Mapping the Human Genome Genetic Mapping Physical Mapping


















- Slides: 52

Mapping the Human Genome Genetic Mapping Physical Mapping DNA Sequencing

Physical Mapping Systems Yeast Artificial Chromosomes (YACs) 200 -2000 kb Bacteriophage P 1 90 kb Cosmids 40 kb Bacteriophage l 9 -23 kb

Large Fragment Cloning --- Hybrid cells : multiple chromosomes : 100 -3000 Mbp --- Mono-chromosomal hybrids : 50 - 300 Mbp --- Sub-chromosomal hybrids : 1 -50 Mbp --- Double-minutes : 0. 3 -2 Mbp --- Yeast Artificial Chromosomes : YACs : 0. 05 -2 Mbp Insert Size --- BACs (F-plasmid derived) : - 300 kb --- PACs (large P 1 clones) : - 300 kb --- T 4 -packaging system : - 120 kb ( - 300 kb) --- EBV derived vectors : - 180 kb --- P 1 + P 1 packaging : - 80 - 90 kb --- Cosmid vectors : 35 - 45 kb --- Lambda replacement vectors : 8 - 22 kb --- Micro-dissection clones : - 1. 5 kb

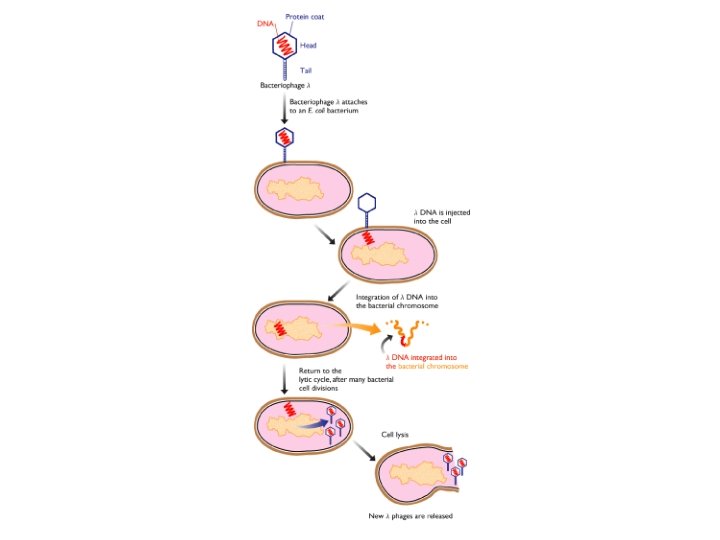
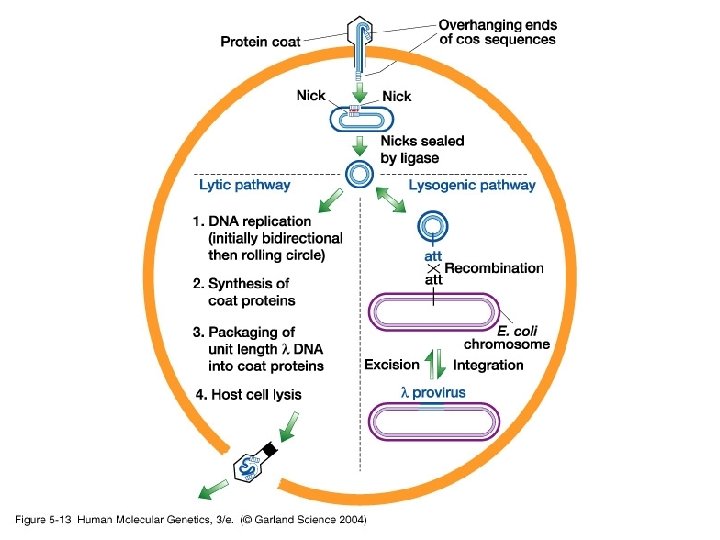
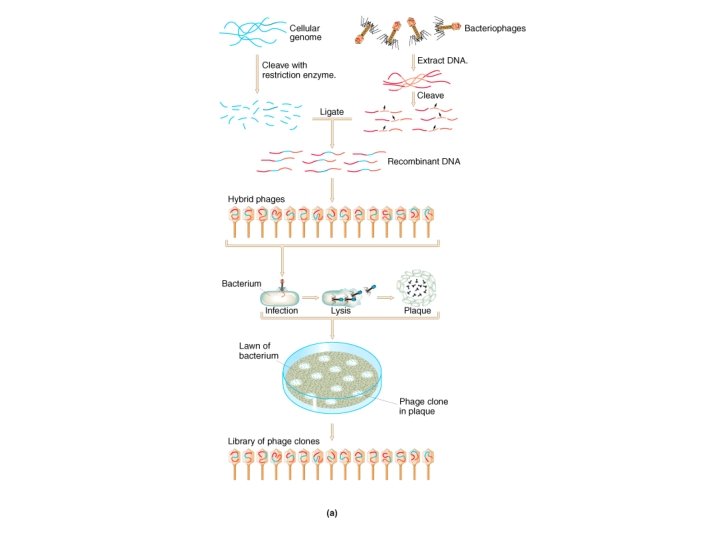
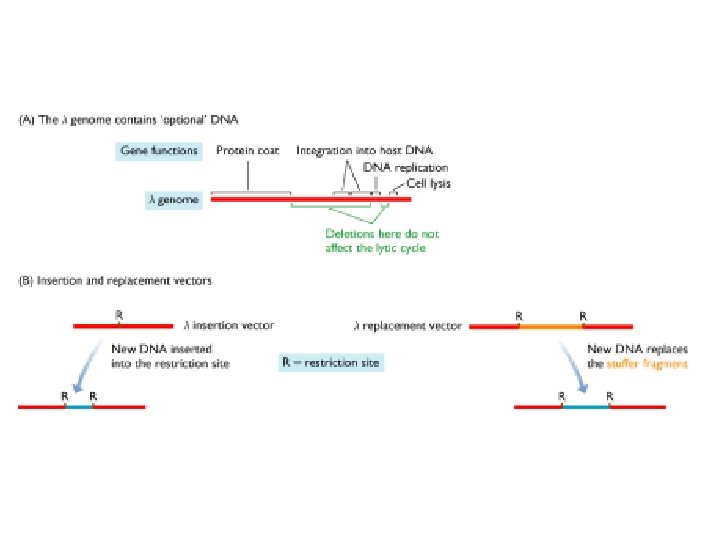
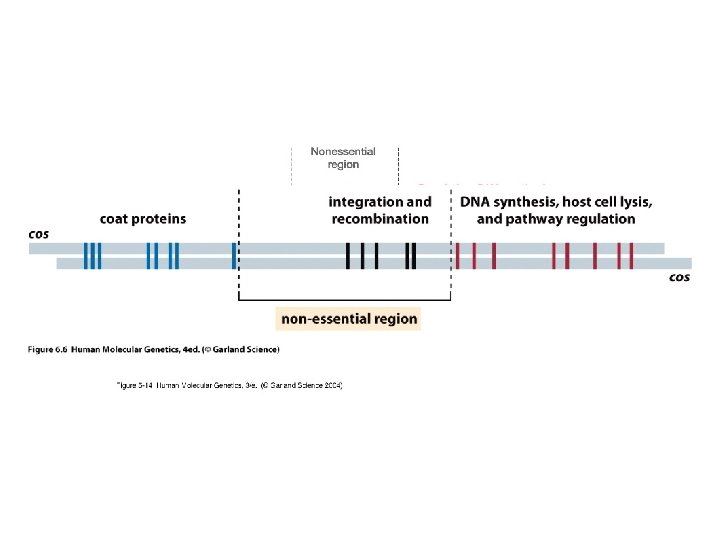
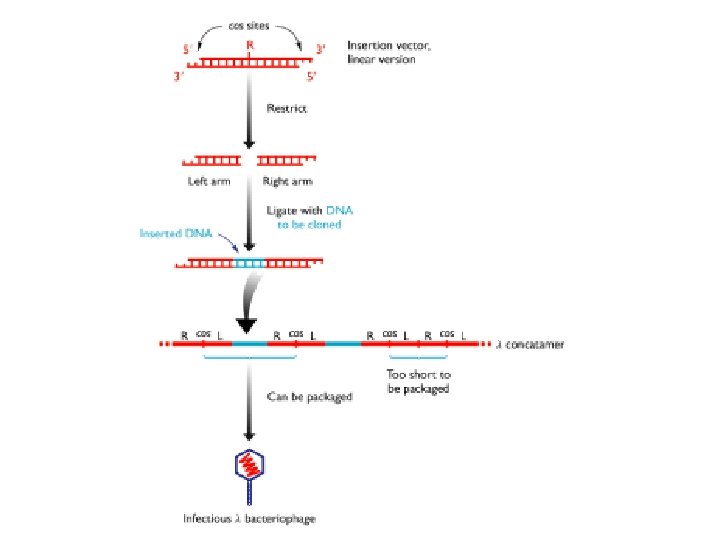
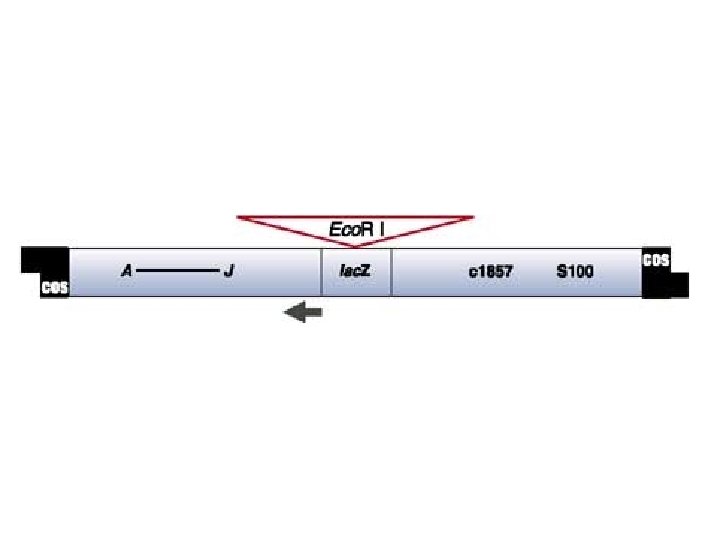
Bacteriophage Lambda • Insertion Vectors – c. DNA cloning and expression GT 10, GT 11, Zap • Replacement Vectors – Genomic cloning EMBL 3, EMBL 4








Lambda gt 11 • Insertion vector • Small inserts (c. DNAs) • Expression cloning


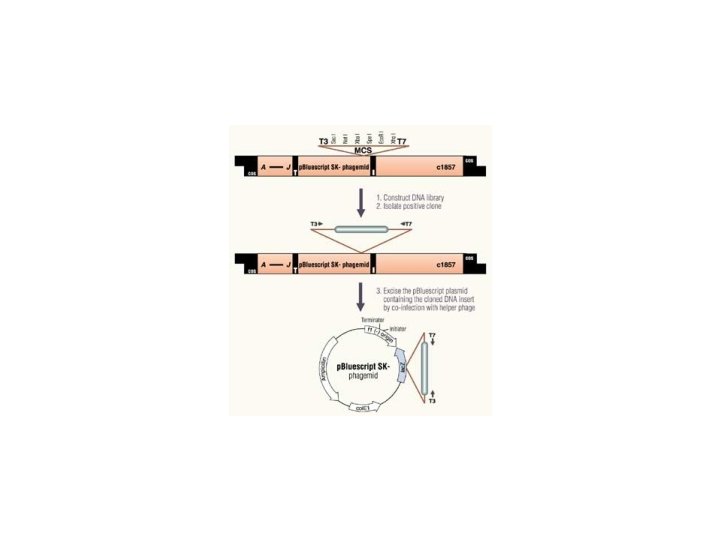
Lambda Zap • • Insertion vector Small inserts (c. DNAs) Expression cloning In vivo excision of inserts



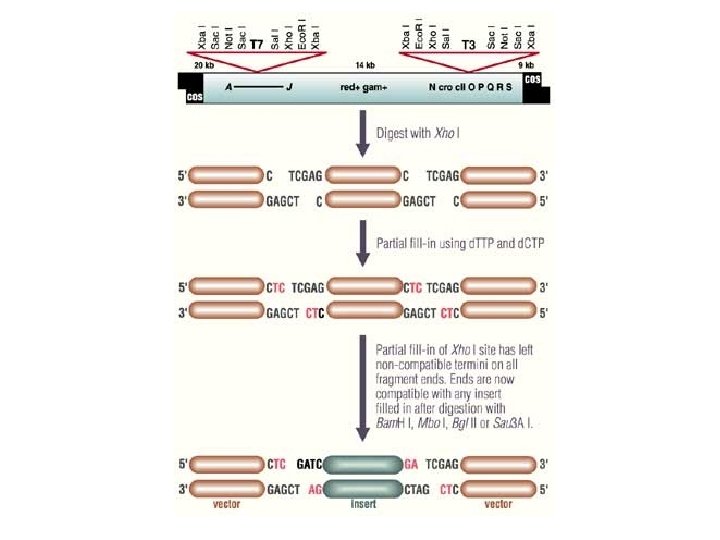
Lambda Fix • • Replacement vector Larger inserts Genomic DNA Spi selection


Lambda EMBL 3 • • Replacement vector Larger inserts Genomic DNA Spi selection


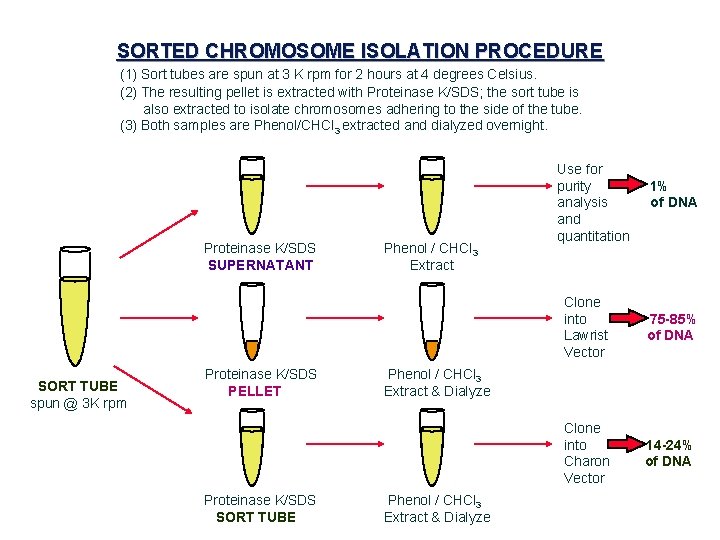
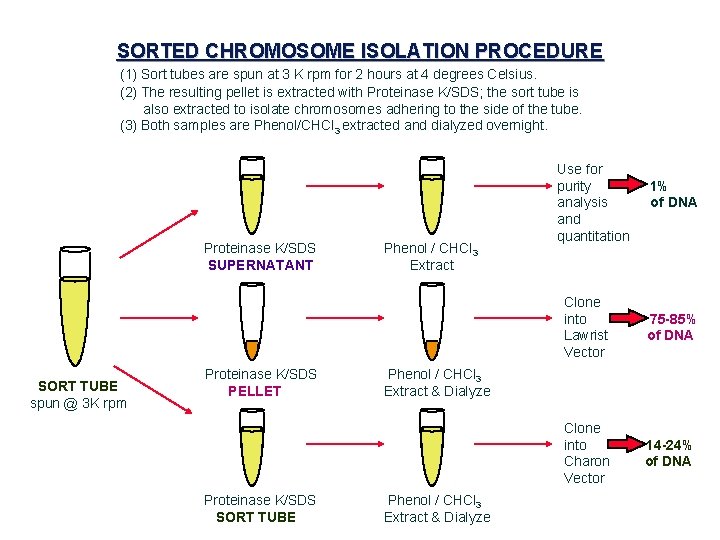
SORTED CHROMOSOME ISOLATION PROCEDURE (1) Sort tubes are spun at 3 K rpm for 2 hours at 4 degrees Celsius. (2) The resulting pellet is extracted with Proteinase K/SDS; the sort tube is also extracted to isolate chromosomes adhering to the side of the tube. (3) Both samples are Phenol/CHCl 3 extracted and dialyzed overnight. Proteinase K/SDS SUPERNATANT Phenol / CHCl 3 Extract Use for purity analysis and quantitation 1% of DNA Clone into ==> 75 -85% Lawrist of DNA Vector SORT TUBE spun @ 3 K rpm Proteinase K/SDS PELLET Phenol / CHCl 3 Extract & Dialyze Clone into ==> 14 -24% Charon of DNA Vector Proteinase K/SDS SORT TUBE Phenol / CHCl 3 Extract & Dialyze

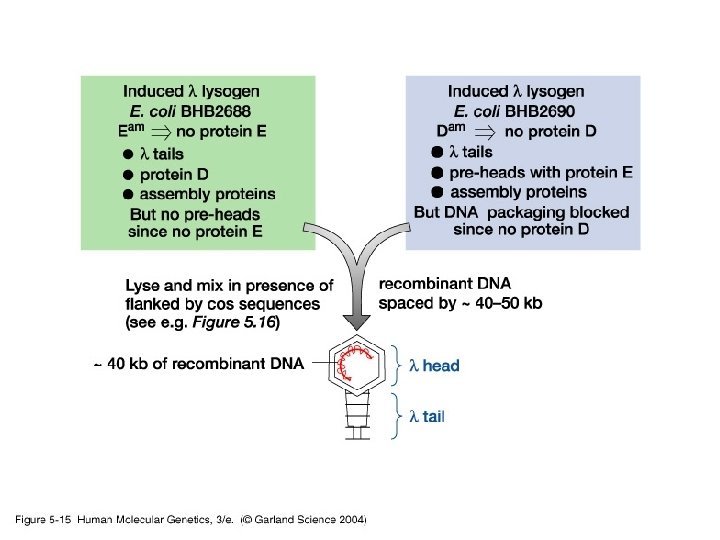
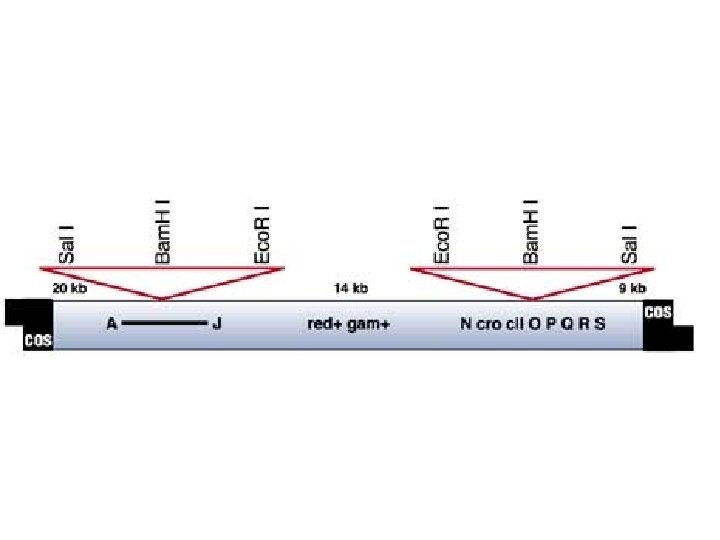
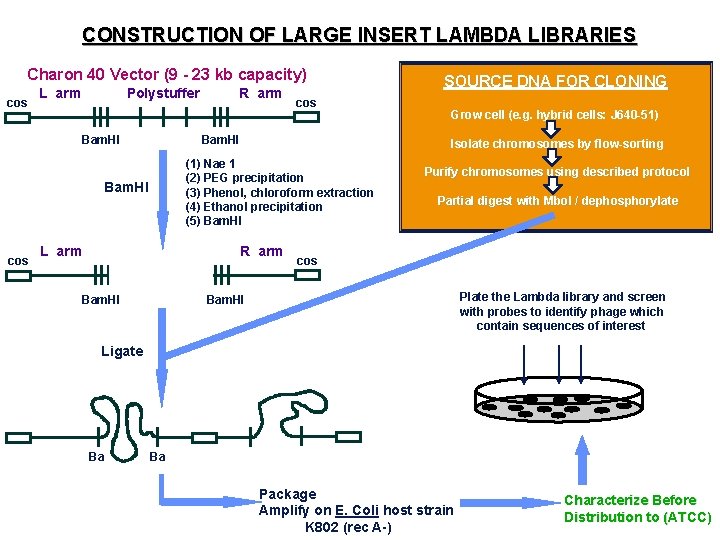
CONSTRUCTION OF LARGE INSERT LAMBDA LIBRARIES Charon 40 Vector (9 - 23 kb capacity) cos Polystuffer L arm Bam. HI SOURCE DNA FOR CLONING Grow cell (e. g. hybrid cells: J 640 -51) Isolate chromosomes by flow-sorting (1) Nae 1 (2) PEG precipitation (3) Phenol, chloroform extraction (4) Ethanol precipitation (5) Bam. HI Polystuffer L arm cos Bam. HI cos R arm Purify chromosomes using described protocol Partial digest with Mbo. I / dephosphorylate cos Plate the Lambda library and screen with probes to identify phage which contain sequences of interest Bam. HI Ligate Ba Ba Package Amplify on E. Coli host strain K 802 (rec A-) Characterize Before Distribution to (ATCC)

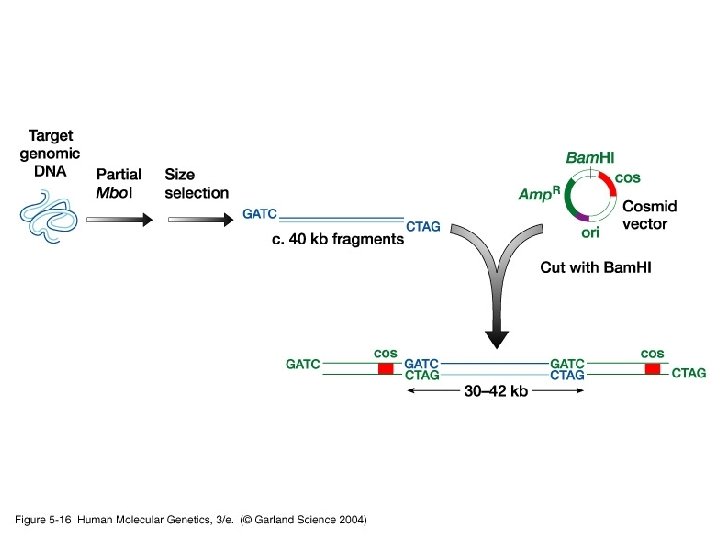
Cosmids and Fosmids • Cosmids – High-copy number replicon – Limited size based on packaging reactions – Chromosome-specific libraries • Fosmids – Low-copy replicon (F factor) – Limited size based on packaging reactions – Chromosome-specific libraries


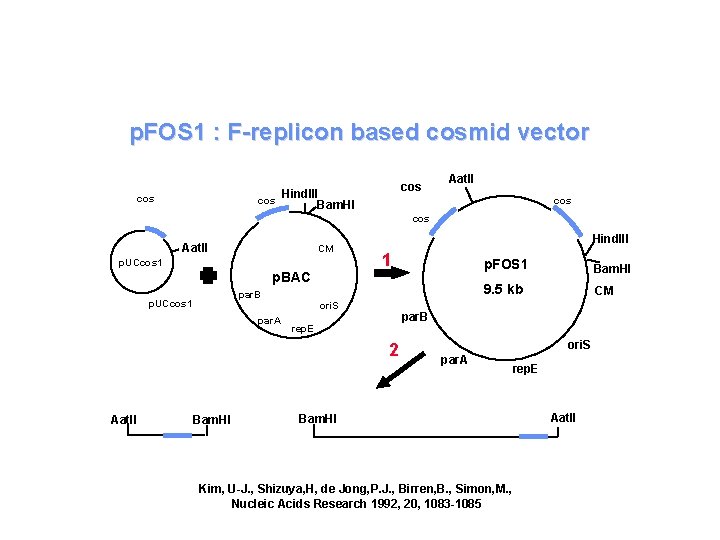
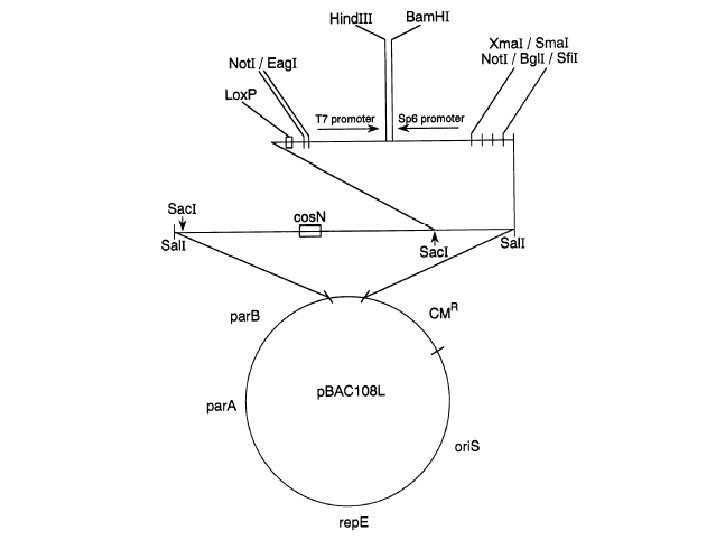
p. FOS 1 : F-replicon based cosmid vector cos cos Hind. III Bam. HI Aat. II cos Aat. II CM p. UCcos 1 Hind. III 1 p. BAC par. B p. UCcos 1 par. A ori. S Bam. HI 9. 5 kb CM par. B rep. E 2 Aat. II p. FOS 1 ori. S par. A rep. E Bam. HI Kim, U-J. , Shizuya, H, de Jong, P. J. , Birren, B. , Simon, M. , Nucleic Acids Research 1992, 20, 1083 -1085 Aat. II

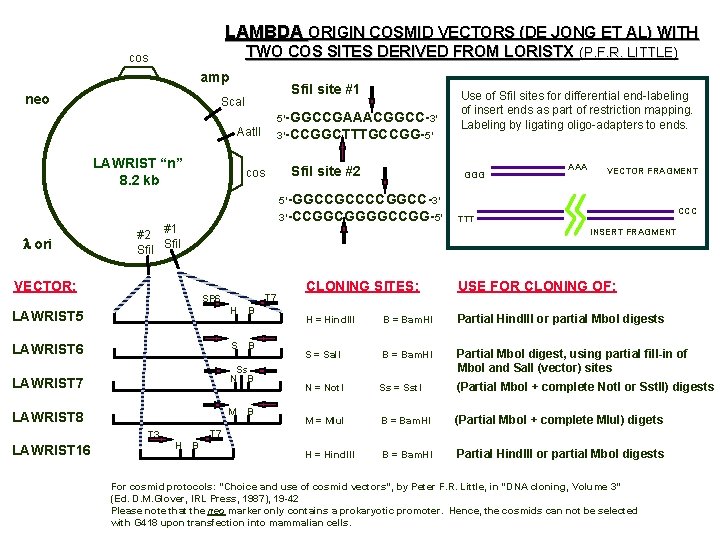
LAMBDA ORIGIN COSMID VECTORS (DE JONG ET AL) WITH TWO COS SITES DERIVED FROM LORISTX (P. F. R. LITTLE) cos amp neo Sfi. I site #1 Sca. I 5’-GGCCGAAACGGCC-3’ Aat. II LAWRIST “n” 8. 2 kb 3’-CCGGCTTTGCCGG-5’ Sfi. I site #2 cos Use of Sfi. I sites for differential end-labeling of insert ends as part of restriction mapping. Labeling by ligating oligo-adapters to ends. GGG AAA VECTOR FRAGMENT 5’-GGCCGCCCCGGCC-3’ l ori 3’-CCGGCGGGGCCGG-5’ #2 #1 Sfi. I VECTOR: INSERT FRAGMENT T 7 SP 6 B LAWRIST 5 H LAWRIST 6 S B Ss N B LAWRIST 7 M LAWRIST 8 B CLONING SITES: USE FOR CLONING OF: H = Hind. III B = Bam. HI Partial Hind. III or partial Mbo. I digests S = Sal. I B = Bam. HI Partial Mbo. I digest, using partial fill-in of Mbo. I and Sal. I (vector) sites N = Not. I Ss = Sst. I (Partial Mbo. I + complete Not. I or Sst. II) digests M = Mlul B = Bam. HI (Partial Mbo. I + complete Mlul) digets H = Hind. III B = Bam. HI Partial Hind. III or partial Mbo. I digests T 7 T 3 LAWRIST 16 CCC TTT H B For cosmid protocols: “Choice and use of cosmid vectors”, by Peter F. R. Little, in “DNA cloning, Volume 3” (Ed. D. M. Glover, IRL Press, 1987), 19 -42 Please note that the neo marker only contains a prokaryotic promoter. Hence, the cosmids can not be selected with G 418 upon transfection into mammalian cells.

SORTED CHROMOSOME ISOLATION PROCEDURE (1) Sort tubes are spun at 3 K rpm for 2 hours at 4 degrees Celsius. (2) The resulting pellet is extracted with Proteinase K/SDS; the sort tube is also extracted to isolate chromosomes adhering to the side of the tube. (3) Both samples are Phenol/CHCl 3 extracted and dialyzed overnight. Proteinase K/SDS SUPERNATANT Phenol / CHCl 3 Extract Use for purity analysis and quantitation 1% of DNA Clone into ==> 75 -85% Lawrist of DNA Vector SORT TUBE spun @ 3 K rpm Proteinase K/SDS PELLET Phenol / CHCl 3 Extract & Dialyze Clone into ==> 14 -24% Charon of DNA Vector Proteinase K/SDS SORT TUBE Phenol / CHCl 3 Extract & Dialyze

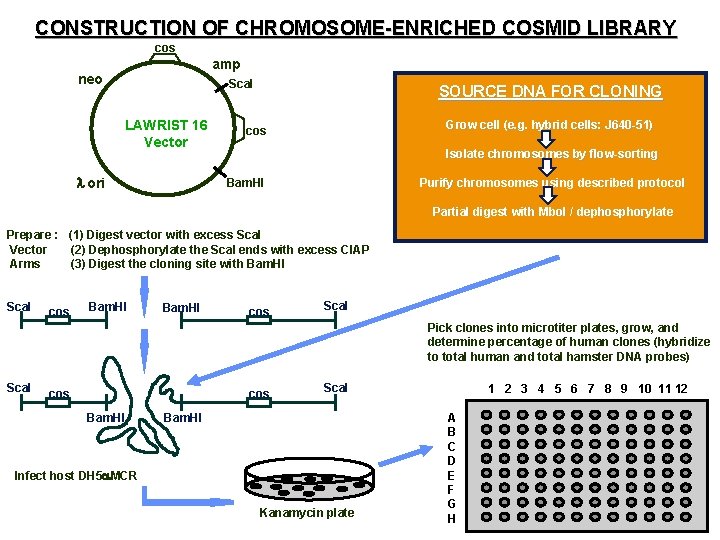
CONSTRUCTION OF CHROMOSOME-ENRICHED COSMID LIBRARY cos amp cos neo Sca. I LAWRIST 16 Vector l ori SOURCE DNA FOR CLONING Grow cell (e. g. hybrid cells: J 640 -51) cos Isolate chromosomes by flow-sorting Purify chromosomes using described protocol Bam. HI Partial digest with Mbo. I / dephosphorylate Prepare : (1) Digest vector with excess Sca. I Vector (2) Dephosphorylate the Sca. I ends with excess CIAP Arms (3) Digest the cloning site with Bam. HI Sca. I cos Bam. HI cos Sca. I Pick clones into microtiter plates, grow, and determine percentage of human clones (hybridize to total human and total hamster DNA probes) Sca. I cos Bam. HI Sca. I Bam. HI Infect host DH 5 a. MCR Kanamycin plate 1 2 3 4 5 6 7 8 9 10 11 12 A B C D E F G H

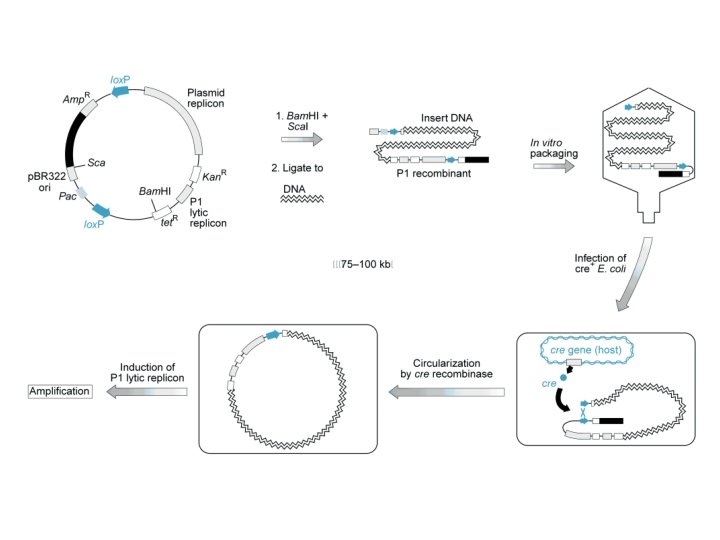
Bacteriophage P 1 • Insertion Vectors – Large Inserts – Limited to “HEADFULL” – Packaging reaction followed by plasmid propagation

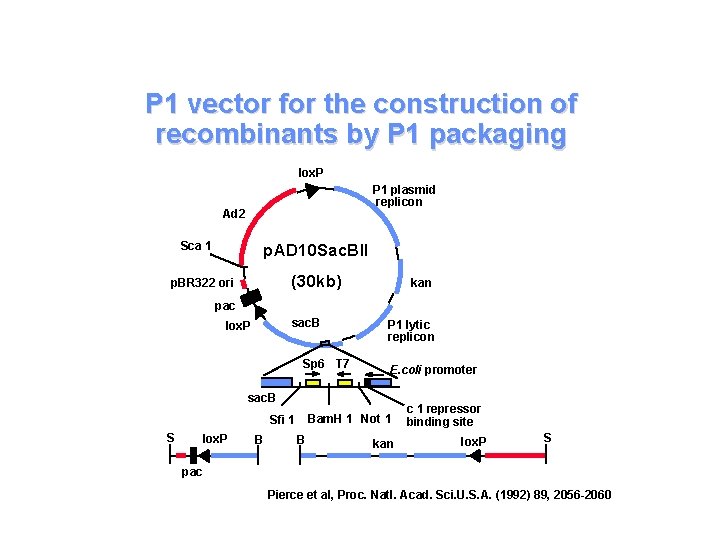
P 1 vector for the construction of recombinants by P 1 packaging lox. P P 1 plasmid replicon Ad 2 Sca 1 p. AD 10 Sac. BII (30 kb) p. BR 322 ori kan pac sac. B lox. P Sp 6 T 7 P 1 lytic replicon E. coli promoter sac. B Bam. H 1 Not 1 Sfi 1 S lox. P B B kan c 1 repressor binding site lox. P S pac Pierce et al, Proc. Natl. Acad. Sci. U. S. A. (1992) 89, 2056 -2060


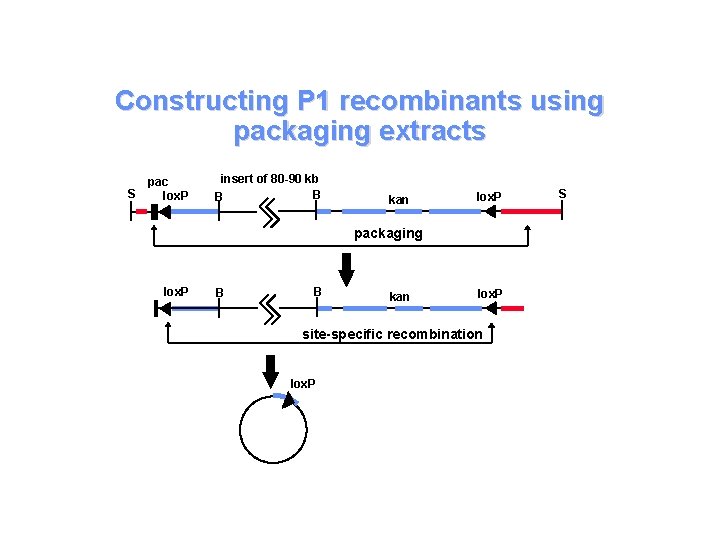
Constructing P 1 recombinants using packaging extracts S pac lox. P insert of 80 -90 kb B B kan lox. P packaging lox. P B B kan lox. P site-specific recombination lox. P S

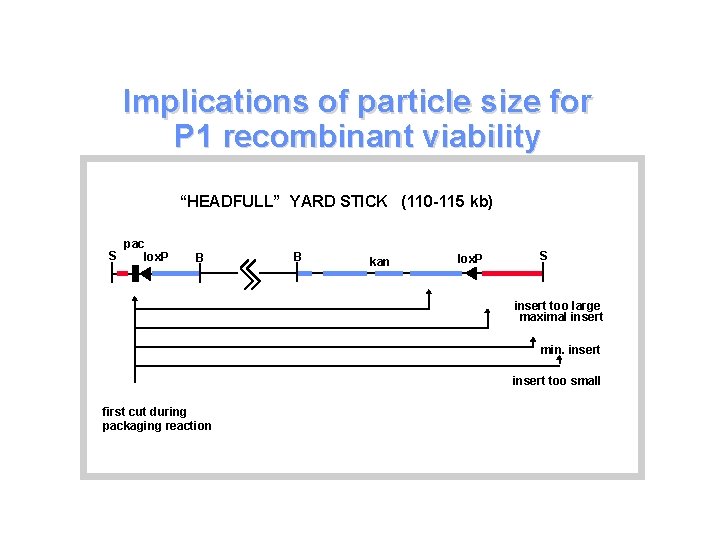
Implications of particle size for P 1 recombinant viability “HEADFULL” YARD STICK (110 -115 kb) S pac lox. P B B kan lox. P S insert too large maximal insert min. insert too small first cut during packaging reaction

P 1 Derived Artificial Chromosomes (PACs) • Electroporation Based System – – – Large insert size Low copy number origin for propagation High copy origin for DNA production Negative selection against non-recombinants Very stable inserts

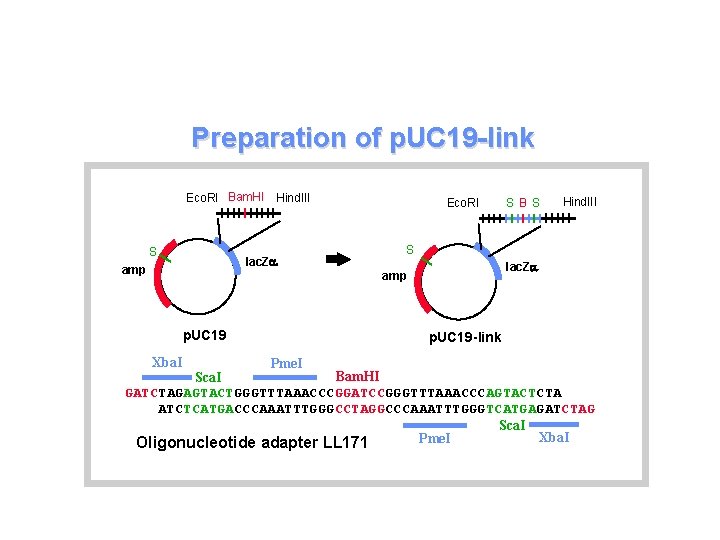
Preparation of p. UC 19 -link Eco. RI Bam. HI Hind. III S lac. Za amp Eco. RI Xba. I Hind. III S lac. Za amp p. UC 19 S BS p. UC 19 -link Pme. I Bam. HI Sca. I GATCTAGAGTACTGGGTTTAAACCCGGATCCGGGTTTAAACCCAGTACTCTA ATCTCATGACCCAAATTTGGGCCTAGGCCCAAATTTGGGTCATGAGATCTAG Sca. I Xba. I Pme. I Oligonucleotide adapter LL 171

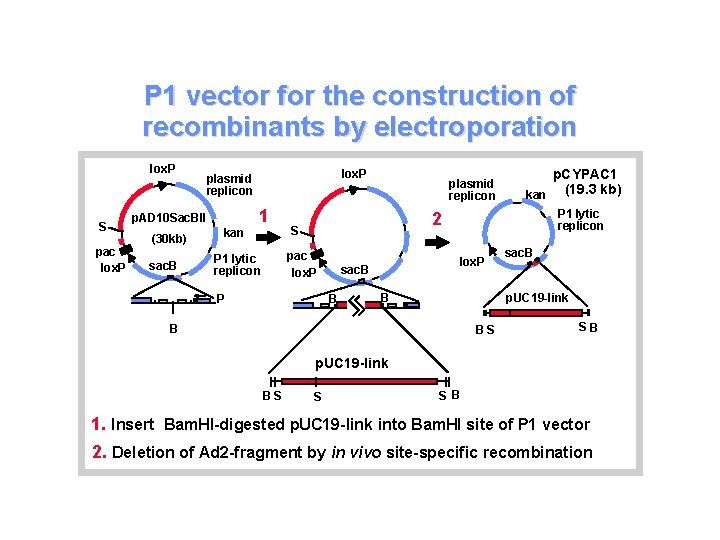
P 1 vector for the construction of recombinants by electroporation lox. P S pac lox. P p. AD 10 Sac. BII (30 kb) sac. B lox. P plasmid replicon kan 1 p. CYPAC 1 kan (19. 3 kb) P 1 lytic replicon 2 S pac lox. P P 1 lytic replicon plasmid replicon P lox. P sac. B B B sac. B p. UC 19 -link B BS SB p. UC 19 -link BS S S B 1. Insert Bam. HI-digested p. UC 19 -link into Bam. HI site of P 1 vector 2. Deletion of Ad 2 -fragment by in vivo site-specific recombination

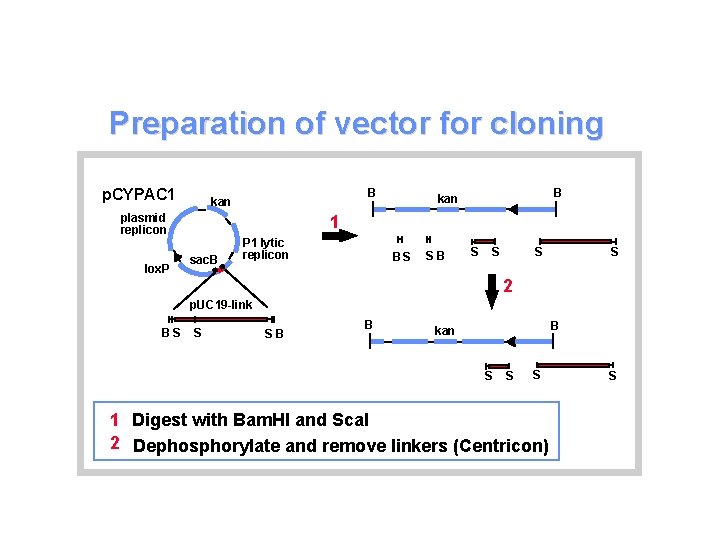
Preparation of vector for cloning p. CYPAC 1 plasmid replicon lox. P B kan 1 sac. B P 1 lytic replicon BS SB S S 2 p. UC 19 -link BS S SB B B kan S S S 1 Digest with Bam. HI and Sca. I 2 Dephosphorylate and remove linkers (Centricon) S

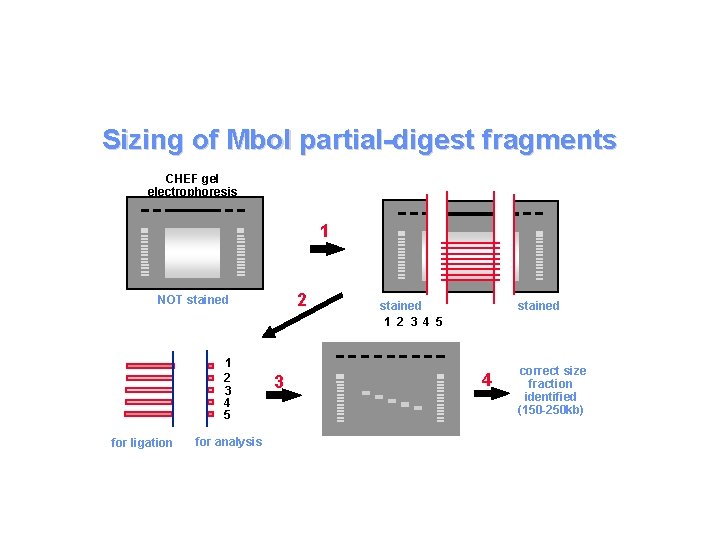
Sizing of Mbo. I partial-digest fragments CHEF gel electrophoresis 1 2 NOT stained 1 2 3 4 5 for ligation for analysis 3 stained 1 2 3 4 5 4 correct size fraction identified (150 -250 kb)

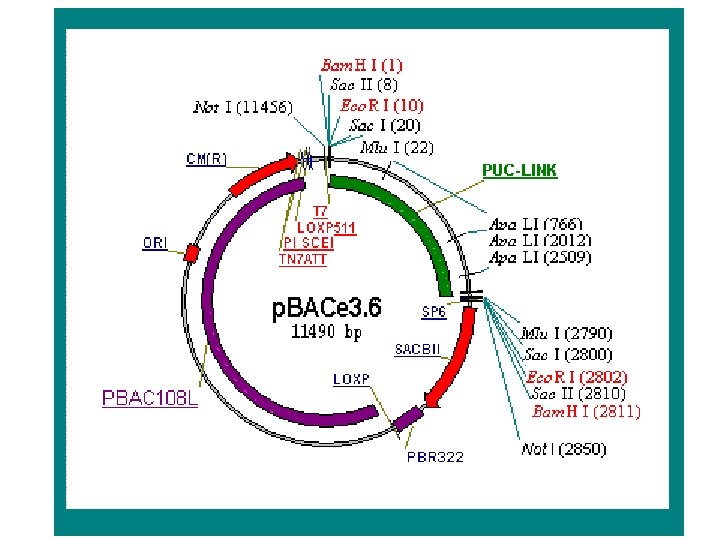
Bacterial Artificial Chromosomes (BACs) • Electroporation Based System – Large insert size – Low copy number origin for propagation – Very stable inserts



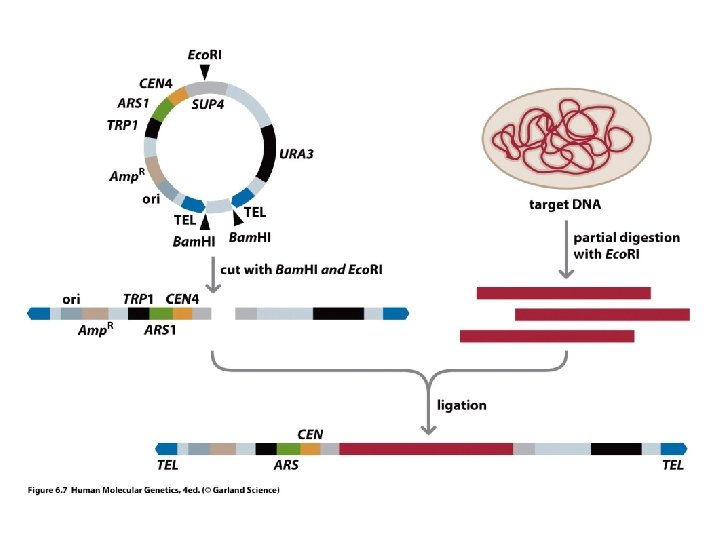
Yeast Artificial Chromosomes • Large Inserts • Not Stable • Chimeric inserts


Problems with YACs! 1) Approximately 40 -60% of the YACs from most libraries are chimeric. 2) Approximately 40 % of the YACs from most libraries are deleted. 3) Low transformation efficiencies. 4) YACs are very difficult to manipulate.

Physical Mapping Systems Yeast Artificial Chromosomes (YACs) 200 -2000 kb Bacteriophage P 1 90 kb Cosmids 40 kb Bacteriophage l 9 -23 kb

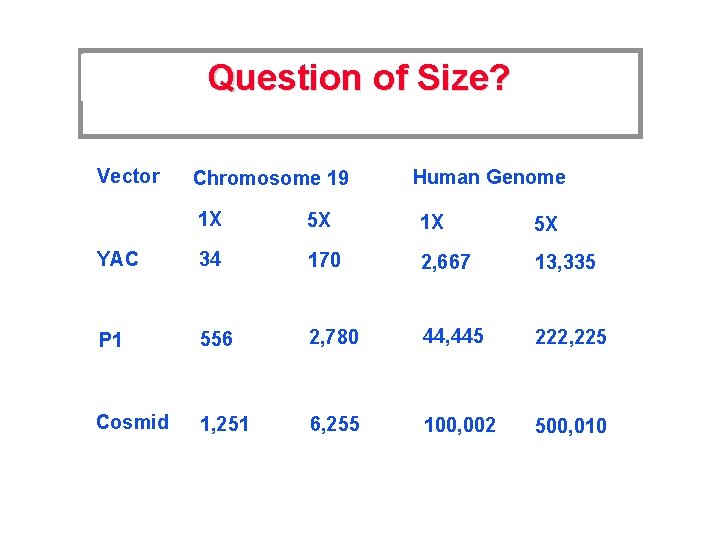
Question of Size? Vector Chromosome 19 Human Genome 1 X 5 X YAC 34 170 2, 667 13, 335 P 1 556 2, 780 44, 445 222, 225 Cosmid 1, 251 6, 255 100, 002 500, 010

Storage Space? Vector Chromosome 19 Human Genome 1 X 5 X YAC 1 2 28 139 P 1 6 29 463 2, 315 Cosmid 14 66 1, 042 5, 209

Advantages of BACs & PACs 1) Bacterial based systems that are easy to manipulate. 2) Libraries are generated using bacterial hosts with well defined properties. 3) Transformation efficiency is higher than that obtained for YACs. 4) BACs and PACs are non-chimeric, very stable and do not delete sequences.

Comparison of BACs & PACs 1) Both allow replication of clones at one copy/cell. 2) Both systems replicate clones faithfully across 60 -100 generations. 3) PACs also have a negative selection against non-recombinants. 4) PACs have an IPTG inducible high copy number origin of replication.

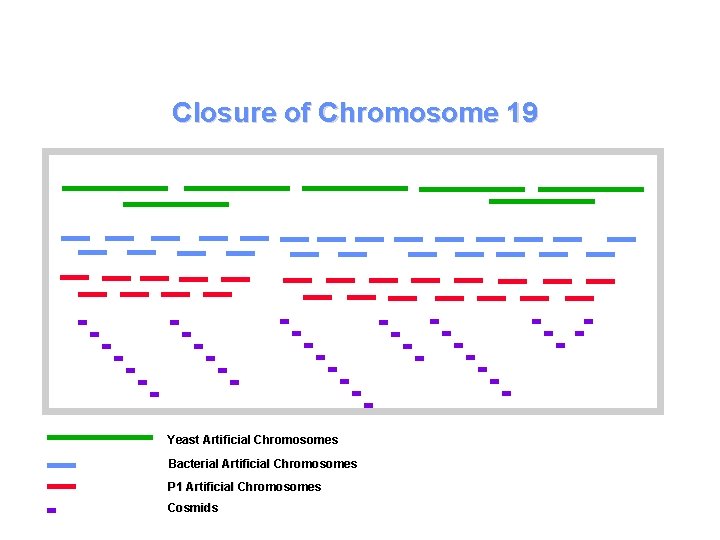
Closure of Chromosome 19 Type of Approach A) Bacterial Artificial Chromosomes (BACs). B) P 1 Artificial Chromosomes (PACs). C) Yeast Artificial Chromosomes (YACs). D) Cosmids. E)All of the above.

Closure of Chromosome 19 Yeast Artificial Chromosomes Bacterial Artificial Chromosomes P 1 Artificial Chromosomes Cosmids

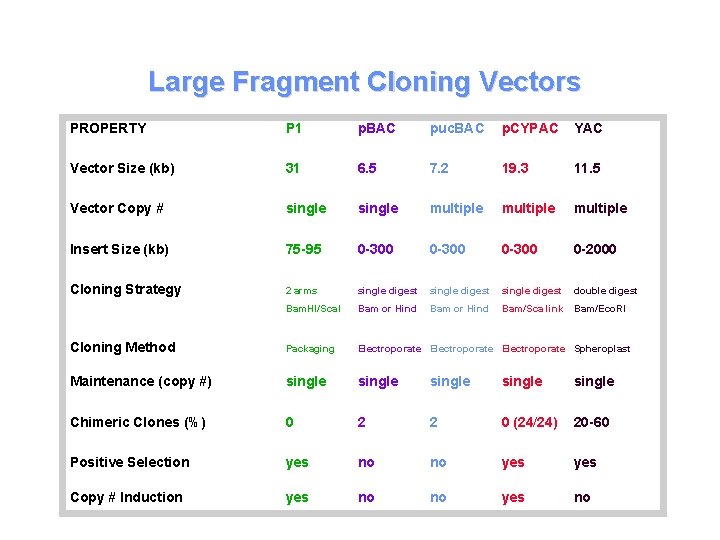
Large Fragment Cloning Vectors PROPERTY P 1 p. BAC puc. BAC p. CYPAC YAC Vector Size (kb) 31 6. 5 7. 2 19. 3 11. 5 Vector Copy # single multiple Insert Size (kb) 75 -95 0 -300 0 -2000 Cloning Strategy 2 arms single digest double digest Bam. HI/Sca. I Bam or Hind Bam/Sca link Bam/Eco. RI Cloning Method Packaging Electroporate Spheroplast Maintenance (copy #) single single Chimeric Clones (%) 0 2 2 0 (24/24) 20 -60 Positive Selection yes no no yes Copy # Induction yes no no yes no