Mass Spectrometry based Proteomics and Metabolomics Guoan Zhang

Mass Spectrometry based Proteomics and Metabolomics Guoan Zhang Director Proteomics and Metabolomics Core Facility Proteomics & Metabolomics Core Facility @ WCM

Proteomics & Metabolomics Core Facility @Weill Cornell • MS- based protein/metabolite analysis • Opened for services – Oct 2016 for proteomics – May 2017 for metabolomics Proteomics & Metabolomics Core Facility @
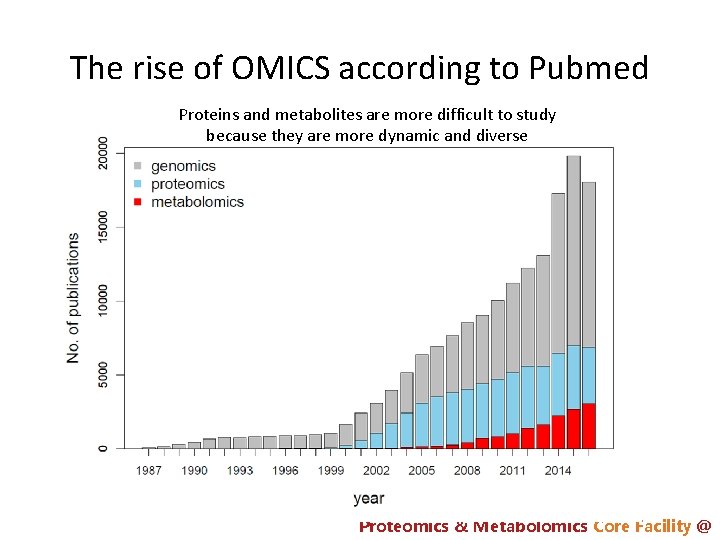
The rise of OMICS according to Pubmed Proteins and metabolites are more difficult to study because they are more dynamic and diverse Proteomics & Metabolomics Core Facility @

Coverage comparison: genomics > proteomics >> metabolomics • Genomics – 100% coverage • Proteomics – Can identify ~10, 000 protein vs ~20, 000 genes (human) – Low sequence coverage • Metabolomics – Can detect ~1000 metabolites vs ? total metabolites – Needs multiple platforms Proteomics & Metabolomics Core Facility @

Mass spectrometry Proteomics & Metabolomics Core Facility @
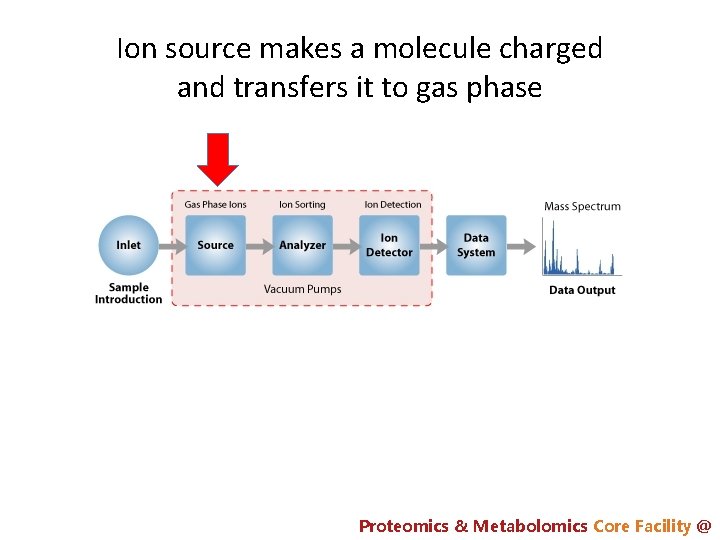
Ion source makes a molecule charged and transfers it to gas phase Proteomics & Metabolomics Core Facility @
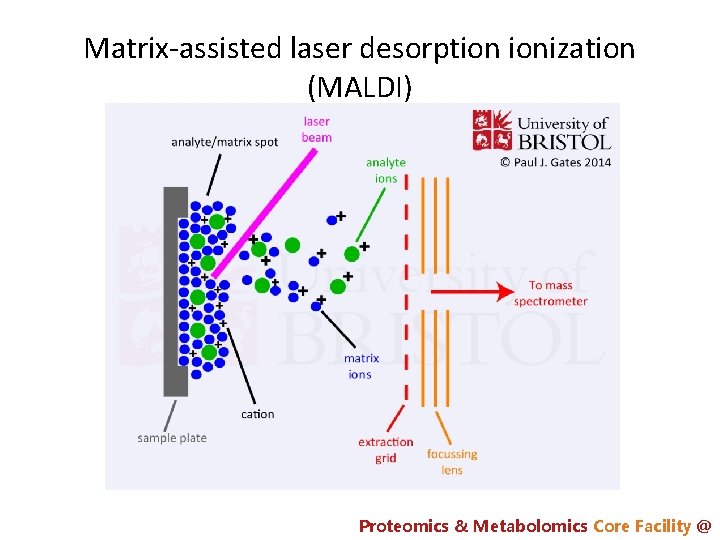
Matrix-assisted laser desorption ionization (MALDI) Proteomics & Metabolomics Core Facility @
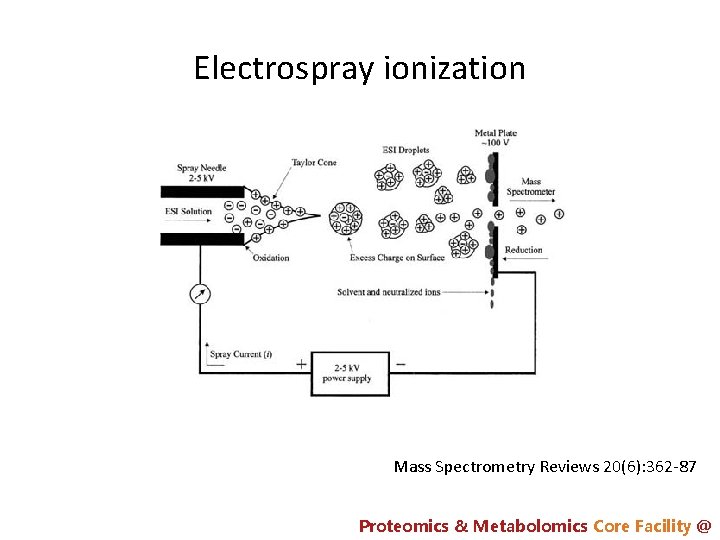
Electrospray ionization Mass Spectrometry Reviews 20(6): 362 -87 Proteomics & Metabolomics Core Facility @
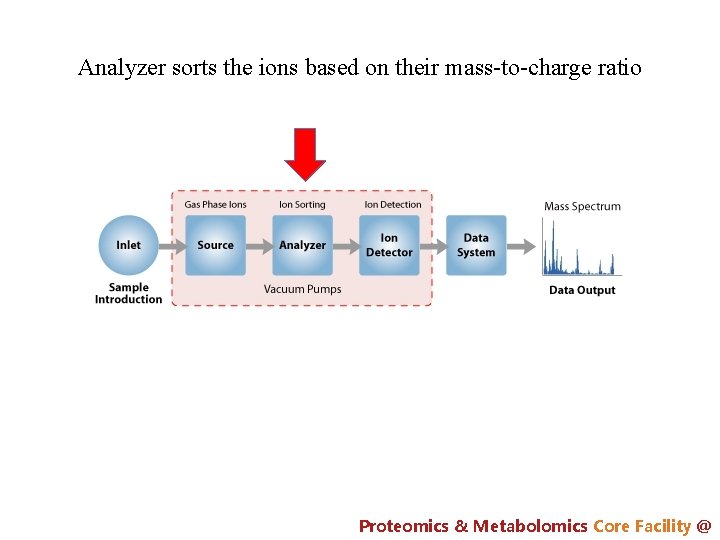
Analyzer sorts the ions based on their mass-to-charge ratio Proteomics & Metabolomics Core Facility @
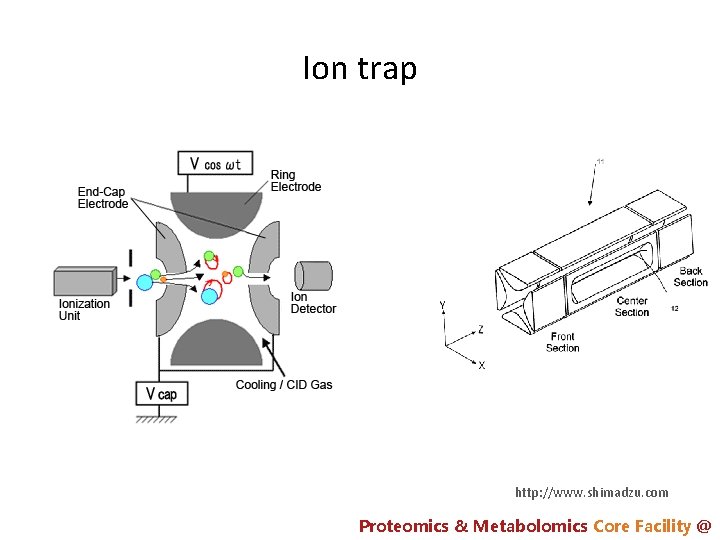
Ion trap http: //www. shimadzu. com Proteomics & Metabolomics Core Facility @
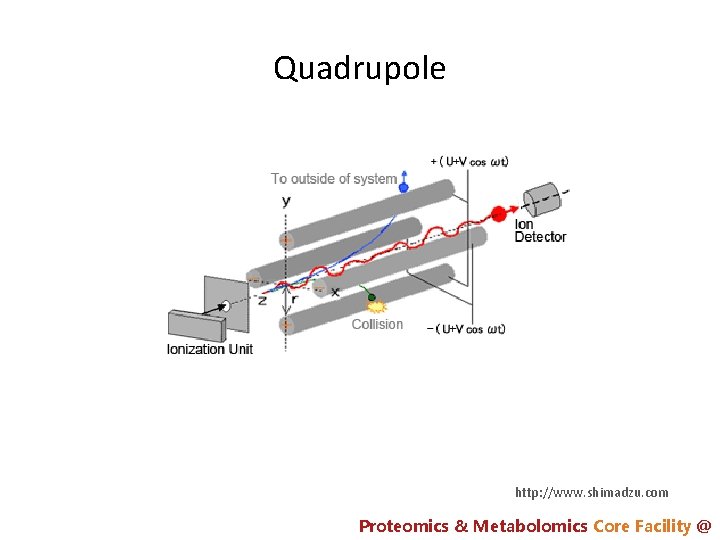
Quadrupole http: //www. shimadzu. com Proteomics & Metabolomics Core Facility @
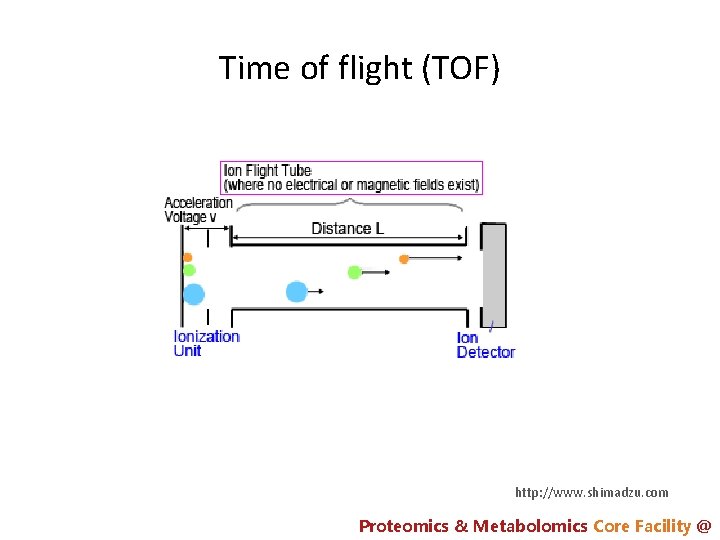
Time of flight (TOF) http: //www. shimadzu. com Proteomics & Metabolomics Core Facility @
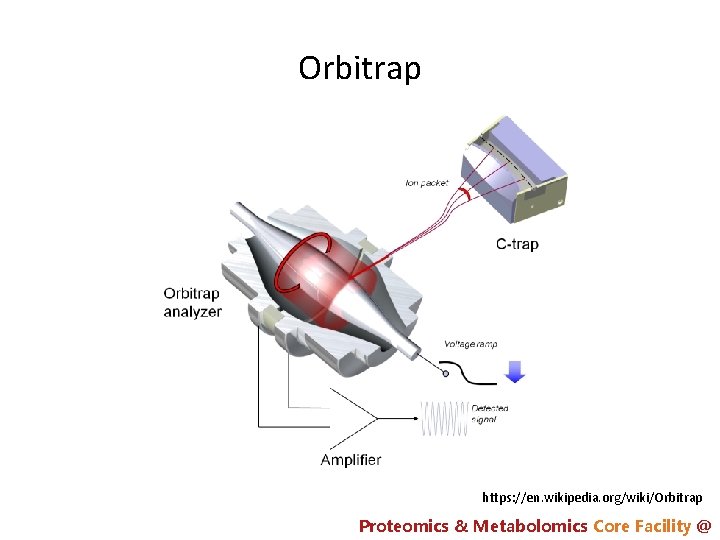
Orbitrap https: //en. wikipedia. org/wiki/Orbitrap Proteomics & Metabolomics Core Facility @
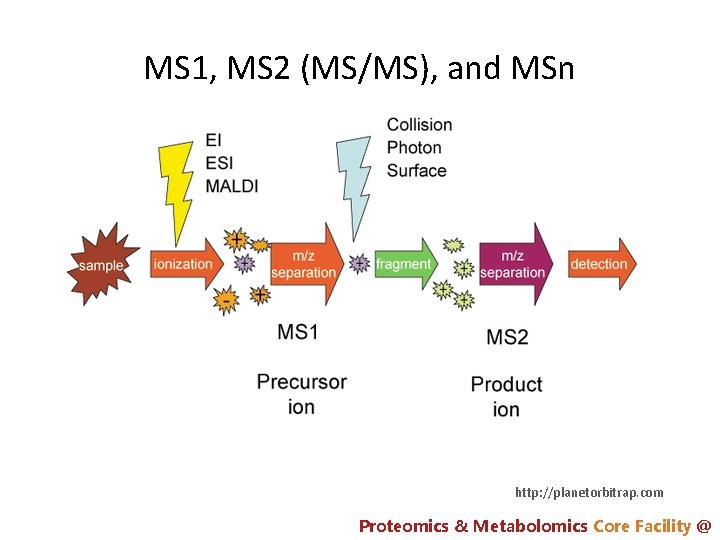
MS 1, MS 2 (MS/MS), and MSn http: //planetorbitrap. com Proteomics & Metabolomics Core Facility @
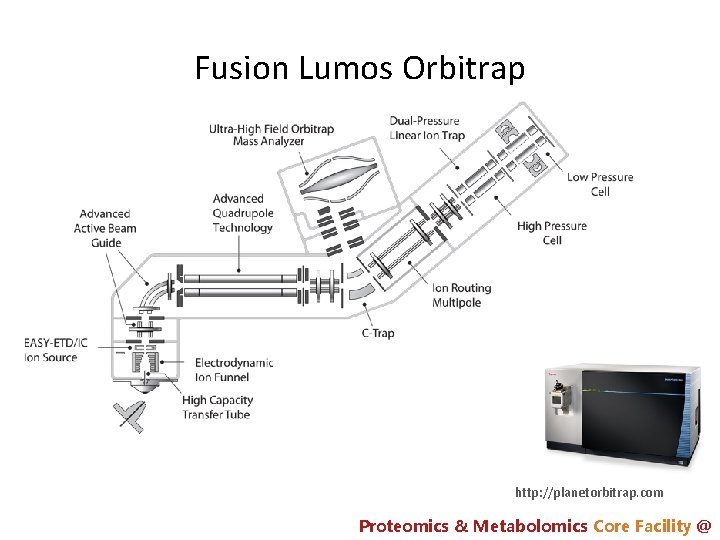
Fusion Lumos Orbitrap http: //planetorbitrap. com Proteomics & Metabolomics Core Facility @
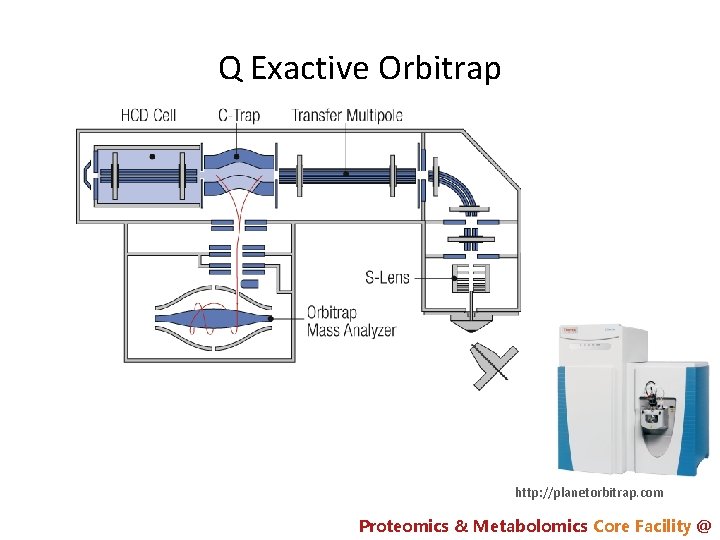
Q Exactive Orbitrap http: //planetorbitrap. com Proteomics & Metabolomics Core Facility @
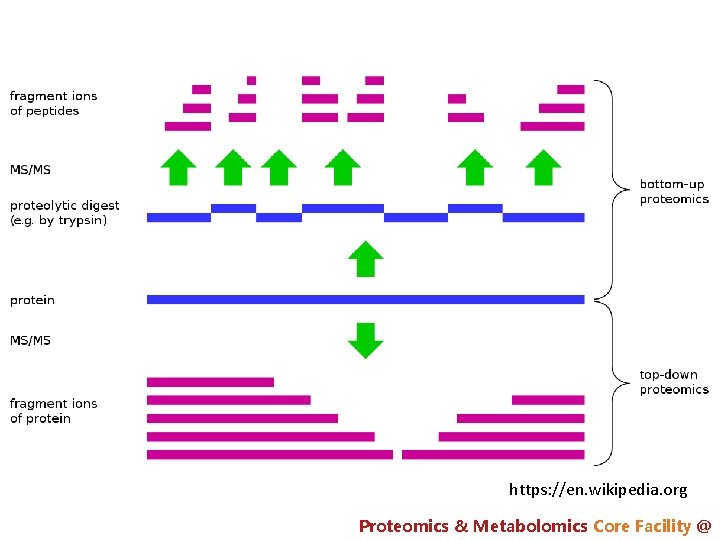
https: //en. wikipedia. org Proteomics & Metabolomics Core Facility @
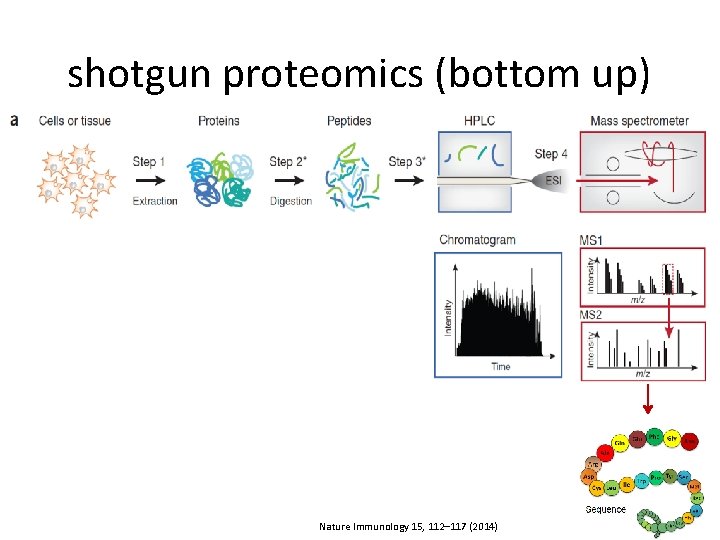
shotgun proteomics (bottom up) Nature Immunology 15, 112– 117 (2014)

Proteomics Services • Protein Identification – ID of proteins from SDS-PAGE and of proteins in-solution. • Post-Translational Modification Analysis – – Phosphorylation; ubiquitination; acetylation; other PTMs. • Protein Quantitation – – Label-free (intensity based / spectral counting); stable isotope labeling by amino acids in cell culture (SILAC); tandem mass tags (TMT); spiked standards. • MS Data Analysis Proteomics & Metabolomics Core Facility @

PTM analysis: affinity enrichment is often required (peptide level) • Phosphorylation (p. Ser, p. Thr, p. Tyr): – Ti. O 2 – IMAC • p. Tyr: – p. Tyr antibody • acetylation: – Ac-antibody • Ubiquitination – di. Gly antibody Proteomics & Metabolomics Core Facility @
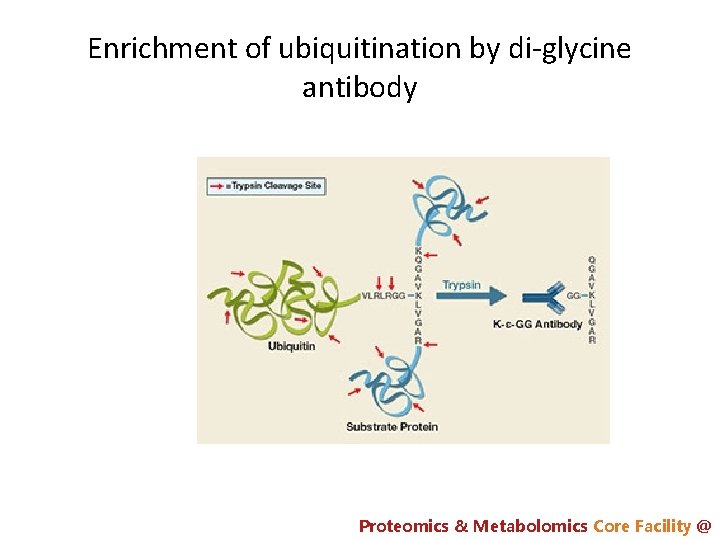
Enrichment of ubiquitination by di-glycine antibody Proteomics & Metabolomics Core Facility @

Quantitation methods • SILAC (Stable Isotope Labeling with Amino acids in Cell culture) • TMT (tandem mass tag) • Label free Proteomics & Metabolomics Core Facility @
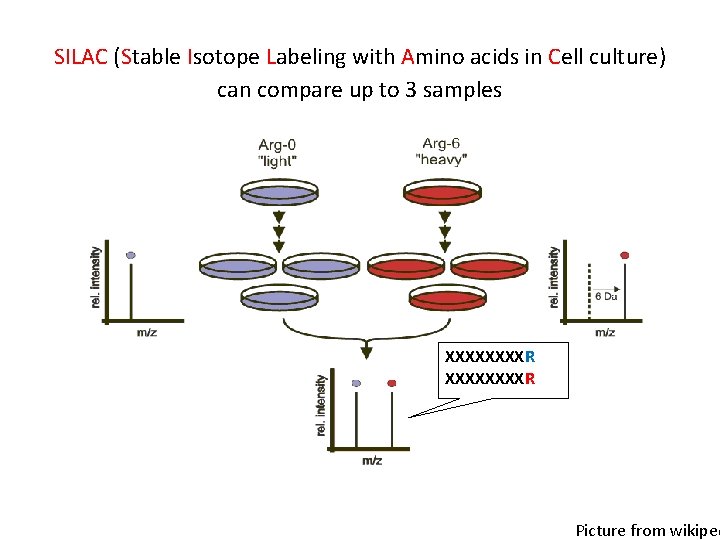
SILAC (Stable Isotope Labeling with Amino acids in Cell culture) can compare up to 3 samples XXXXXXXXR Picture from wikiped
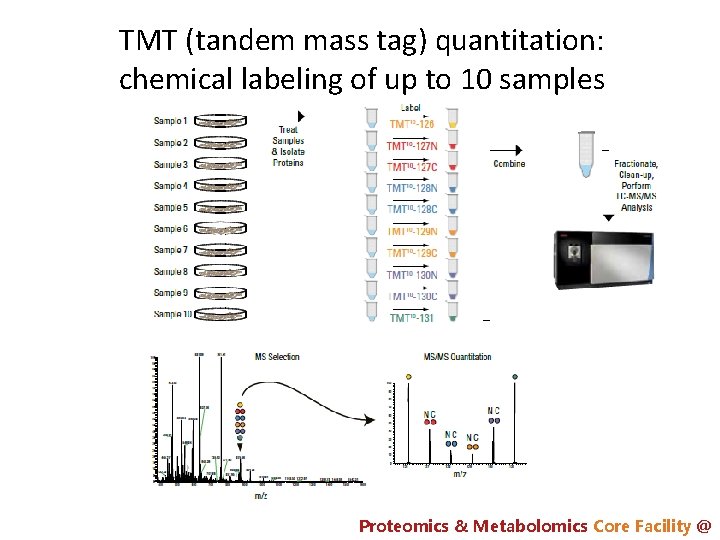
TMT (tandem mass tag) quantitation: chemical labeling of up to 10 samples Proteomics & Metabolomics Core Facility @
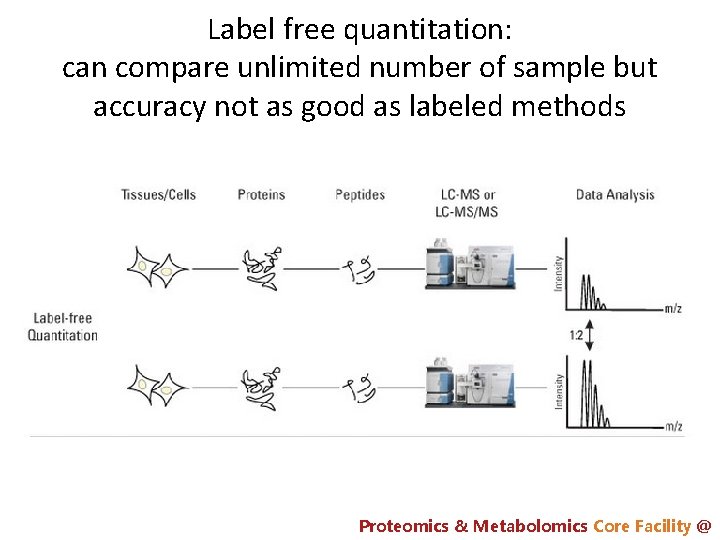
Label free quantitation: can compare unlimited number of sample but accuracy not as good as labeled methods Proteomics & Metabolomics Core Facility @
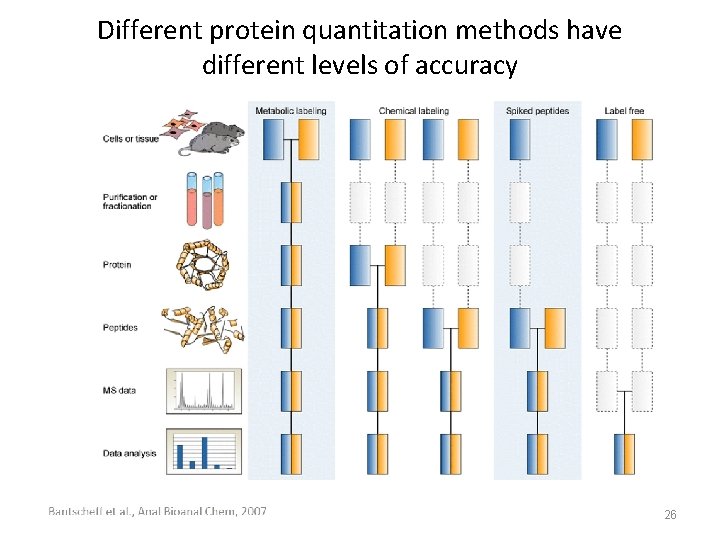
Different protein quantitation methods have different levels of accuracy 26

Comparison of different Quantitation methods Proteomics & Metabolomics Core Facility @
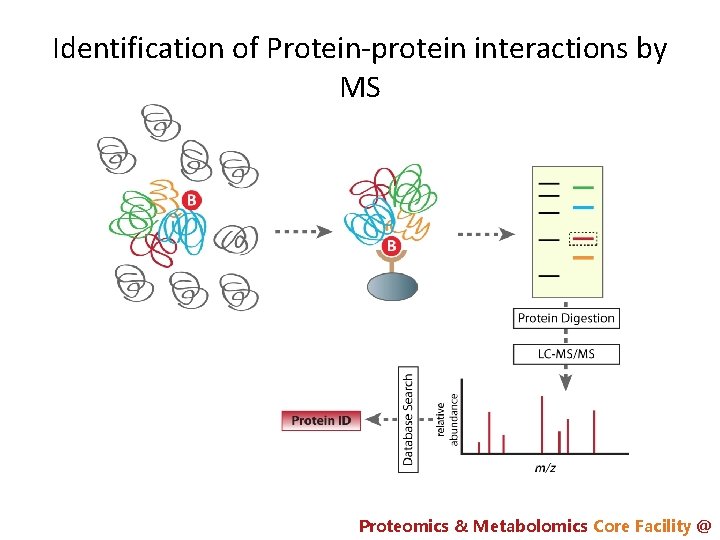
Identification of Protein-protein interactions by MS Proteomics & Metabolomics Core Facility @
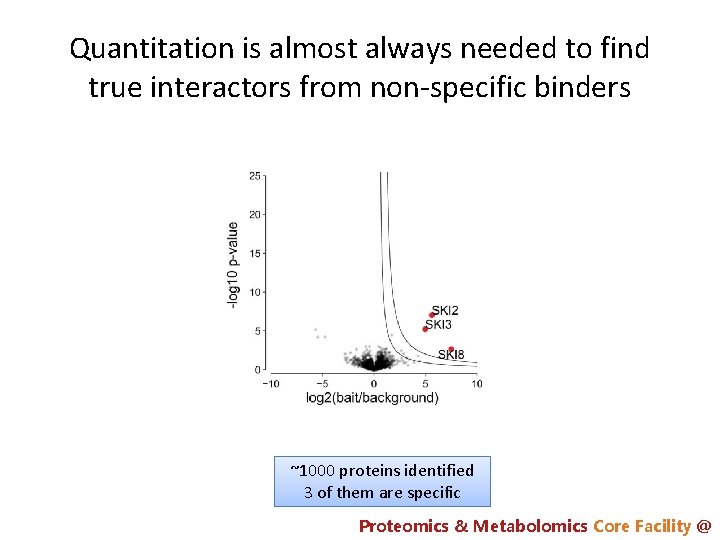
Quantitation is almost always needed to find true interactors from non-specific binders ~1000 proteins identified 3 of them are specific Proteomics & Metabolomics Core Facility @
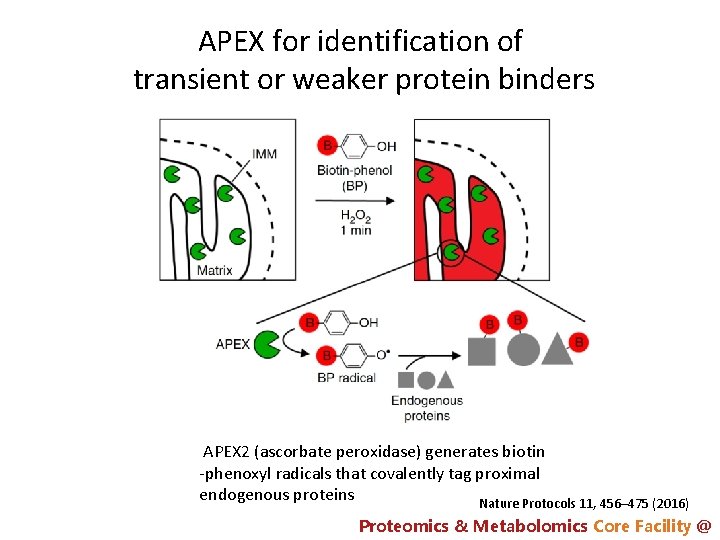
APEX for identification of transient or weaker protein binders APEX 2 (ascorbate peroxidase) generates biotin -phenoxyl radicals that covalently tag proximal endogenous proteins Nature Protocols 11, 456– 475 (2016) Proteomics & Metabolomics Core Facility @
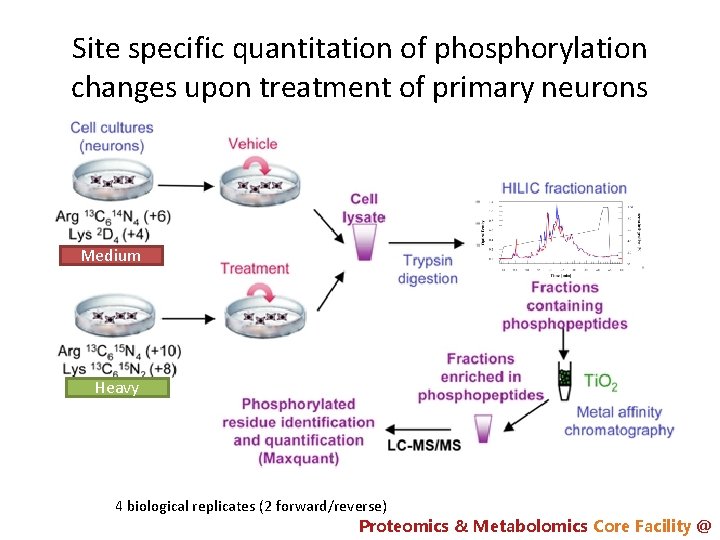
Site specific quantitation of phosphorylation changes upon treatment of primary neurons Medium Heavy 4 biological replicates (2 forward/reverse) Proteomics & Metabolomics Core Facility @
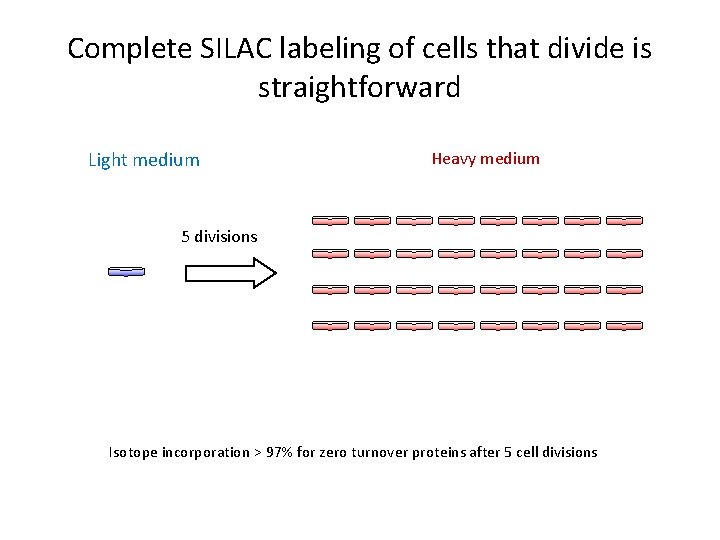
Complete SILAC labeling of cells that divide is straightforward Light medium Heavy medium 5 divisions Isotope incorporation > 97% for zero turnover proteins after 5 cell divisions
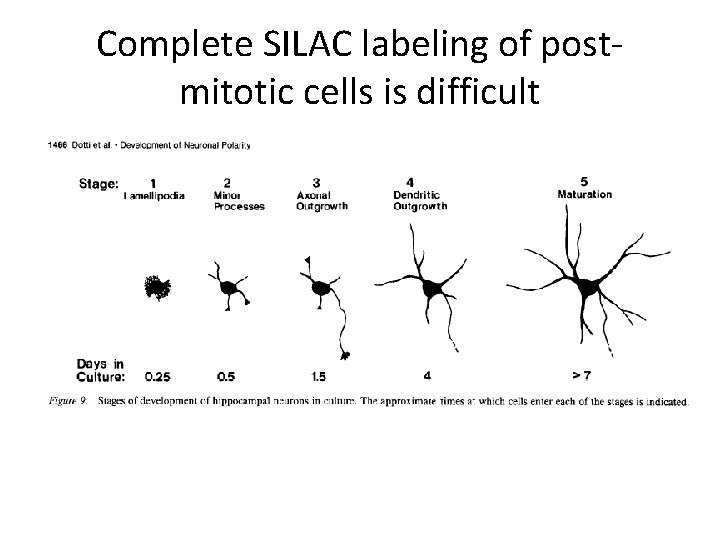
Complete SILAC labeling of postmitotic cells is difficult
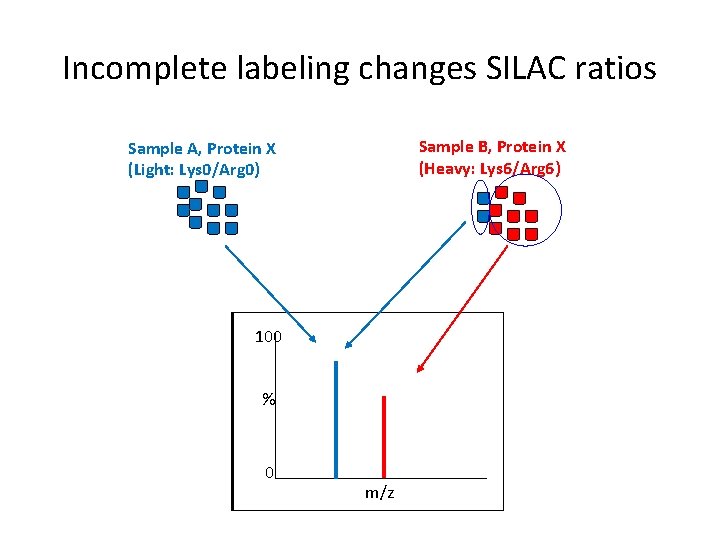
Incomplete labeling changes SILAC ratios Sample B, Protein X (Heavy: Lys 6/Arg 6) Sample A, Protein X (Light: Lys 0/Arg 0) 100 % 0 m/z
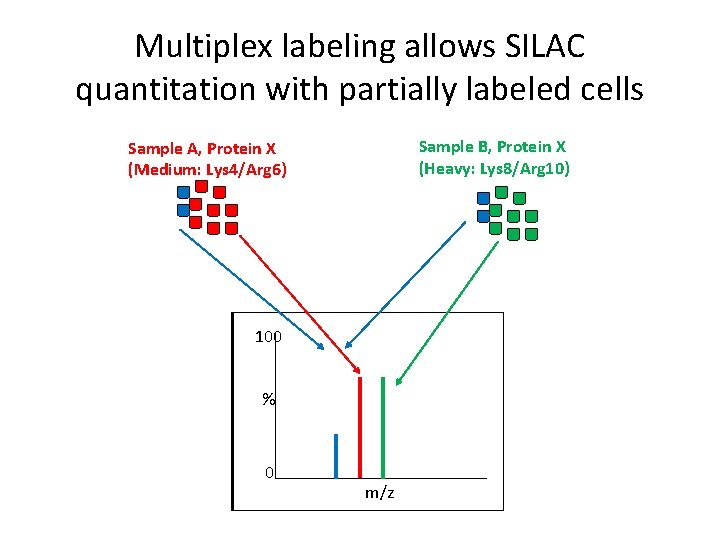
Multiplex labeling allows SILAC quantitation with partially labeled cells Sample B, Protein X (Heavy: Lys 8/Arg 10) Sample A, Protein X (Medium: Lys 4/Arg 6) 100 % 0 m/z
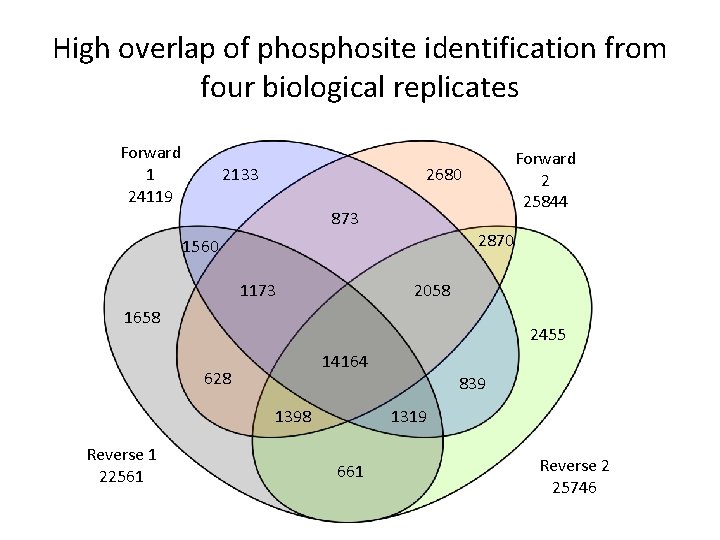
High overlap of phosite identification from four biological replicates Forward 1 24119 2133 Forward 2 25844 2680 873 2870 1560 1173 2058 1658 2455 14164 628 1398 Reverse 1 22561 839 1319 661 Reverse 2 25746
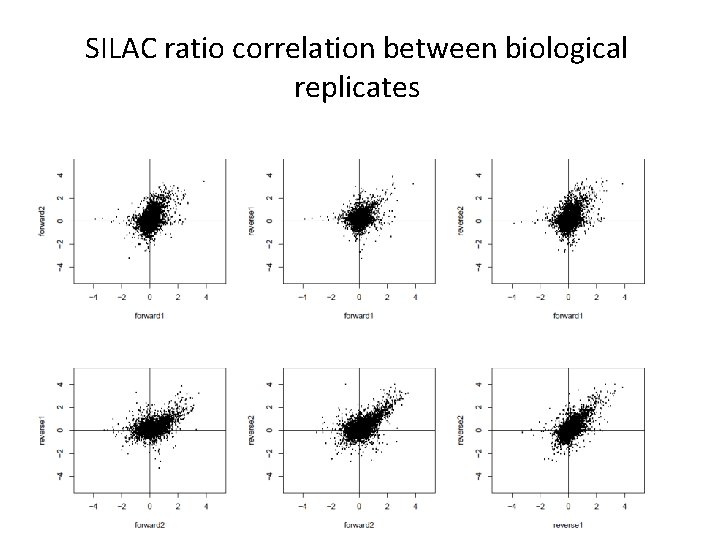
SILAC ratio correlation between biological replicates
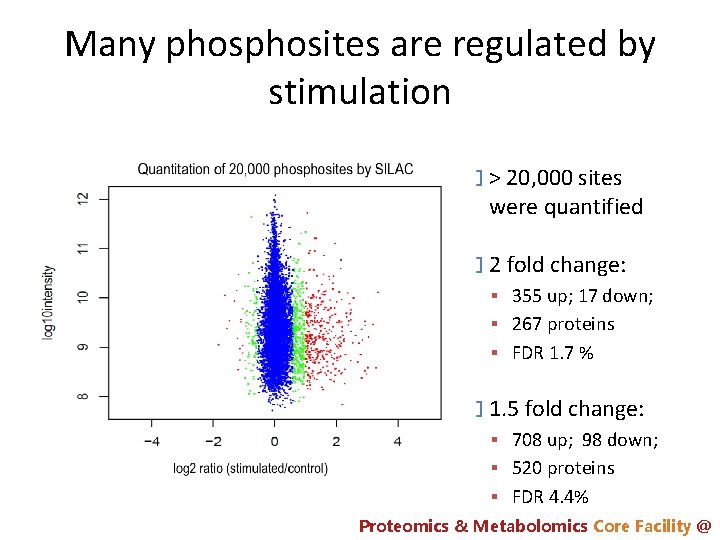
Many phosites are regulated by stimulation � > 20, 000 sites were quantified � 2 fold change: 355 up; 17 down; 267 proteins FDR 1. 7 % � 1. 5 fold change: 708 up; 98 down; 520 proteins FDR 4. 4% Proteomics & Metabolomics Core Facility @
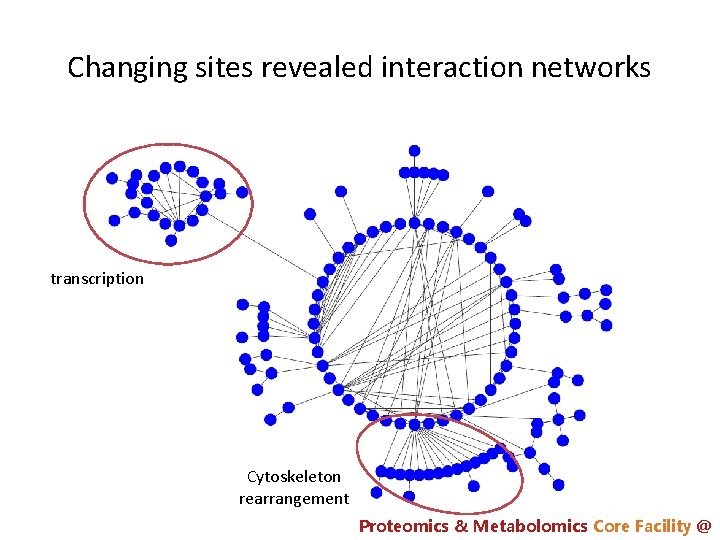
Changing sites revealed interaction networks transcription Cytoskeleton rearrangement Proteomics & Metabolomics Core Facility @
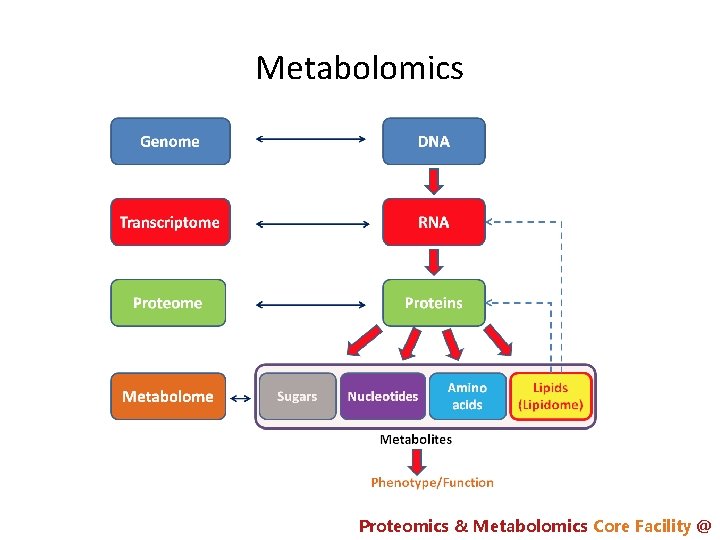
Metabolomics Proteomics & Metabolomics Core Facility @

Many analytical platforms are used for metabolomics • • NMR GC/MS LC/MS CE/MS • LC/MS has the best coverage and dynamic range

Untargeted metabolite profiling • Attempts to identify and quantify all compounds • Needs standard MS/MS library Proteomics & Metabolomics Core Facility @

Targeted polar metabolite profiling Sample extraction 80% methanol LCMS HILIC (hydrophilic interaction chromatography) Data analysis A list of >200 compounds with known m/z and RT Proteomics & Metabolomics Core Facility @
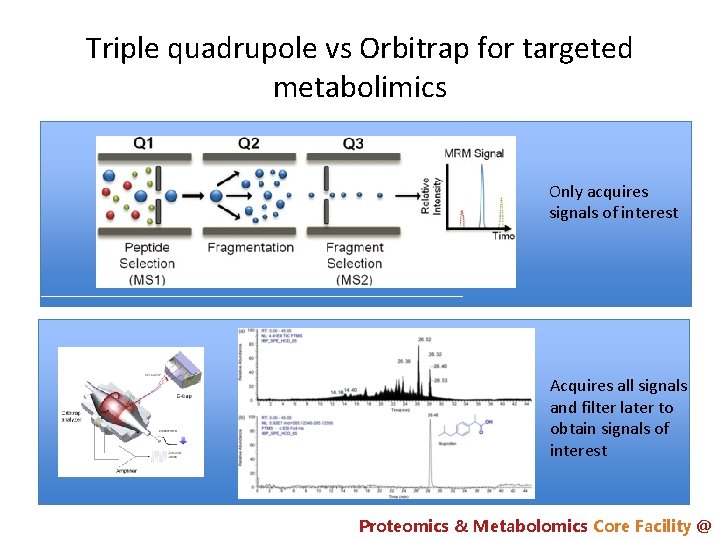
Triple quadrupole vs Orbitrap for targeted metabolimics Only acquires signals of interest Acquires all signals and filter later to obtain signals of interest Proteomics & Metabolomics Core Facility @
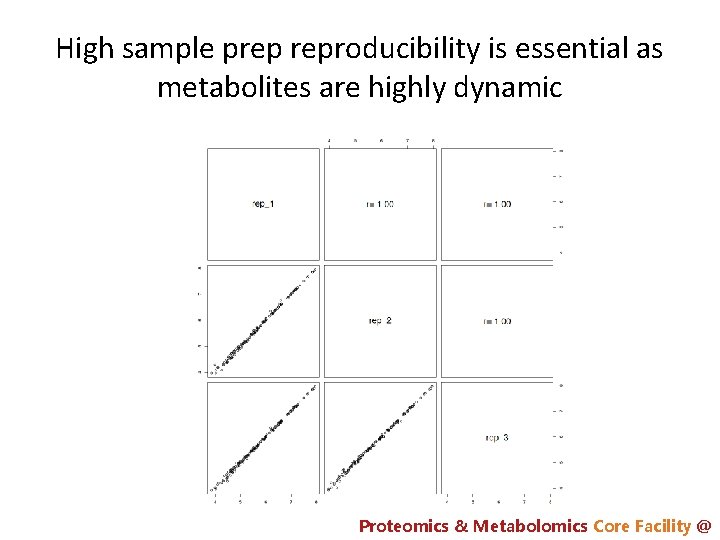
High sample prep reproducibility is essential as metabolites are highly dynamic Proteomics & Metabolomics Core Facility @
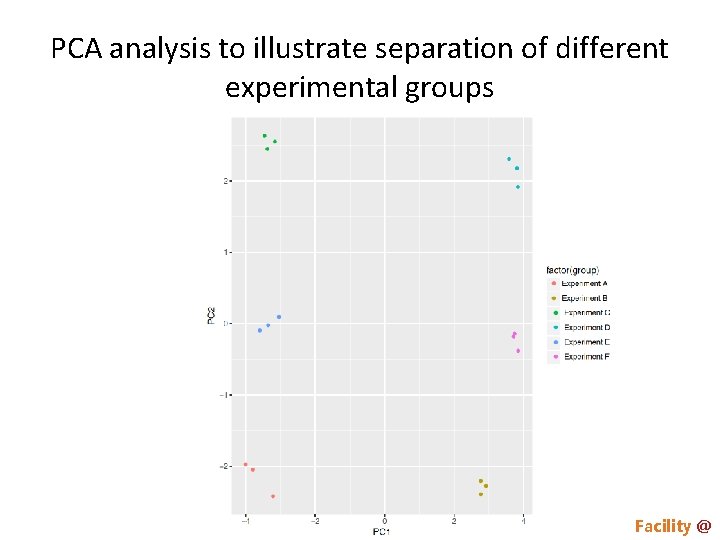
PCA analysis to illustrate separation of different experimental groups Proteomics & Metabolomics Core Facility @
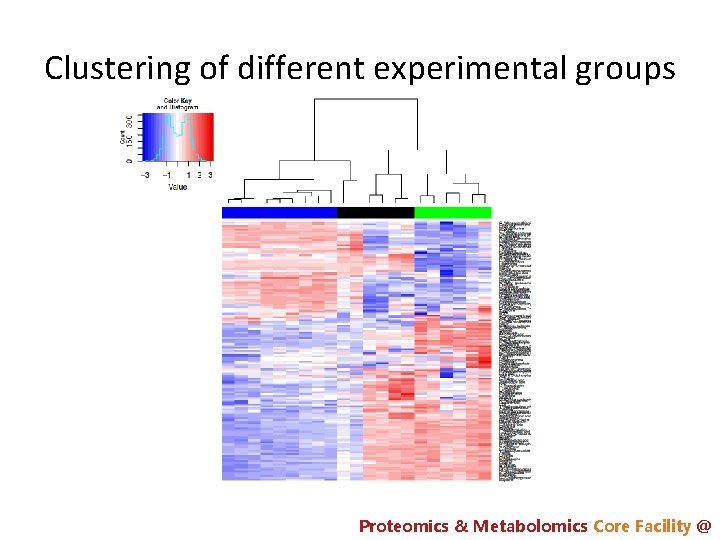
Clustering of different experimental groups Proteomics & Metabolomics Core Facility @
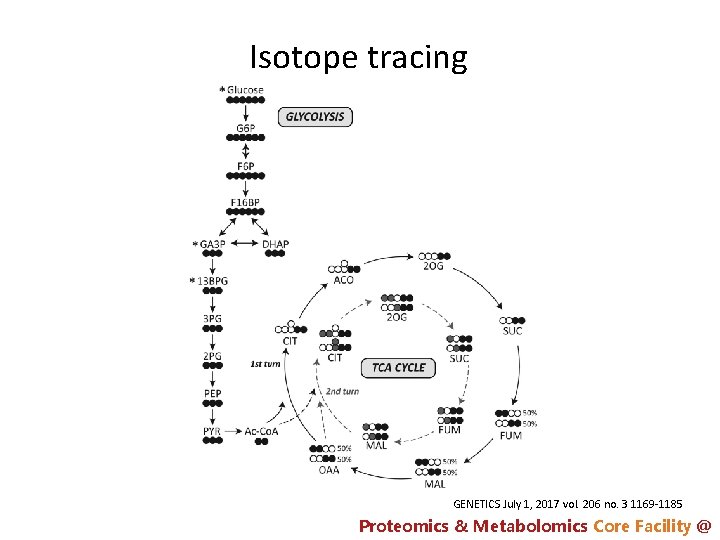
Isotope tracing GENETICS July 1, 2017 vol. 206 no. 3 1169 -1185 Proteomics & Metabolomics Core Facility @
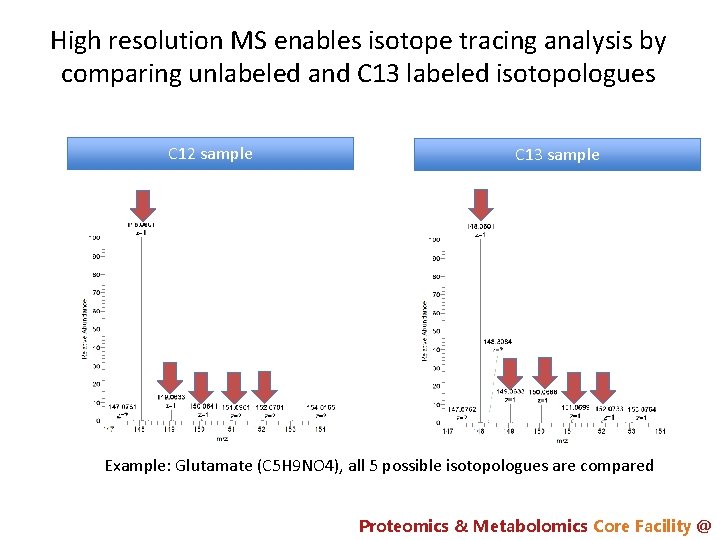
High resolution MS enables isotope tracing analysis by comparing unlabeled and C 13 labeled isotopologues C 12 sample C 13 sample Example: Glutamate (C 5 H 9 NO 4), all 5 possible isotopologues are compared Proteomics & Metabolomics Core Facility @
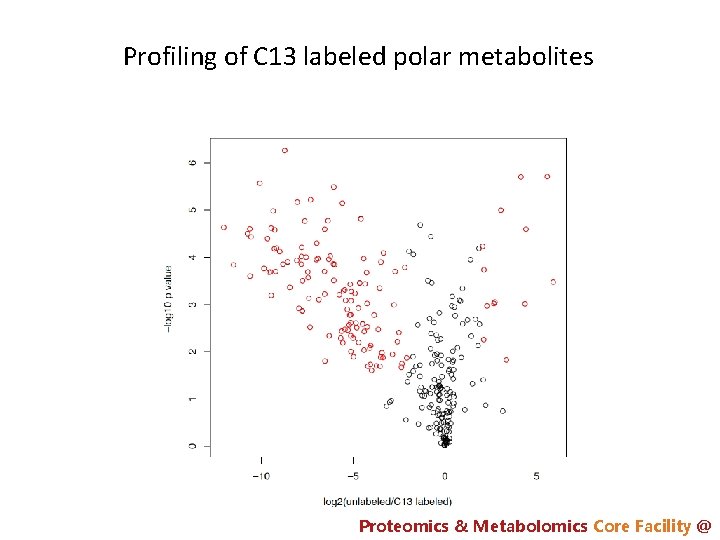
Profiling of C 13 labeled polar metabolites Proteomics & Metabolomics Core Facility @
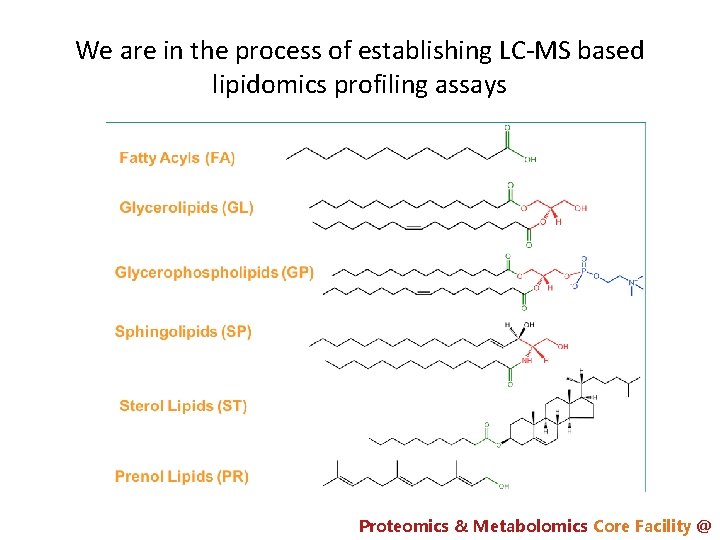
We are in the process of establishing LC-MS based lipidomics profiling assays Proteomics & Metabolomics Core Facility @

Contact us • Website – https: //wcmc. corefacilities. org/service_center/show_ external/3474 • Director – Guoan Zhang, Ph. D. – Office: BRB-1616 – Tel: (646) 962 -6222 – Email: guz 2004@med. cornell. edu • Lab – BRB-1552 – Tel: 646 -962 -9739 Proteomics & Metabolomics Core Facility @
- Slides: 52