Mapping Quantitative Trait Loci QTLs PBG 430 Mapping

Mapping Quantitative Trait Loci (QTLs) PBG 430

Mapping QTLs: two approaches § Goal: search for associations between markers (loci/genes) and phenotypes in a population § Two approaches: § Biparental mapping population: population studied is the progeny of/traces to a single designed mating (cross is designed to segregate for the trait of interest) § Genome wide association study (GWAS): population studied is a panel of individuals selected to represent the gene pool of interest § Both approaches employ an array of statistical and bioinformatics tools

Biparental QTL mapping – Steps 1. Select two parents that are genetically diverse for the trait(s) of interest 2. Cross parents and develop a progeny population § In self-pollinated crops, progeny is often designed to be homozygous plants (replicated experimental design) § Doubled haploids § Recombinant inbred lines § Large population size Oregon Wolfe Barley 3. Measure phenotypes 4. Score molecular markers 5. Construct a linkage map based on marker data 6. Identify QTLs that are linked to the markers Dominant Recessive

The OWB example – full dataset is publicly available Paper Full dataset
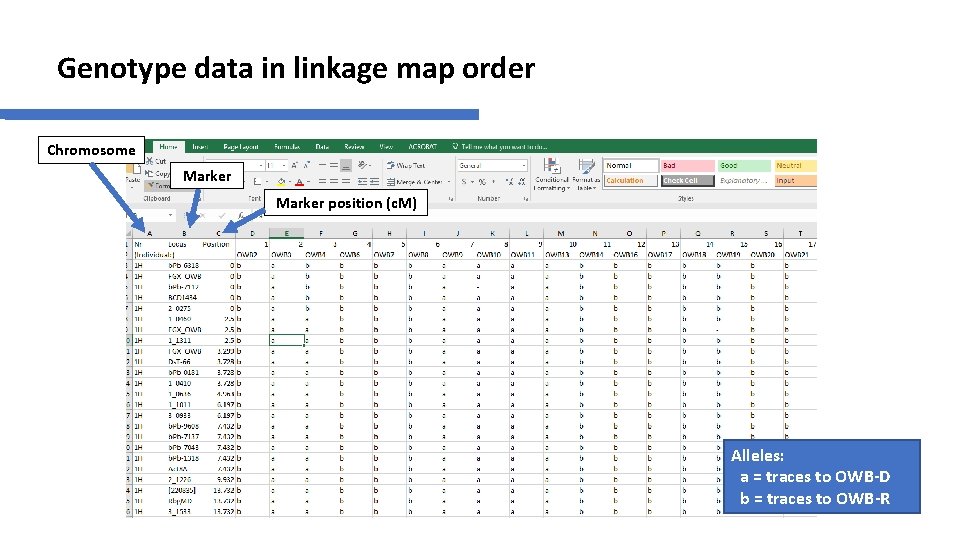
Genotype data in linkage map order Chromosome Marker position (c. M) Alleles: a = traces to OWB-D b = traces to OWB-R
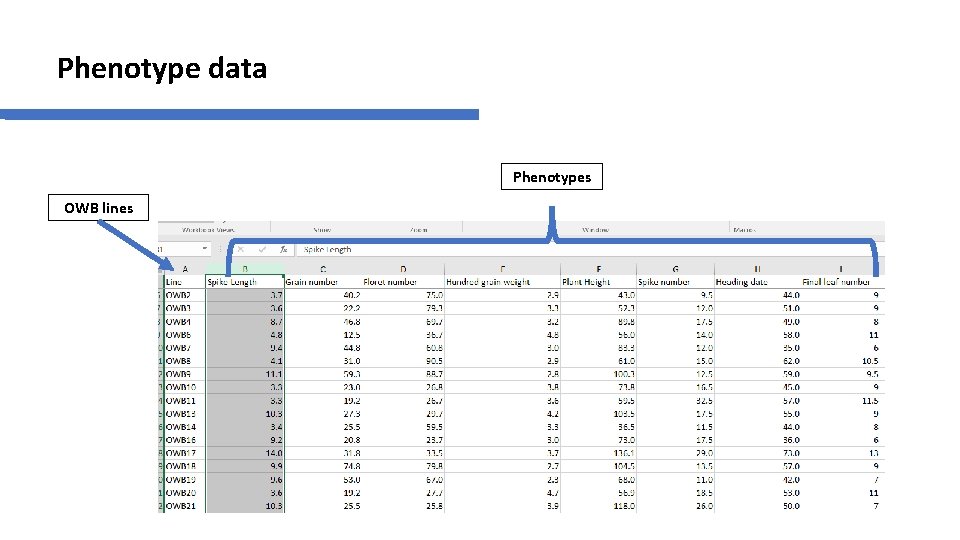
Phenotype data Phenotypes OWB lines

Biparental QTL mapping software: so many choices § Perform statistical tests to look for associations between phenotype and genotype § Example software: QTL Cartographer
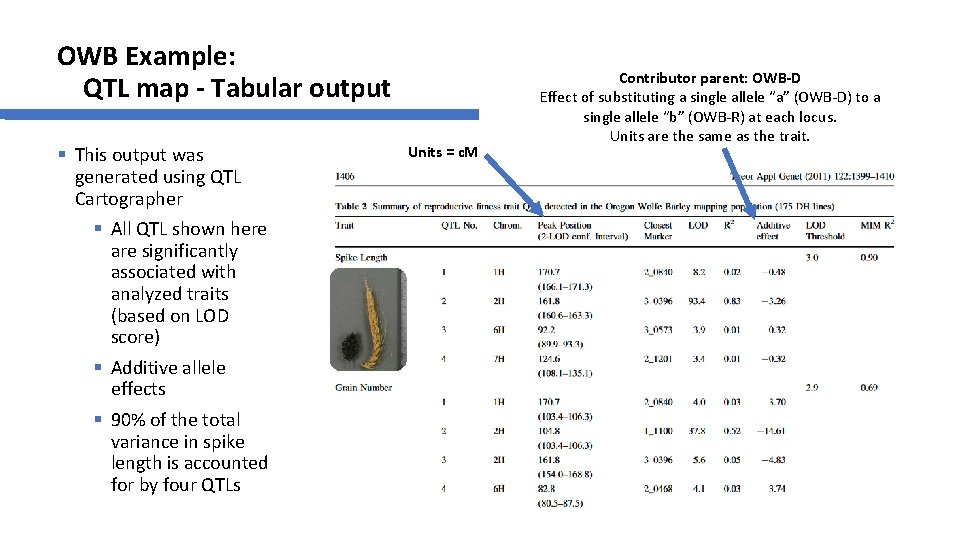
OWB Example: QTL map - Tabular output § This output was generated using QTL Cartographer § All QTL shown here are significantly associated with analyzed traits (based on LOD score) § Additive allele effects § 90% of the total variance in spike length is accounted for by four QTLs Units = c. M Contributor parent: OWB-D Effect of substituting a single allele “a” (OWB-D) to a single allele “b” (OWB-R) at each locus. Units are the same as the trait.
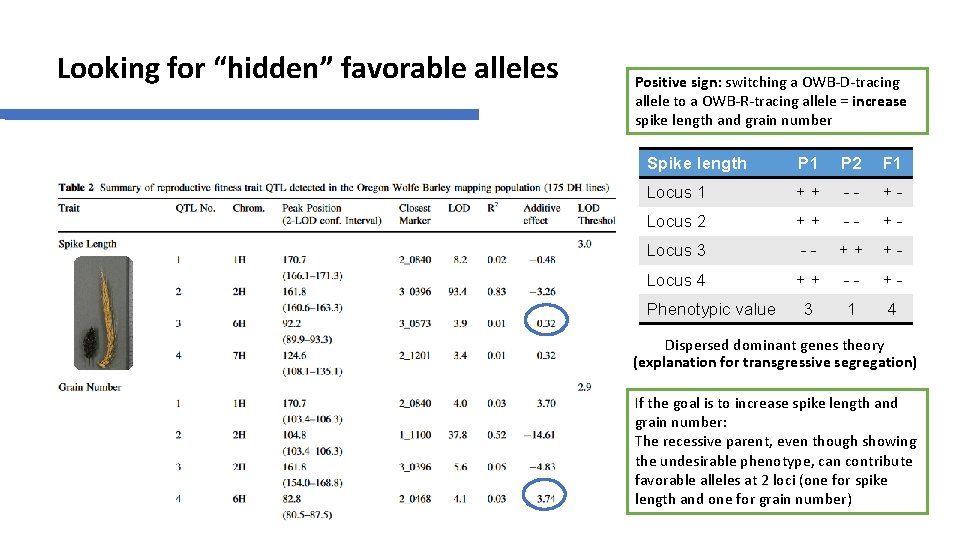
Looking for “hidden” favorable alleles Positive sign: switching a OWB-D-tracing allele to a OWB-R-tracing allele = increase spike length and grain number Spike length P 1 P 2 F 1 Locus 1 ++ -- +- Locus 2 ++ -- +- Locus 3 -- ++ +- Locus 4 ++ -- +- 3 1 4 Phenotypic value Dispersed dominant genes theory (explanation for transgressive segregation) If the goal is to increase spike length and grain number: The recessive parent, even though showing the undesirable phenotype, can contribute favorable alleles at 2 loci (one for spike length and one for grain number)
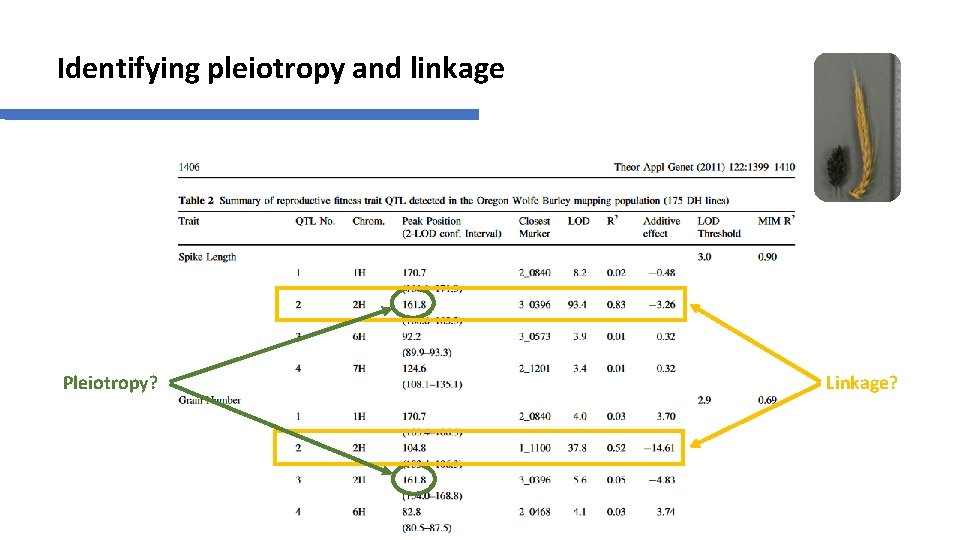
Identifying pleiotropy and linkage Pleiotropy? Linkage?
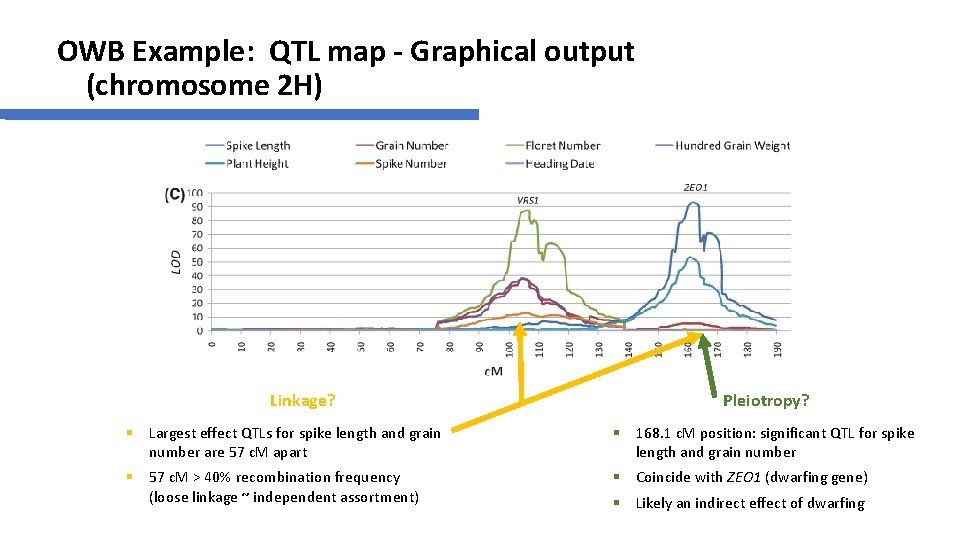
OWB Example: QTL map - Graphical output (chromosome 2 H) Linkage? Pleiotropy? § Largest effect QTLs for spike length and grain number are 57 c. M apart § 168. 1 c. M position: significant QTL for spike length and grain number § 57 c. M > 40% recombination frequency (loose linkage ~ independent assortment) § Coincide with ZEO 1 (dwarfing gene) § Likely an indirect effect of dwarfing
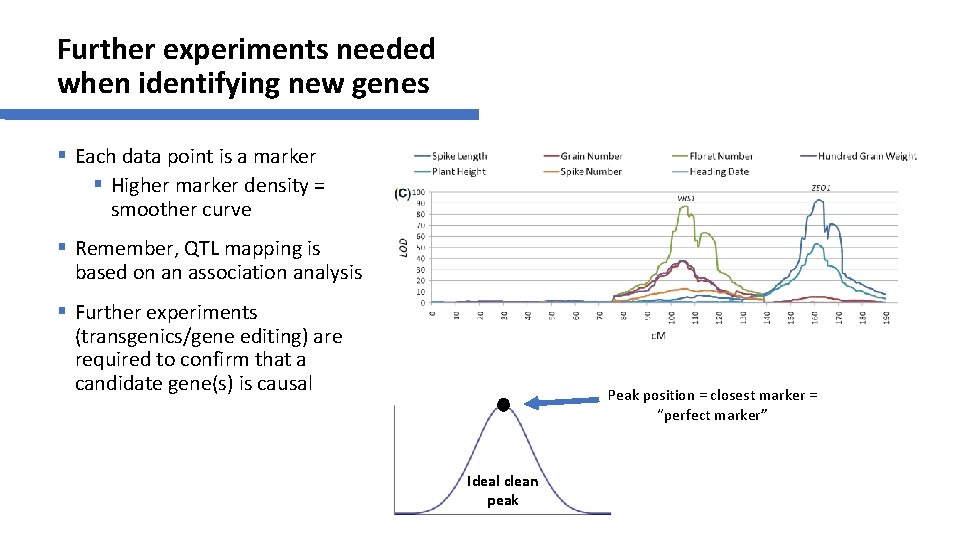
Further experiments needed when identifying new genes § Each data point is a marker § Higher marker density = smoother curve § Remember, QTL mapping is based on an association analysis § Further experiments (transgenics/gene editing) are required to confirm that a candidate gene(s) is causal Peak position = closest marker = “perfect marker” Ideal clean peak

Mapping Quantitative Trait Loci (QTLs) - Summary § Integrate phenotypic data and genotypic data § Reduce the complexity of quantitative traits to map coordinates (linkage or physical map) § Describe phenotype/genotype associations in terms of: § Statistical significance (LOD) § % of phenotypic variance explained (R 2) § Linked or perfect markers § Additive allele effect (QTL allele value)

Why map QTLs? § Understand how genes control quantitative traits (trait dissection) § Starting point for map-based cloning of genes (identifying new genes) § Identify genetic markers for indirect selection for expensive/challenging phenotypes (Marker Assisted Selection = MAS) § Determine if correlation of phenotypes is due to linkage or pleiotropy § Applicable to all types of plants

By now you should be able to… § Describe the goal of QTL mapping and differentiate between the two approaches that can be used to achieve it. § Describe the steps in a biparental QTL mapping. § Why are doubled haploids or recombinant inbred lines commonly used in selfpollinated species? § Interpret QTL map data in tabular form. § Remember that “unexpected” QTL allele values (i. e. additive effect) indicate the possibility of transgressive segregation and that the effect of a QTL is measured by both the LOD score and the R 2 § Interpret a QTL map in graphical form. § Why are further experiments required to confirm that a candidate gene is causing the trait? § Describe how the identification of markers through QTL mapping can be used for Marker Assisted Selection.
- Slides: 15