Mapping Quantitative Trait Loci Two approaches 1 Biparental

Mapping Quantitative Trait Loci Two approaches 1. Biparental mapping populations • Linkage analysis in unselected progeny of a cross 2. Genome wide association studies (GWAS) • Search for associations between markers and phenotypes in a panel of individuals that represent the gene pool of interest
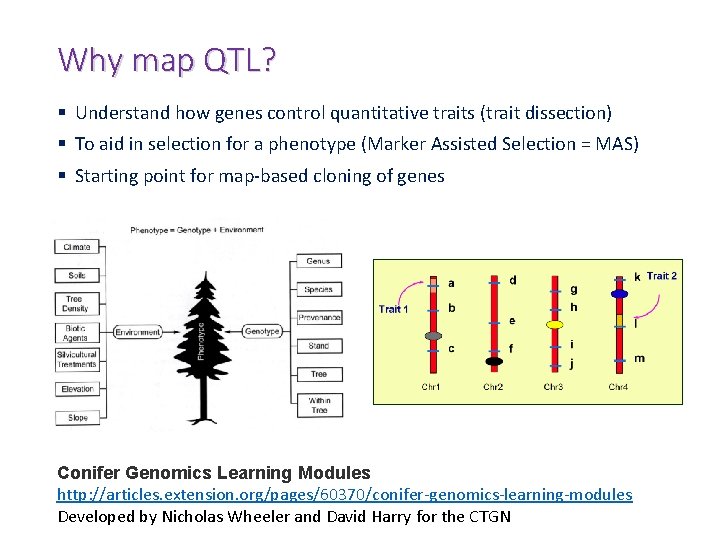
Why map QTL? § Understand how genes control quantitative traits (trait dissection) § To aid in selection for a phenotype (Marker Assisted Selection = MAS) § Starting point for map-based cloning of genes Conifer Genomics Learning Modules http: //articles. extension. org/pages/60370/conifer-genomics-learning-modules Developed by Nicholas Wheeler and David Harry for the CTGN
QTL mapping – linkage analysis § Select two parents that are genetically diverse for the trait(s) of interest Oregon Wolfe Barley § Cross parents and develop a population of segregating progeny § Measure phenotypes § Score molecular markers § Construct a linkage map for the markers § Identify QTL that are linked to the markers • Observe segregation of genetic markers • See if individuals with different marker genotypes have different phenotypes Dom Rec www. barleyworld. org

Advantages of biparental mapping populations § Derived from an F 1 population, so for unlinked loci, “linkage” equilibrium is obtained with one generation of recombination Ø Effects of QTL are not correlated (confounded) with the effects of unlinked genes in the population § Simplicity • No more than two alleles per locus • For polymorphic loci, p = q = 0. 5 (no minor alleles) Note: • Other types of pedigreed populations can also be used for linkage mapping • Results from multiple mapping populations may be pooled together • These approaches may provide additional benefits, but analysis may be more complex

Disadvantages of linkage analysis approach § There is often only one opportunity for recombination, so estimates of map distances are imprecise • Large populations are needed to obtain rare recombinants (in practice, existing populations are often too small) • Alternatively, can random-mate the F 2 for several generations before extracting lines, but this requires more time and expense § Must develop and maintain mapping populations that are often used solely for experimental purposes • e. g. , resistant x susceptible (rather than resistant x resistant) • additional time and expense § Results are specific for each mapping population • only 2 alleles per locus are sampled

Common types of mapping populations § F 2 population • Individual plants • F 2: 3 or F 2: 4 families – families of F 3 or F 4 plants that trace back to an F 2 individual § Backcross population § Recombinant inbred lines (RILs)** • Inbred lines obtained from an F 2 population through single seed descent (without selection) F 6 or later generation § Doubled haploids (DH)** • Derived from male or female gametes of an F 1 plant **Populations of fixed haplotypes can be efficiently genotyped and permanently maintained
Linkage mapping - steps 1. Genotype the mapping population and prepare data 2. Calculate recombination fractions (RFs) • Maximum likelihood estimates of pairwise RFs 3. Group loci into linkage groups • Based on maximum allowable RF threshold • Apply LOD threshold (e. g. , LOD > 3 indicates linkage) 4. Locus ordering • Begin with smallest RFs, and add others to minimize total size • Highest multipoint LOD among possible orders • Computer intensive 5. Multilocus estimation of map distances • Apply appropriate mapping function (Haldane, Kosambi)
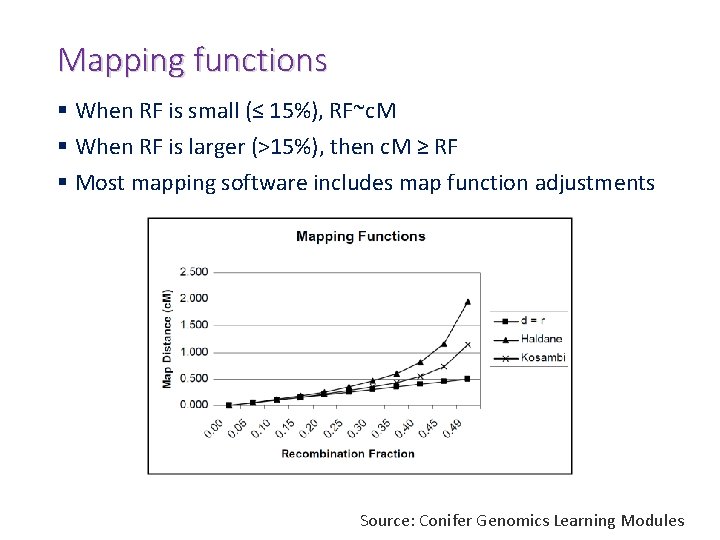
Mapping functions § When RF is small (≤ 15%), RF~c. M § When RF is larger (>15%), then c. M ≥ RF § Most mapping software includes map function adjustments Source: Conifer Genomics Learning Modules

Software § >100 genetic analyses software packages (linkage analysis and QTL mapping) § Distinguishing features • • Computer platform Types of populations that can be handled Mapping methods Price Common examples for linkage maps • Mapmaker/EXP • Join. Map Common examples for QTL analysis • QTL Cartographer • R/qtl

Linkage map 1 H 2 H Oregon Wolfe Barley • 175 DH lines • 1328 SNP markers • Same map order with ─ 82 maternal DH ─ 93 paternal DH Cistué et al. 2011. Theoretical and Applied Genetics 122: 1399– 1410. 3 H 4 H 5 H 6 H 7 H

QTL detection methods § Single-marker analysis (SMA) • Consider one marker at a time • Simple statistical tests are used to compare phenotypes of marker classes (e. g. AA, Aa, aa) ─ Need to make adjustments for multiple tests • Does not require a linkage map • Can’t tell how close the marker is to the QTL Ø Observed phenotypes for marker classes will depend on the extent of recombination as well as the genotypic effect of the QTL • Can’t evaluate interactions among QTLs
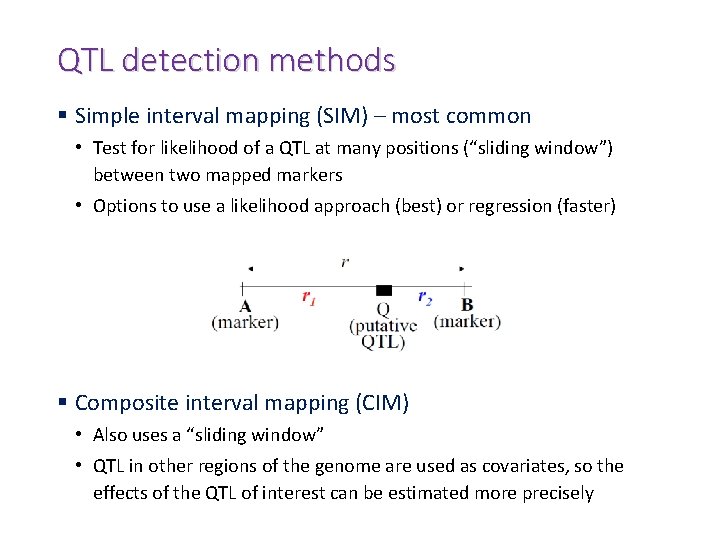
QTL detection methods § Simple interval mapping (SIM) – most common • Test for likelihood of a QTL at many positions (“sliding window”) between two mapped markers • Options to use a likelihood approach (best) or regression (faster) § Composite interval mapping (CIM) • Also uses a “sliding window” • QTL in other regions of the genome are used as covariates, so the effects of the QTL of interest can be estimated more precisely

Cercospora zeae-maydis Y 32 Q 11 1 3 5 7 9 § 161 F 2: 3 families § 2 replicates, 2 locations § 183 polymorphic SSR markers § Constructed linkage map § 4 QTL identified using CIM § 15 F 2 recombinants within a major QTL were used for fine mapping
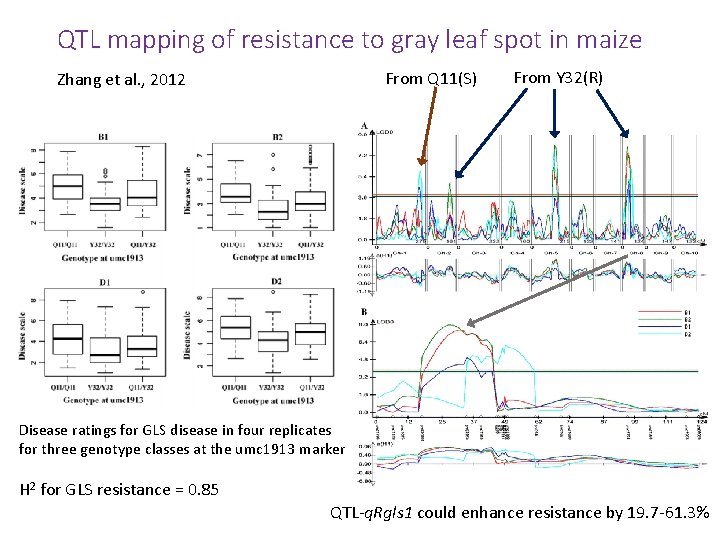
QTL mapping of resistance to gray leaf spot in maize From Q 11(S) Zhang et al. , 2012 From Y 32(R) Disease ratings for GLS disease in four replicates for three genotype classes at the umc 1913 marker H 2 for GLS resistance = 0. 85 QTL-q. Rgls 1 could enhance resistance by 19. 7 -61. 3%
- Slides: 14