Manual Annotation of Human Genome at Broad Institute

Manual Annotation of Human Genome at Broad Institute Chinnappa Kodira April 2004 GMOD 2004, Cambridge, MA

Goals n Accurate and comprehensive catalog of genes and gene products n Robust annotation system for annotation of all sequenced genomes

Annotation Strategy: Evidence-based Annotation CSMD 1 gene: Gene Size: 2065, 608 bases Transcript Length: 11, 297 bases Protein Length: 3565 aa No of Exons: 68 Average length of Exons : 166 bases Fgensh 20 Genscan 25 Blat_EST 179 m. RNA 3

Rule-based Annotation FL-m. RNA Species-specific ESTs Cross-species ESTs Protein homology Decreasing order of confidence level Ecores + Gene. Predictions
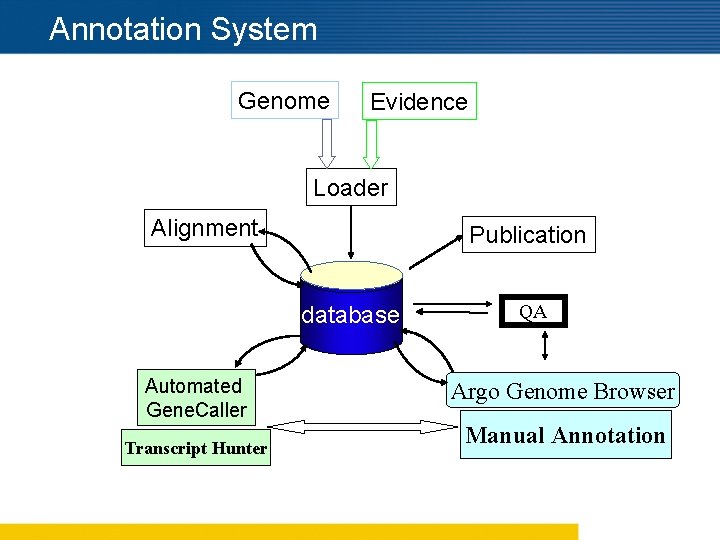
Annotation System Genome Evidence Loader Alignment Publication database Automated Gene. Caller Transcript Hunter QA Argo Genome Browser Manual Annotation

Critical Steps in our Annotation Process n Running Computes n Selection and Filtering Evidence n Intelligent Automated Gene Caller n Genome Browser and Editor n Annotation Rules n Trained Manual Annotators n Annotation QA Process
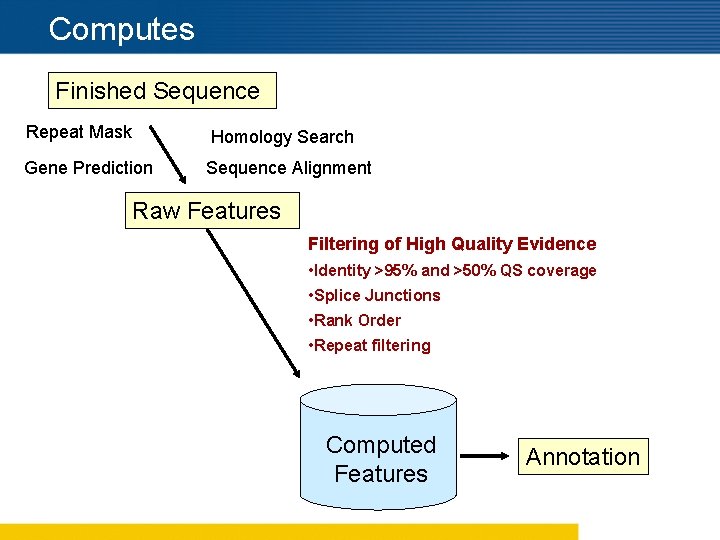
Computes Finished Sequence Repeat Mask Homology Search Gene Prediction Sequence Alignment Raw Features Filtering of High Quality Evidence • Identity >95% and >50% QS coverage • Splice Junctions • Rank Order • Repeat filtering Computed Features Annotation
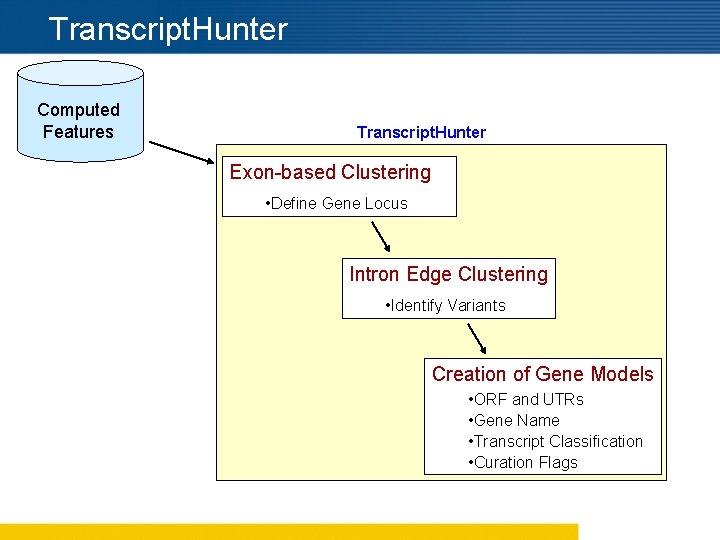
Transcript. Hunter Computed Features Transcript. Hunter Exon-based Clustering • Define Gene Locus Intron Edge Clustering • Identify Variants Creation of Gene Models • ORF and UTRs • Gene Name • Transcript Classification • Curation Flags

Screening of spliced ESTs contained within repeat elements Alu. Yb 8 Repeat Spliced ESTs
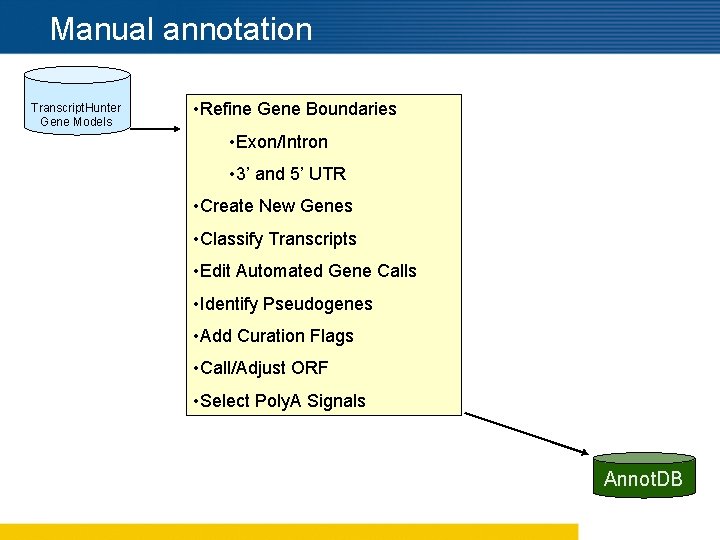
Manual annotation Transcript. Hunter Gene Models • Refine Gene Boundaries • Exon/Intron • 3’ and 5’ UTR • Create New Genes • Classify Transcripts • Edit Automated Gene Calls • Identify Pseudogenes • Add Curation Flags • Call/Adjust ORF • Select Poly. A Signals Annot. DB

Features of Argo n Attaching primary and supplemental evidence n Cluster feature display n Filtering and customizing evidence list n Display poly A signals and splice junctions n Alerting discrepancies before updating n Highlighting parent and child features n Real-time interactive analysis n ORF selection options n Tabular dump of selected features n Roll back and save work n Customization of feature display
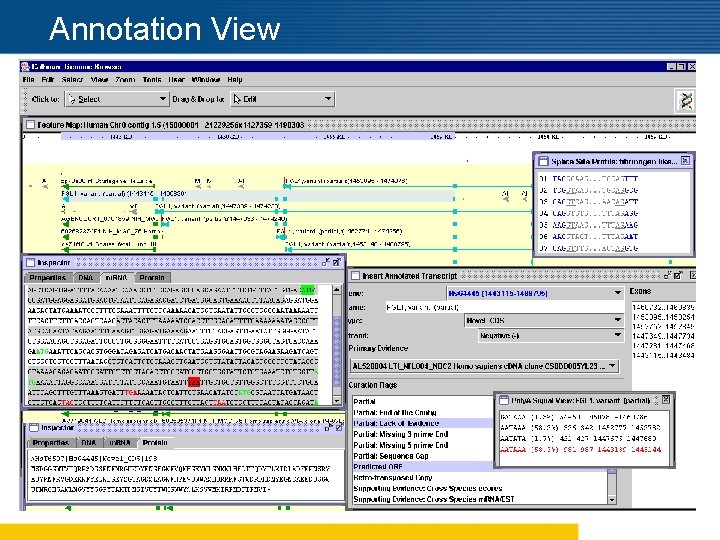
Annotation View

Confidence levels of our gene models n Classification of transcripts –Hawk standards n Known, Novel_CDS, Novel, Putative, Pseudogene n Association of primary and supplemental evidence with annotated feature n Rank order in selection of supporting evidence n Curation flags n Free text comments
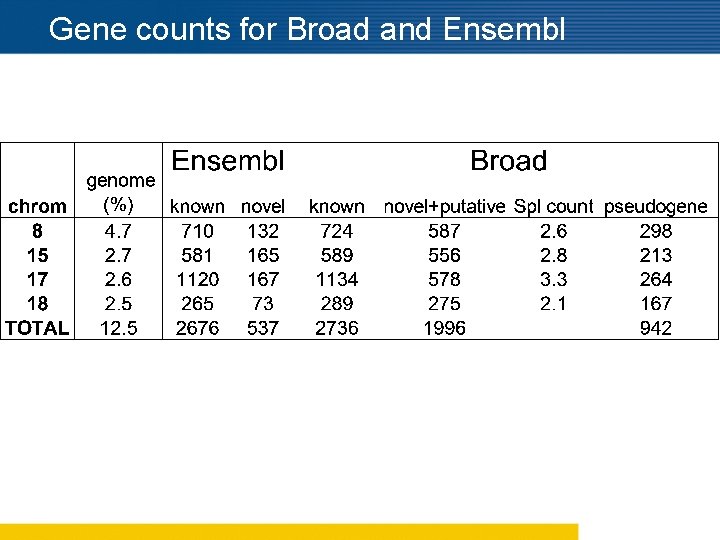
Gene counts for Broad and Ensembl
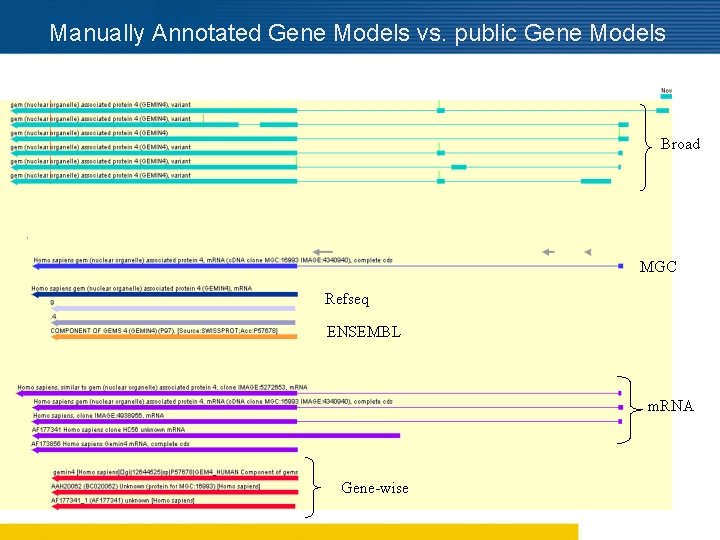
Manually Annotated Gene Models vs. public Gene Models Broad MGC Refseq ENSEMBL m. RNA Gene-wise
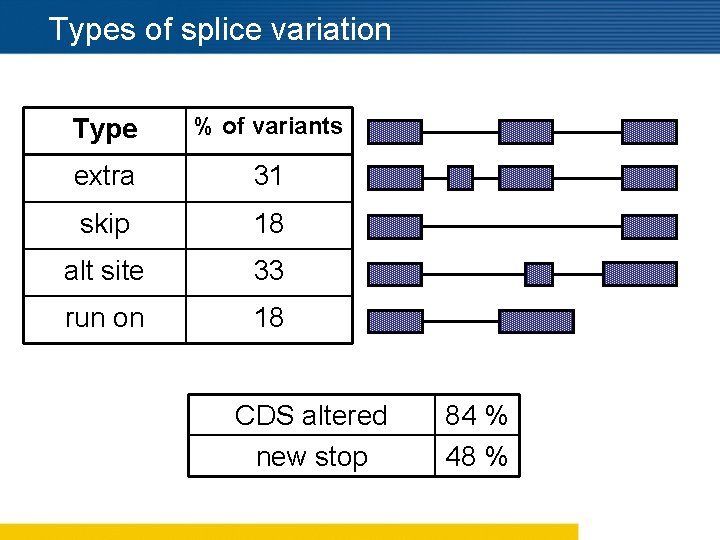
Types of splice variation Type % of variants extra 31 skip 18 alt site 33 run on 18 CDS altered new stop 84 % 48 %
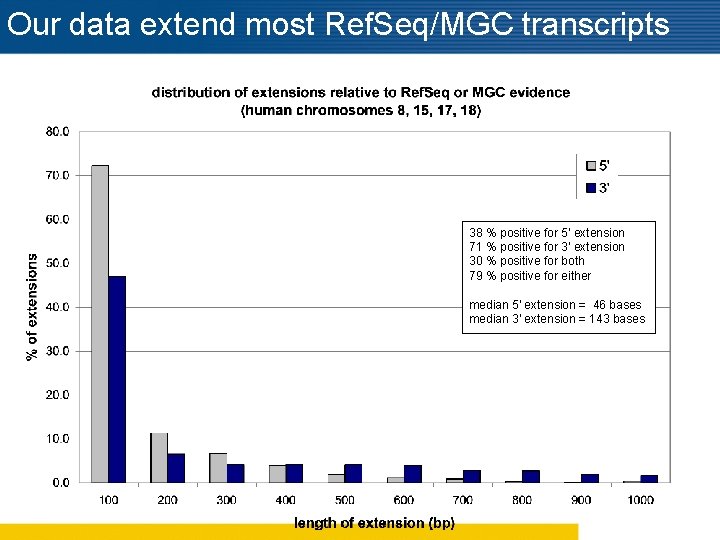
Our data extend most Ref. Seq/MGC transcripts 38 % positive for 5' extension 71 % positive for 3' extension 30 % positive for both 79 % positive for either median 5' extension = 46 bases median 3' extension = 143 bases
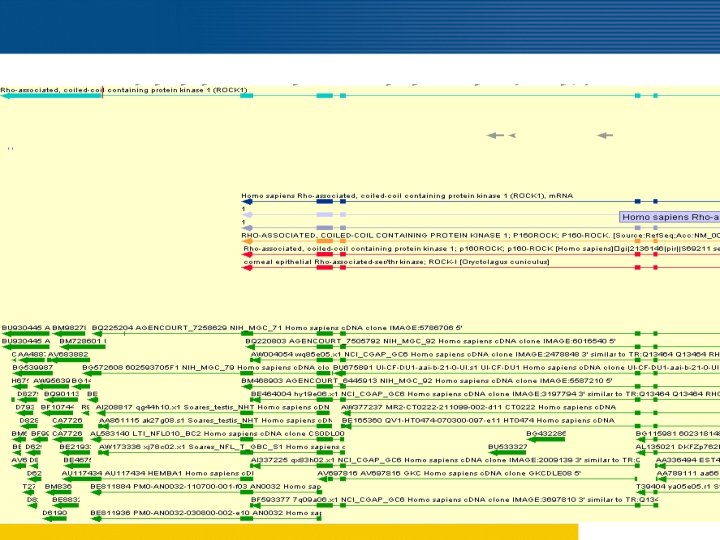
Complete 3 end as compared to Refseq m. RNA and ENSEMBL gene
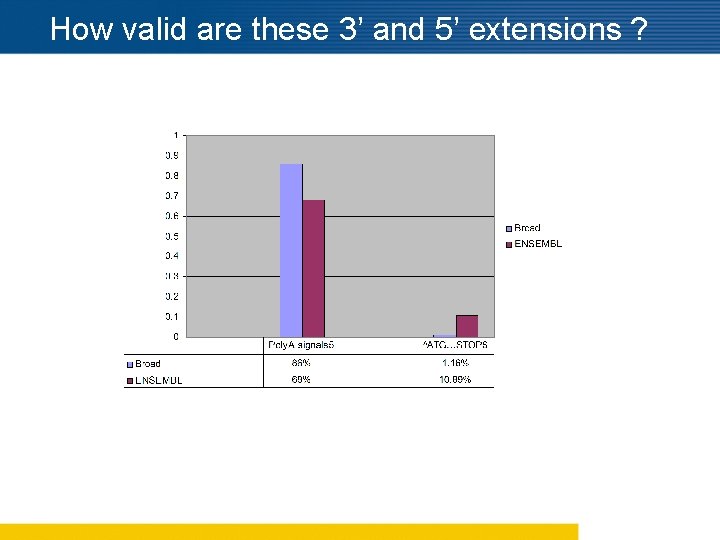
How valid are these 3’ and 5’ extensions ?
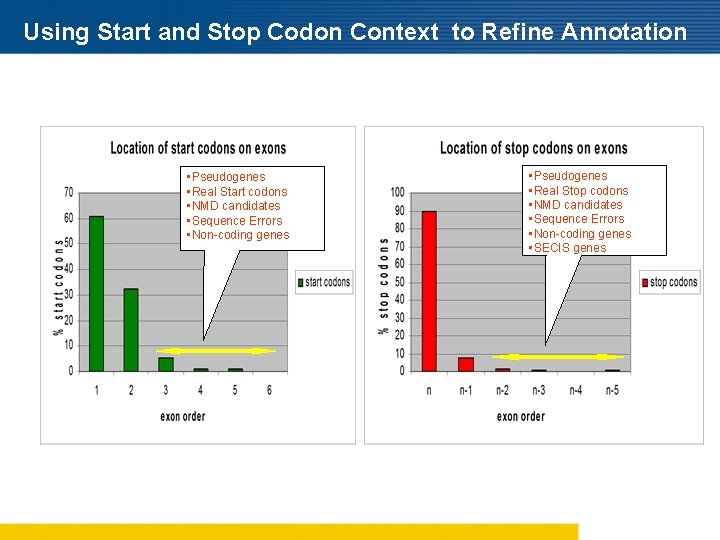
Using Start and Stop Codon Context to Refine Annotation • Pseudogenes • Real Start codons • NMD candidates • Sequence Errors • Non-coding genes • Pseudogenes • Real Stop codons • NMD candidates • Sequence Errors • Non-coding genes • SECIS genes
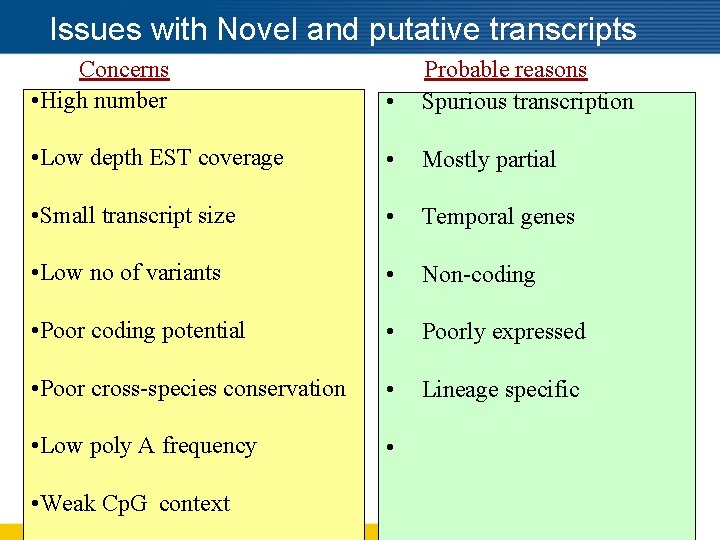
Issues with Novel and putative transcripts Concerns • High number • Probable reasons Spurious transcription • Low depth EST coverage • Mostly partial • Small transcript size • Temporal genes • Low no of variants • Non-coding • Poor coding potential • Poorly expressed • Poor cross-species conservation • Lineage specific • Low poly A frequency • • Weak Cp. G context
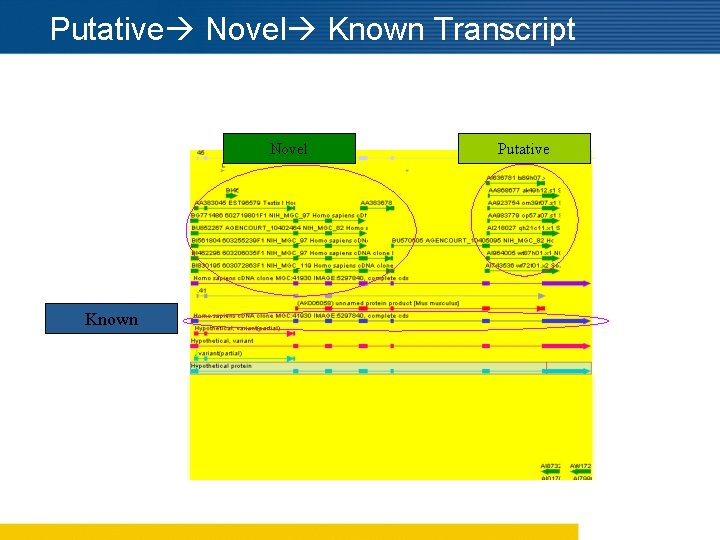
Putative Novel Known Transcript Novel Known Putative

Annotating Non-coding m. RNAs is still a challenge !!! Sno RNAs

Challenges Ahead…. n Establishing Common Standards n Validating Novel Transcripts n Single Exon Expressed Sequences n Determination of Accurate ORFs n Annotation of Functionally Relevant Alternative Splice Forms n Finding Sparsely Expressed Genes n Annotation of New Types of Non-coding Functional m. RNAs n Incremental Update of Annotation n Capturing Biological Exceptions

Acknowledgements Annotation Pipeline Annotation and Analysis • Charlie Whittaker • Reinhard Engels • Mark Borowsky • Shunguang Wang • Sinead O’leary • Seth Purcell • Tim Elkins • James Galagan • Yuhong Wu • Jill Mesirov • Serge Smirnov • Eric Lander • Sarah Calvo • Sequencing, Finishing and Closure Teams • David Dicaprio

Manually Annotated Gene Models vs. public Gene Models Comparison of alternative splice forms between ENSEMBL and Broad annotation db. EST nrnt-m. RNA ENSEMBL Refseq Broad

Novel Transcript Variants of Known Genes Poly. A signal MANUAL ANNOTATION Transcript Hunter REFSEQ GENEWISE ENSEMBL ESTs
- Slides: 27