 101935 -01, NIH 7 R 01 AI 039071. • Resources: – Notre Dame Center for Research Computing – Stanford Institute for Computation and Mathematical Engineering 18

Feature Requests
- Slides: 19