Massively Distributed Computing and An NRPGM Project on

Massively Distributed Computing and An NRPGM Project on Protein Structure and Function Computation Biology Lab Physics Dept & Life Science Dept National Central University
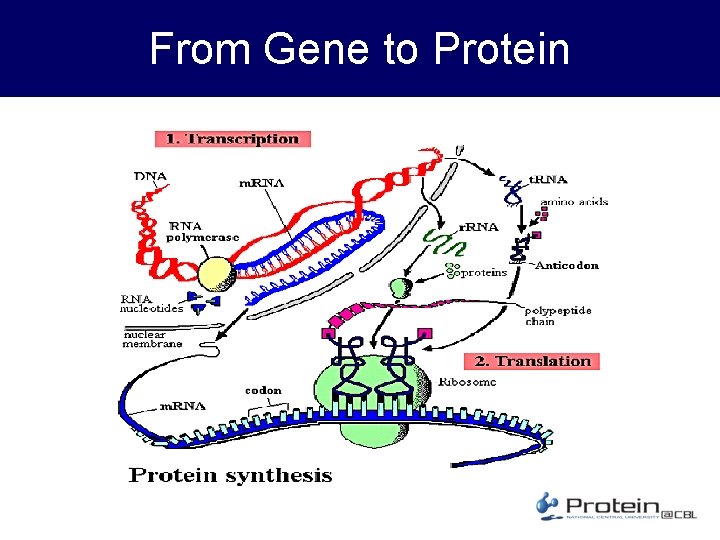
From Gene to Protein

About Protein • Function – Storage, Transport, Messengers, Regulation… Everything that sustains life – Structure: shell, silk, spider-silk, etc. • Structure – String of amino acid with 3 D structure – Homology and Topology • Importance – Science, Health & Medicine – Industry – enzyme, detergent, etc. • An example – 3 hvt. pdb

Problem Structure & Function • Primary sequence Native state with 3 D structure – Structure function – Expensive and time consuming • Misfolding means malfunction – Mad cow disease (“prion” misfolds)
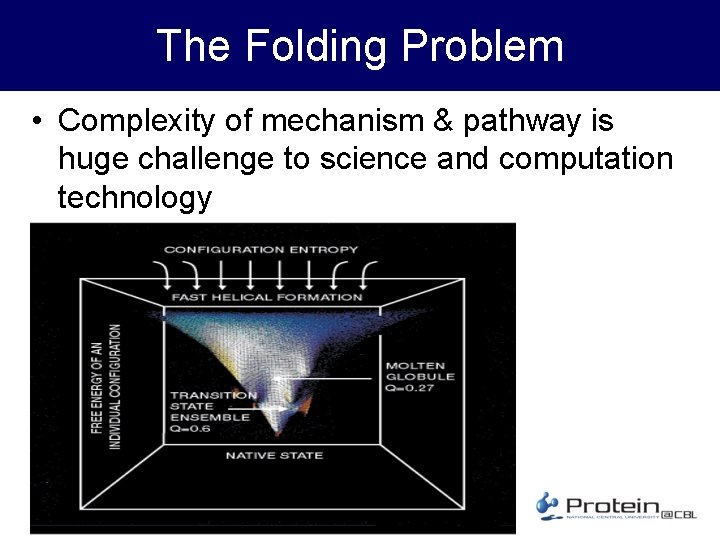
The Folding Problem • Complexity of mechanism & pathway is huge challenge to science and computation technology

Molecular Dynamics (MD) • Molecular’s behavior determined by – Ensemble statistics – Newtonian mechanics • Experiment in silico • All-atom w. water – Huge number of particles • Super-heavyduty computation • Software for macromolecular MD available – CHARMm, AMBER, GROMACS

Basic Statistics for Protein MD Simulation • Atoms in a small protein plus surrounding water (N) 32000 • Approximate number of interactions in force calculation (N 2/2 ) 0. 5 x 109 • Machine instructions per force calculation 1000 • Machine time per calculation (CPU: 3 G) 160 sec • Typical time-step size 0. 5 x 10– 15 sec • Total number of steps for 1 ms folding 0. 5 x 109 steps • Total machine time (160 sec x 0. 5 x 109) 106 days

How to overcome the factor of 1 million • A two-pronged approach – Faster or more CPUs • • – Nature of bottle-neck in protein folding dictated by Boltzmann distribution, can be overcome by large statistics (parallel computing NOT needed) Our solution: Massively distributed computing We seek factor of ~ 10, 000 Note. IBM’s solution: Blue Machine w/ 106 CPUs Shorten computation time • • • Many simulation steps needed b/c short time-scale of fast (vibrational) mode of ~ 10 fs But time-scale of folding motion slow, ~ 1 ns Ideal solution: by-pass or smooth out fast modes

Protein Studies by Massively Distributed Computing A Project in National Research Program on Genomic Medicine • Scientific – Protein folding, structure, function, proteinmolecule interaction – Algorithm, force-field • Computing – Massive distributive computing • Education – Everyone and Anyone with a personal PC can take part • Industry – collaborative development

Distributive Computing • Concept – Computation through internet – Utilize idle PC power (through screen-saver) • Advantage – Cheap way to acquire huge computation power – Perfectly suited to task • Huge number of runs needed to beat statistics • Parallel computation NOT needed • Massive data - good management necessary • Public education – anyone w/ PC can take part

Hardware Strategies • Parallel computation (we are not this) – PC cluster – IBM (The blue gene), 106 CPU • Massive distributive computing – Grid computing (formal and in the future) – Server to individual client (now in inexpensive) • Examples: SETI, folding@home, genome@home • Our project: protein@CBL

Software Components • Dynamics of macromolecules – Molecular dynamics, all atomistic or mean-field solvent – Computer codes • GROMACS (for distributive comp; freeware) • AMBER and others (for in-house comp; licensed) • Distributed Computing – COSM - a stable, reliable, and secure system for large scale distributed processing (freeware)
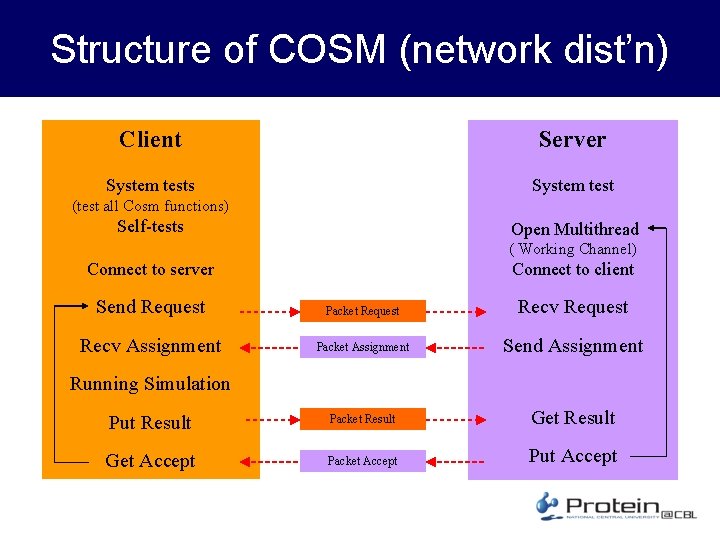
Structure of COSM (network dist’n) Client Server System tests System test (test all Cosm functions) Self-tests Open Multithread ( Working Channel) Connect to client Connect to server Send Request Packet Request Recv Assignment Packet Assignment Send Assignment Put Result Packet Result Get Accept Packet Accept Put Accept Running Simulation
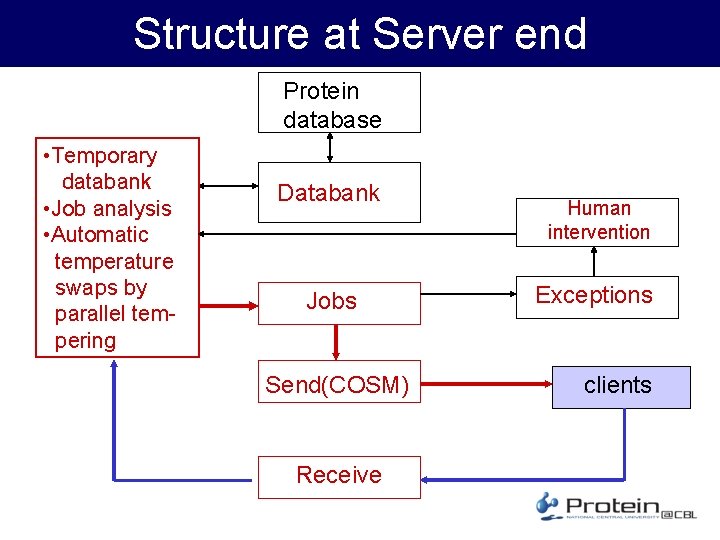
Structure at Server end Protein database • Temporary databank • Job analysis • Automatic temperature swaps by parallel tempering Databank Jobs Send(COSM) Receive Human intervention Exceptions clients
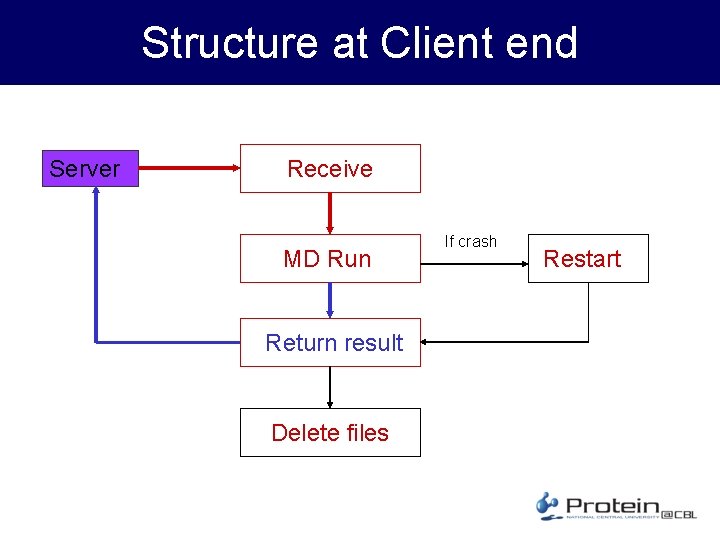
Structure at Client end Server Receive MD Run Return result Delete files If crash Restart

Multi-temperature Annealing • Project suited for multi-temperature runs – Parallel Tempering • Two configurations with energy and temperature (E 1, T 1) and (E 2, T 2) Temperature swapped with probability P = min{1, exp[-(E 2 -E 1)(1/k. T 1 – 1/k. T 2)]} • Mode of operation – Send same peptide at different temperature to many clients; let run; collect; swap T’s by multiple parallel tempering; randomly redistribute peptides with new T’s to clients
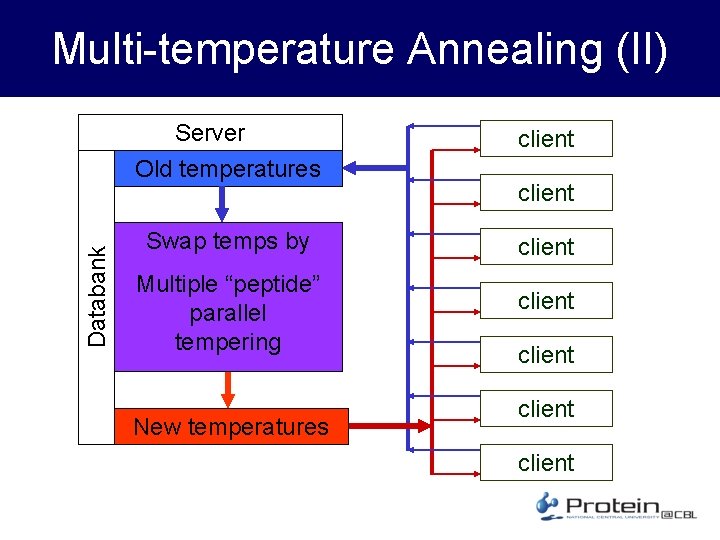
Databank Multi-temperature Annealing (II) Server Old temperatures client Swap temps by client Multiple “peptide” parallel tempering New temperatures client client

Potential of Massive Distributive Computing • Simulation of folding a small peptide for 100 ns – Each run (105 simulation steps; 100 ps) ~100 min PC time – 1000 runs (100 ns) per “fold” ~105 min – Approx. 70 days on single PC running 24 h/day • Ideal client contribute 8 h/day – 100 clients 70 x 3/100 = 2 days per fold – 10, 000 clients 50 folds/day (small peptide) • Mid-sized protein needs > 1 ms to fold – 106 days on single PC – 10, 000 clients ~300 days – 106 clients (!!) ~3 days

Schedule • Launched –August 2002 • Small PC-cluster – October 2002 – In-house runs to learn codes • Infrastructure for Distributive Computation – Installation Gromacs & COSM – January-March 2003 • Test runs – Intra. Laboratory test run – March-October 2003 – NCU test run – July-October 2003 • Launched on WWW – November 20 2003 • Scientific studies – Getting familiar w/ MD and folding of peptides – Looking for ways to increase MD time step

Current status of PAC • Last beta version Pac v 0. 9 – Released on July 15 – To lab CBL members & physics dept – About 25 clients • First alpha version Pac v 1. 0 released October 1 2003 • Current version Pac v 1. 2 – Releases to public on 20 November 2003 – In search of clients • Portal in “Educities” http: //www. educities. edu. tw/ ~3, 700 downloads, ~700 active clients • PC’s in university administrative units • City halls and county government offices • Talks and visits to universities and high schools

Some current Simulations 1 SOL: (20 res. ) A Pip 2 and FActin-Binding Site Of Gelsolin, Residue 150169. One helix. 1 ZDD: (35 res. ) Disulfide-Stabilized Mini Protein A Domain. Two helices. 1 L 2 Y: (20 res. ) NMR Structure Of Trp-Cage Miniprotein Construct Tc 5 B; synthetic.

A small test case – 1 SOL • Target peptide – 1 SOL. pdb – 20 amino acids; 3 -loop helix and 1 hairpin; 352 atoms; ~4000 bonds interaction – Unit time step= 1 fs • Compare constant temperature and paralleltempering – Constant T @ 300 K – Parallel-tempering with about 20 peptides, results returned to server for swapping after each “run”, or 105 time steps (100 ps)
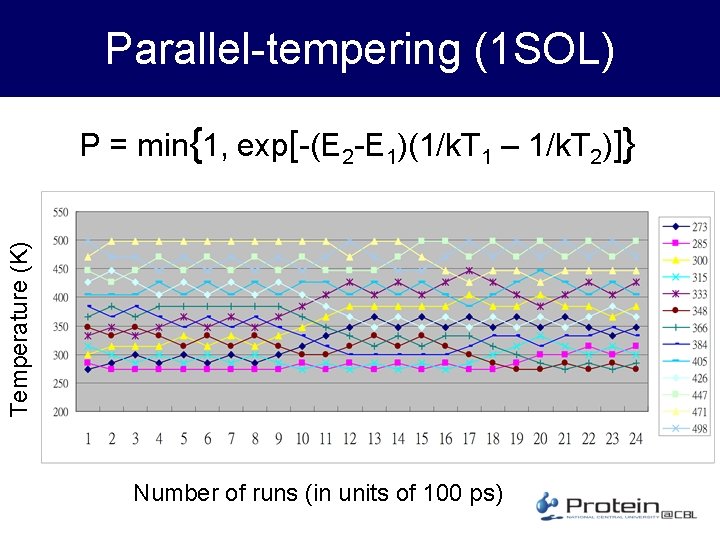
Parallel-tempering (1 SOL) Temperature (K) P = min{1, exp[-(E 2 -E 1)(1/k. T 1 – 1/k. T 2)]} Number of runs (in units of 100 ps)
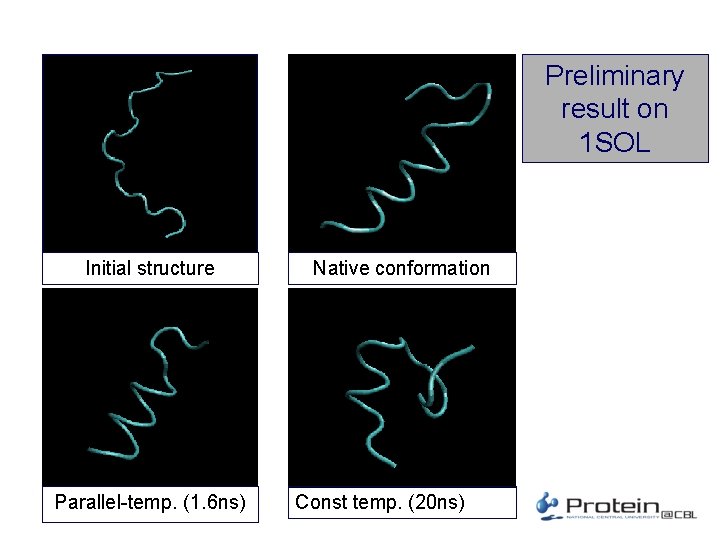
Preliminary result on 1 SOL Initial structure Parallel-temp. (1. 6 ns) Native conformation Const temp. (20 ns)

A second test case – 1 L 2 Y • Simulation target – Trp-Cage • 20 amino acids, 2 helical loops • A short, artificial and fold-by-itself peptide • Have been simulated with AMBER • Folding mechanism not well understood

Temperature (K) A case in swap History (1 L 2 Y) Number of runs (in units of 100 ps)
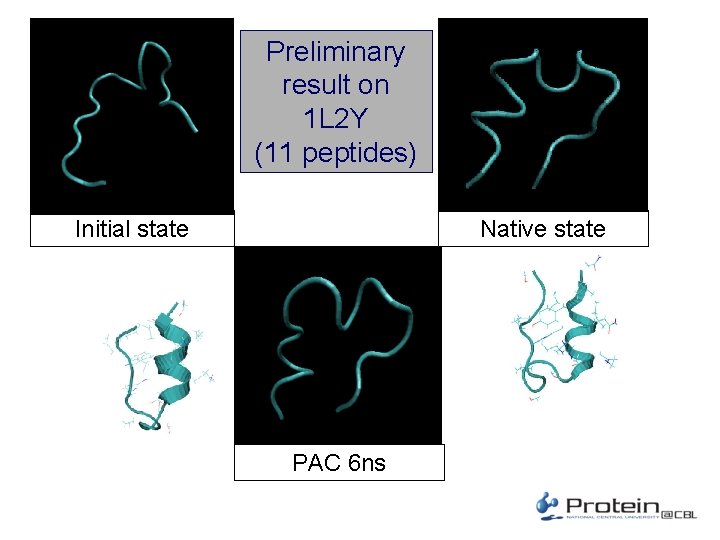
Preliminary result on 1 L 2 Y (11 peptides) Initial state Native state PAC 6 ns

Speeding up simulation - Separating the fast from slow modes • Fast modes associated with bonded interactions – Bond-stretching vibrations ~ 10 -20 fs – Bond-angle bending vibrations ~ 20 -40 fs • Slow modes associated with dihedral angles – – Of the order of 0. 1 ns Alpha-helix folds in ~ 1 -10 ns Beta-sheets folds in ~ 10 -100 ns Native structure ~ 1 ms -1 s
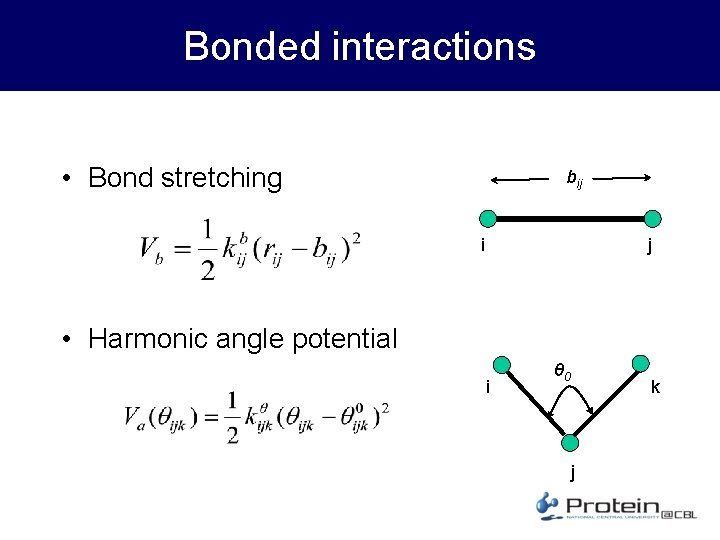
Bonded interactions • Bond stretching bij i j • Harmonic angle potential i θ 0 k j
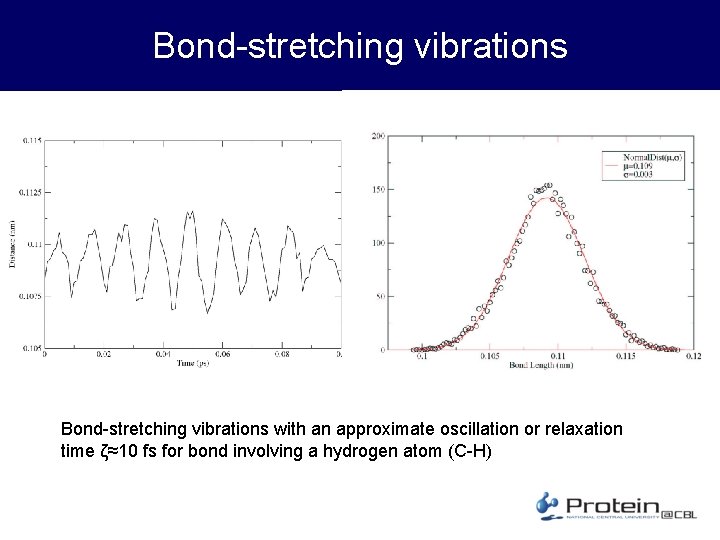
Bond-stretching vibrations with an approximate oscillation or relaxation time ζ≈10 fs for bond involving a hydrogen atom (C-H)
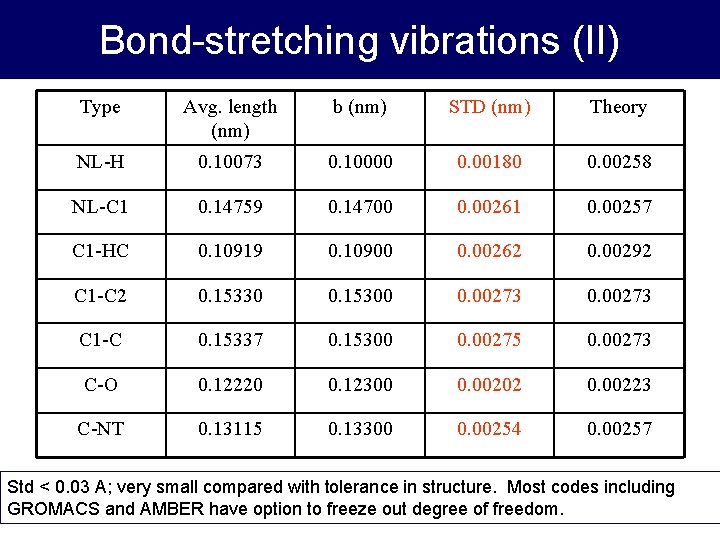
Bond-stretching vibrations (II) Type Avg. length (nm) b (nm) STD (nm) Theory NL-H 0. 10073 0. 10000 0. 00180 0. 00258 NL-C 1 0. 14759 0. 14700 0. 00261 0. 00257 C 1 -HC 0. 10919 0. 10900 0. 00262 0. 00292 C 1 -C 2 0. 15330 0. 15300 0. 00273 C 1 -C 0. 15337 0. 15300 0. 00275 0. 00273 C-O 0. 12220 0. 12300 0. 00202 0. 00223 C-NT 0. 13115 0. 13300 0. 00254 0. 00257 Std < 0. 03 A; very small compared with tolerance in structure. Most codes including GROMACS and AMBER have option to freeze out degree of freedom.
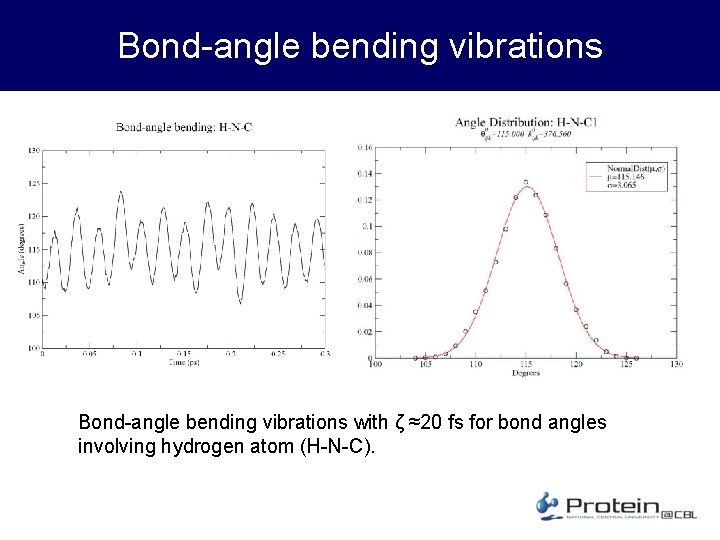
Bond-angle bending vibrations with ζ ≈20 fs for bond angles involving hydrogen atom (H-N-C).
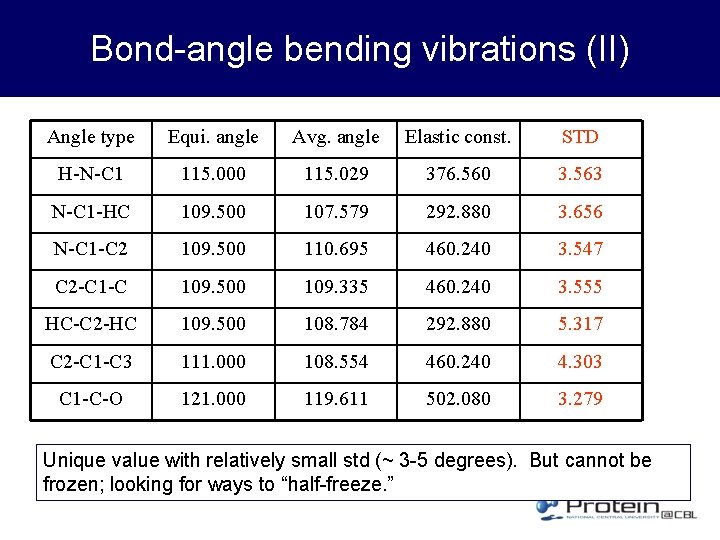
Bond-angle bending vibrations (II) Angle type Equi. angle Avg. angle Elastic const. STD H-N-C 1 115. 000 115. 029 376. 560 3. 563 N-C 1 -HC 109. 500 107. 579 292. 880 3. 656 N-C 1 -C 2 109. 500 110. 695 460. 240 3. 547 C 2 -C 109. 500 109. 335 460. 240 3. 555 HC-C 2 -HC 109. 500 108. 784 292. 880 5. 317 C 2 -C 1 -C 3 111. 000 108. 554 460. 240 4. 303 C 1 -C-O 121. 000 119. 611 502. 080 3. 279 Unique value with relatively small std (~ 3 -5 degrees). But cannot be frozen; looking for ways to “half-freeze. ”
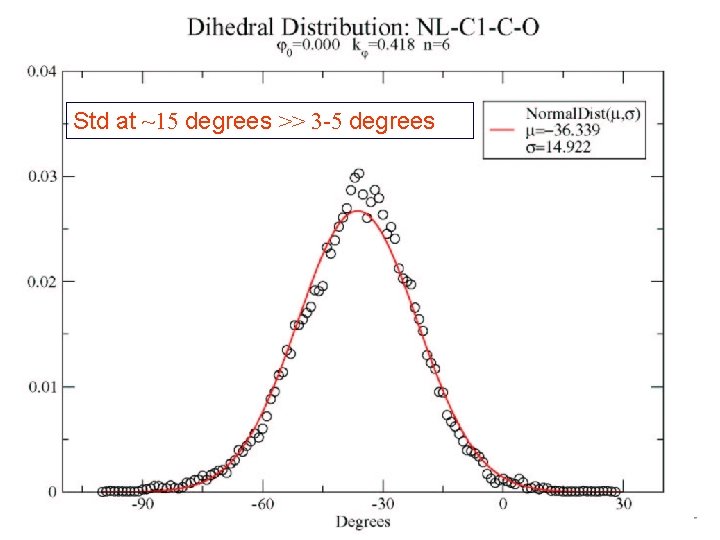
Std at ~15 degrees >> 3 -5 degrees
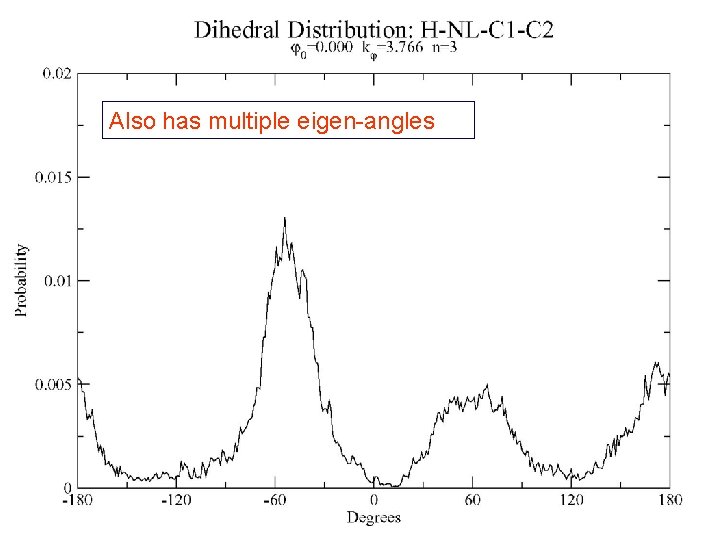
Also has multiple eigen-angles

Current and future efforts • Computing facility – expand the base of PAC clients; target 10, 000 • Data management – efficient server-client protocol – efficient management and analysis of data when client number is large • Running simulations – optimum implementation of parallel tempering – reduce size of water box • Dealing with fast modes – freeze bond stretching – isolate bond-angle bending deg. of freedom for special treatment; new (heavy) code-writing – target time-step: > 20 fs; ultimately 100 fs

The Team • Funded by NRPGM/NSC • Computational Biology Laboratory Physics Dept & Life Sciences Dept National Central University – – – – PI: Professor HC Lee (Phys & LS/NCU) Jia-Lin Lo (Ph. D student) Jun-Ping Yiu (MSc Res. Assistant) Chien-Hao Wei (MSc RA) Engin Lee ( MSc student ) Dr. Richard Tseng (PDF, since May 2004) Visiting scientist: physicist/computer specialist (TBA)
Website client_stats http: //protein. ncu. edu. tw

Please visit http: //protein. ncu. edu. tw and let your PC take part in this project while you sleep Thank you
- Slides: 39