Mapping and Visualization of Biochemical Networks Stephen Michnick

Mapping and Visualization of Biochemical Networks Stephen Michnick Département de biochimie Université de Montréal

Mapping and Visualization of Biochemical Networks • Theme I: Design and Modularity in Biochemical Networks • Theme II: A general approach to mapping biochemical networks. Design organization and paradoxes in PKB signaling • Theme III: Design Modularity and Evolution of Networks • Theme IV: Towards genome-wide mapping of biochemical networks in living cells.
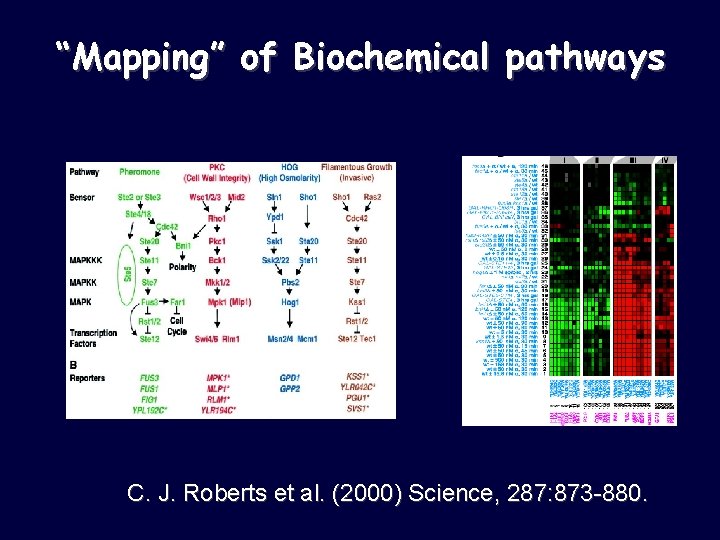
“Mapping” of Biochemical pathways C. J. Roberts et al. (2000) Science, 287: 873 -880.
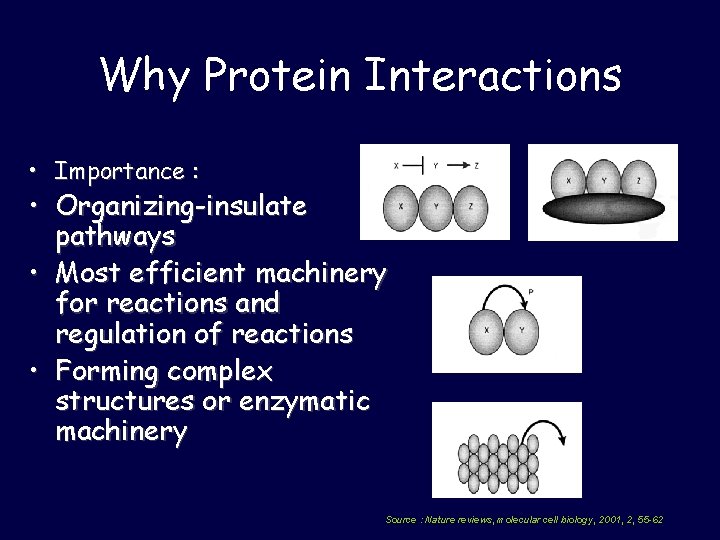
Why Protein Interactions • Importance : • Organizing-insulate pathways • Most efficient machinery for reactions and regulation of reactions • Forming complex structures or enzymatic machinery Source : Nature reviews, molecular cell biology, 2001, 2, 55 -62

Mapping and Visualization of Biochemical Networks • Theme I: Design and Modularity in Biochemical Networks • Theme II: A general approach to mapping biochemical networks. Design organization and paradoxes in PKB signaling • Theme III: Design Modularity and Evolution of Networks The Rosetta Hypothesis and organization of networks • Theme IV: Towards genome-wide mapping of biochemical networks in living cells. The MAP kinase pathways in S. cerevisiae.
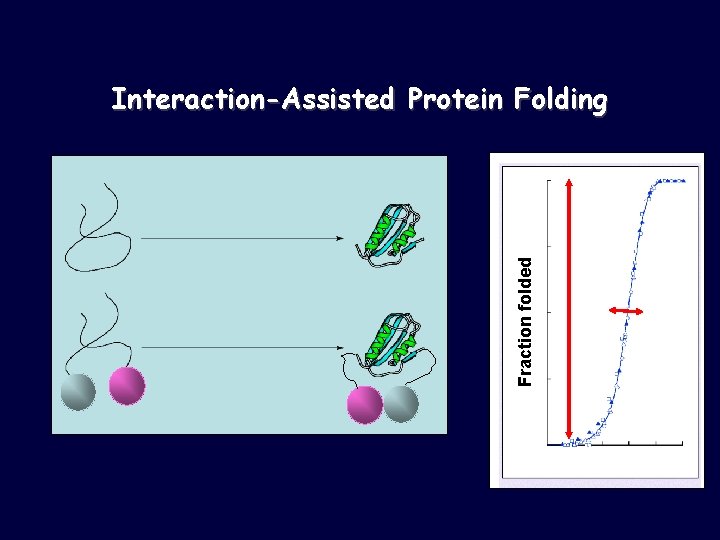
Fraction folded Interaction-Assisted Protein Folding

PCA (Protein fragment Complementation Assays) Screening A B Dynamics Quantitation Pharmacology Localization Trafficking Genes are transfected into cells. The cell makes proteins A and B which have reporter tags on either end (N- or C-terminal). A/B complex formation reconstitutes the reporter and generates a signal. Any gene, cell and reporter protein can be used. Localization Trafficking
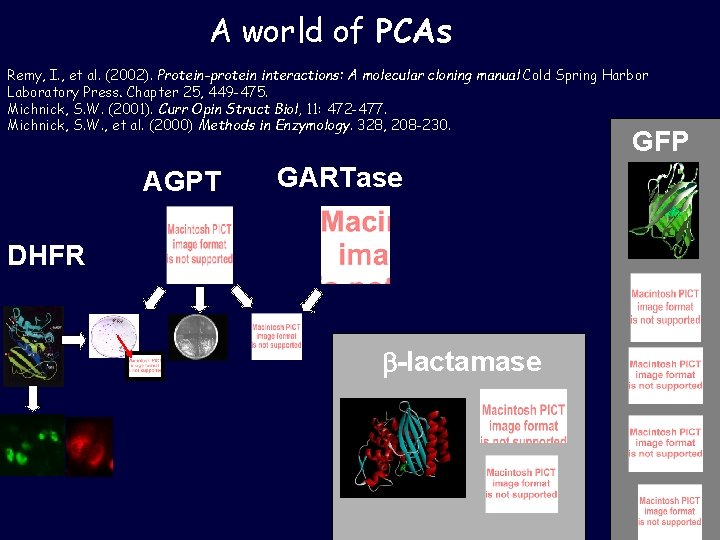
A world of PCAs Remy, I. , et al. (2002). Protein-protein interactions: A molecular cloning manual Cold Spring Harbor Laboratory Press. Chapter 25, 449 -475. Michnick, S. W. (2001). Curr Opin Struct Biol, 11: 472 -477. Michnick, S. W. , et al. (2000) Methods in Enzymology. 328, 208 -230. GFP AGPT GARTase DHFR b-lactamase

PCA in Network Mapping • Molecular interactions are detected directly. • Genes are expressed in a relevant cellular context, in which components of the underlying pathway exist. • Events induced by any pathway purturbation can be detected, linking specific interactions to specific pathways. • Subcellular locations of protein complexes can be determined unambiguously.

Dihydrofolate reductase (DHFR) PNAS Vol. 95, pp. 12141 -12146 - Fluorometirc assay based on binding of fluor-methotrexate to DHFR PNAS Vol. 96, pp. 5394 -5399, May 1999
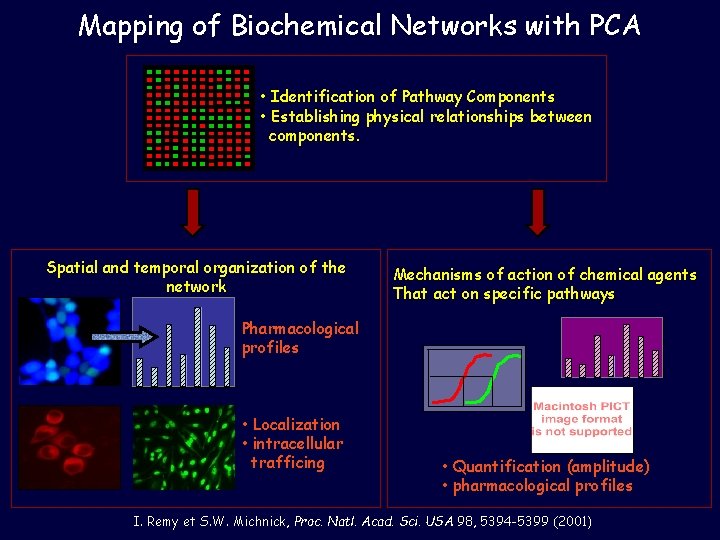
Mapping of Biochemical Networks with PCA • Identification of Pathway Components • Establishing physical relationships between components. Spatial and temporal organization of the network Mechanisms of action of chemical agents That act on specific pathways Pharmacological profiles • Localization • intracellular trafficing • Quantification (amplitude) • pharmacological profiles I. Remy et S. W. Michnick, Proc. Natl. Acad. Sci. USA 98, 5394 -5399 (2001)
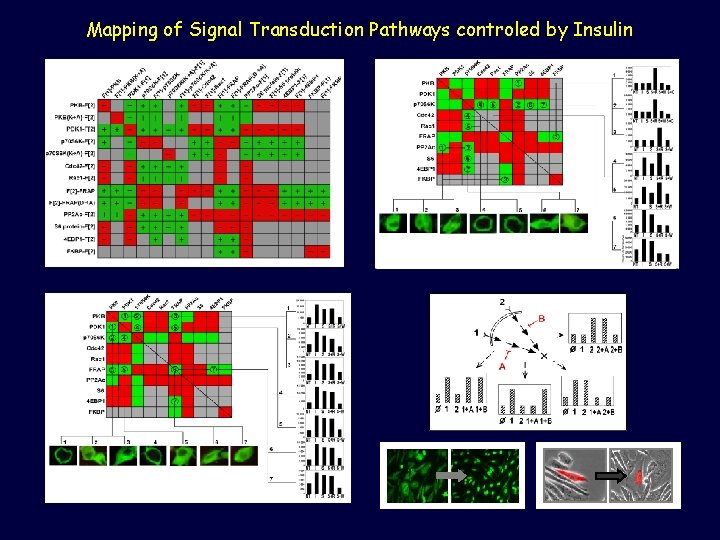
Mapping of Signal Transduction Pathways controled by Insulin
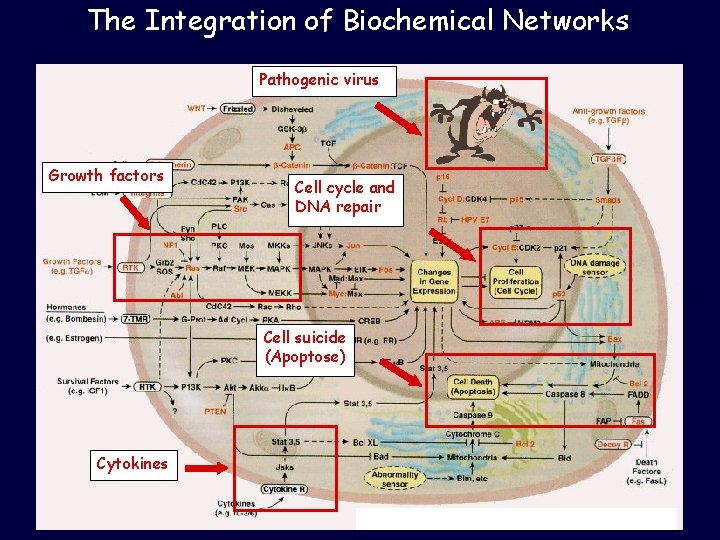
The Integration of Biochemical Networks Pathogenic virus Growth factors Cell cycle and DNA repair Cell suicide (Apoptose) Cytokines
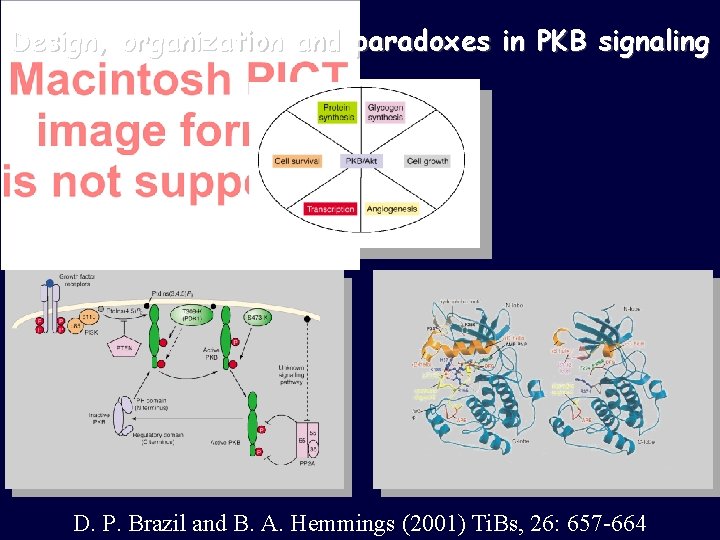
Design, organization and paradoxes in PKB signaling D. P. Brazil and B. A. Hemmings (2001) Ti. Bs, 26: 657 -664

Convergence of RTK and FRAP Pathways

Crosstalk between FRAP and PKB signaling
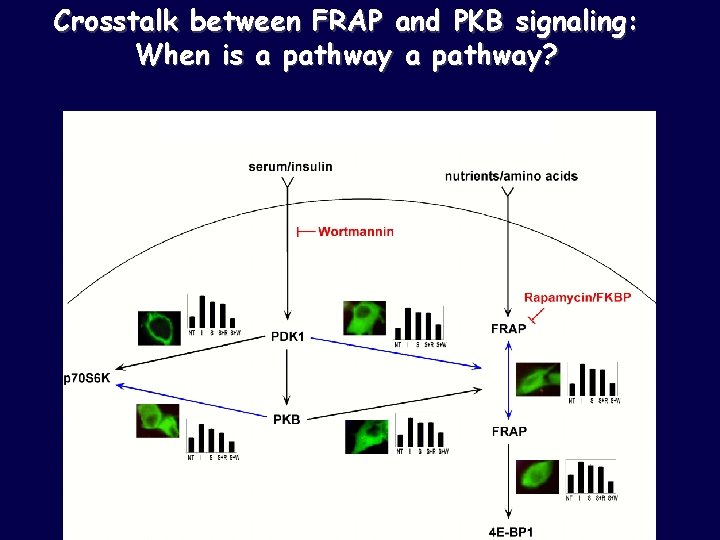
Crosstalk between FRAP and PKB signaling: When is a pathway?
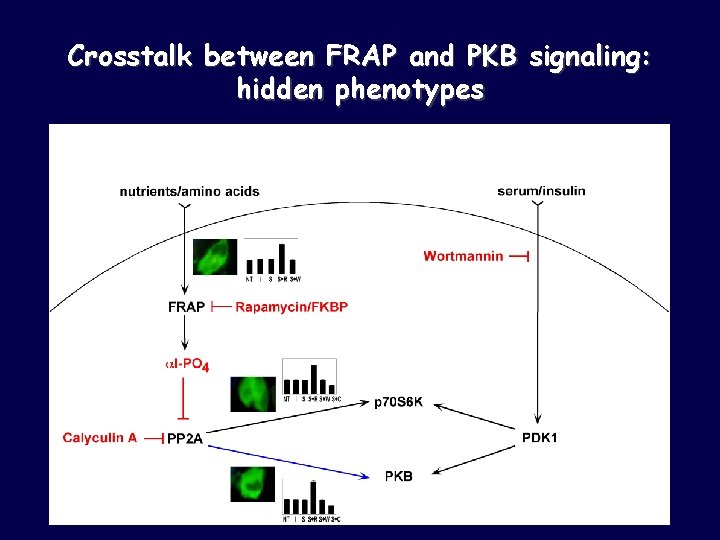
Crosstalk between FRAP and PKB signaling: hidden phenotypes
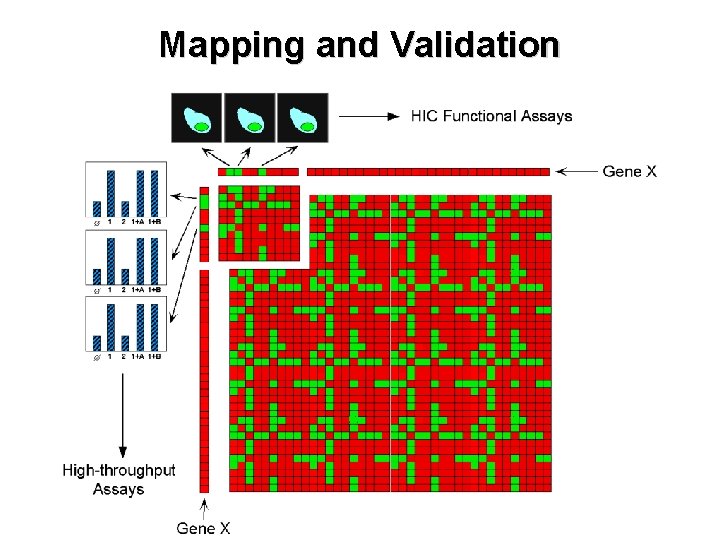
Mapping and Validation

PCA in Network Mapping • Molecular interactions are detected directly. • Genes are expressed in a relevant cellular context, in which components of the underlying pathway exist. • Events induced by any pathway purturbation can be detected, linking specific interactions to specific pathways. • Subcellular locations of protein complexes can be determined unambiguously.

Directional c. DNA library screening with the GFP PCA • Size-fractionated directional c. DNA library from human brain (107 independent clones) • Transiently co-transfected COS-1 cells

Systematic Screening of PKB (AKT 1 and AKT 2)

Mapping biochemical networks The key to resolving specificity and organization of biochemical networks is CELL-BASED ASSAYS. PCA strategy A general approach to mapping pathways that allows for both defining the organization of pathways. Can be used for pure description in an ontological sense (what, where and when) as well as more quantitative descriptions.

PCA in Network Mapping • Molecular interactions are detected directly. • Genes are expressed in a relevant cellular context, in which components of the underlying pathway exist. • Events induced by any pathway purturbation can be detected, linking specific interactions to specific pathways. • Subcellular locations of protein complexes can be determined unambiguously.

Mapping and Visualization of Biochemical Networks • Theme I: Design and Modularity in Biochemical Networks • Theme II: A general approach to mapping biochemical networks. The key to resolving specificity and organization of biochemical networks is CELL-BASED ASSAYS. PCA provides general approach to mapping pathways that allows for both defining the organization of pathways in terms of interactions and their dynamics. • Theme III: Design, Modularity and Evolution of Networks e. g. Rosetta Hypothesis: Predictions of interactions and functional inferences provide crucial and testable hypotheses. • Theme IV: Towards genome-wide mapping of biochemical networks in living cells. Mapping the interactions among all biochemical network components is the only way that we may ultimately know natures design. If there is one.

Thanks to… F. -X. Campbell-Valois André Galarneau Ingrid Remy Alexis Vallée-Belisle Dimitri Sans Helen Yu Jane Lamerdin Hugo Lavois Luciano Vidali Nathalie Bourassa Galia Ghaddar Martin Primeau Jean-François Turcotte Annie Montmarquette Geoffroy Denis Philippe Nissaire Po Hein Ear Stéphane Le Crom Kirill Tarrasov Ludovic Fuzellier Financing CIHR, NIH, The Burroughs-Wellcome Fund, HFSP Génome Québec/Canada, Odyssey Pharmaceuticals Inc.
- Slides: 26