Machine Learning Probabilistic Graphical Models with Scientific Applications

Machine Learning & Probabilistic Graphical Models (with Scientific Applications) Hagit Shatkay MQPEEGTGWLLELLSEVQLQQYFLRLRDDLN VTRLSHFEYVKNEDLEKIGMGRPGQRRLWEA VKRRKALCKRKSWMSKVFSGKRLEAEFPPHH SQSTFRKTSPAPGGPAGEGPLQSLTCLIGEKDL RLLEKLGDGSFGVVRRGEWDAPSGKTVSVAV KCLKPDVLSQPEAMDDFIREVNAMHSLDHRN LIRLYGVVLTPPMKMVTELAPLGSLLDRLRKH QGHFLLGTLSRYAVQVAEGMGYLESKR Computational Biomedicine & Machine Learning Lab Dept. of Computer and Information Sciences, Dept. of Biomed. Eng. University of Delaware Brown CS (Ph. D – ML, Robotics) NIH/NCBI (Post. Doc) Celera Genomics Queen’s, Kingston, ON U. of Delaware
Learning Models from Data (Model Fitting) Data Model Caution: Machine Learning © 2019. Hagit Shatkay, CIS (Deep) Neural Networks 2
Data ECG MQPEEGTGWLLELLSEVQLQQYFLRLRDDLNVTRLSHFEYV KNEDLEKIGMGRPGQRRLWEAVKRRKALCKRKSWMSKVF SGKRLEAEFPPHHSQSTFRKTSPAPGGPAGEGPLQSLTCLIGE KDLRLLEKLGDGSFGVVRRGEWDAPSGKTVSVAVKCLKPD VLSQPEAMDDFIREVNAMHSLDHRNLIRLYGVVLTPPMKM VTELAPLGSLLDRLRKHQGHFLLGTLSRYAVQVAEGMGYL ESKRFIHRDLAARNLLLATRDLVKIGDFGLMRALPQNDDHY VMQEHRKVPFAWCAPESLKTRTFSHASDTWMFGVTLWEM FTYGQEWIGLNGSQILHKIDKEGERLPRPEDCPQDIYNVMVQ CWAHKPEDRPTFVALRDFLLEAQPTDMRALQDEEPDKLHIQ MNDVITVIEGRAENYWWRGQNTRTLCVGPFPRNVVTSVAG LSAQDISQPLQNSFIHTGGDSDPRHCWGFPDYNVMVQCWA HKPEDRPTFVALRDFLLEAQPTDMRALQDFEEPDKLHIQMN DV http: //www. mcl. tulane. edu/sif/protein. jpg EEG © 2019. Hagit Shatkay, CIS 3
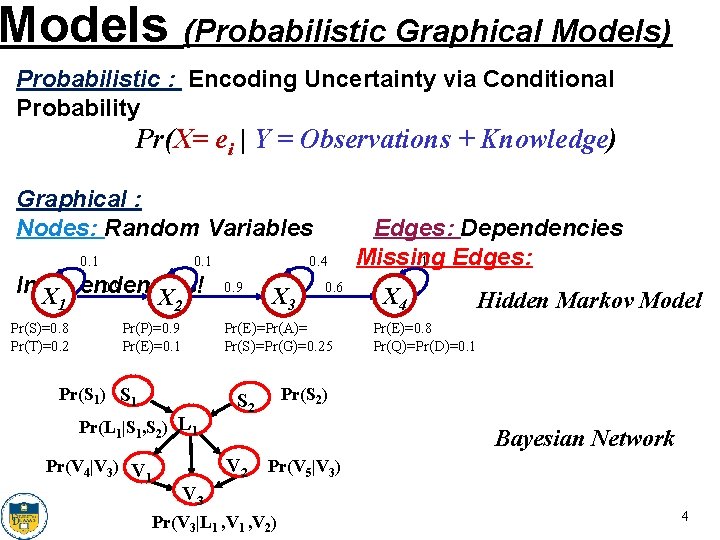
Models (Probabilistic Graphical Models) Probabilistic : Encoding Uncertainty via Conditional Probability Pr(X= ei | Y = Observations + Knowledge) Graphical : Nodes: Random Variables 0. 1 0. 9 Independencies! X 1 X 2 Pr(S)=0. 8 Pr(T)=0. 2 Pr(P)=0. 9 Pr(E)=0. 1 0. 9 X 3 0. 6 Pr(E)=Pr(A)= Pr(S)=Pr(G)=0. 25 Pr(S 1) S 1 Pr(L 1|S 1, S 2) L 1 Pr(V 4|V 3) V 1 0. 4 Edges: Dependencies Missing Edges: 1 X 4 Hidden Markov Model Pr(E)=0. 8 Pr(Q)=Pr(D)=0. 1 Pr(S 2) S 2 Bayesian Network V 2 Pr(V 5|V 3) V 3 Pr(V 3|L 1 , V 2) 4
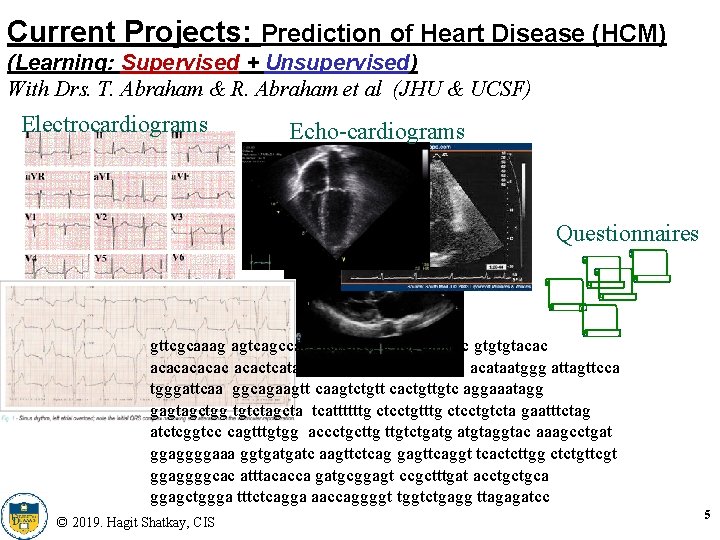
Current Projects: Prediction of Heart Disease (HCM) (Learning: Supervised + Unsupervised) With Drs. T. Abraham & R. Abraham et al (JHU & UCSF) Electrocardiograms Echo-cardiograms Questionnaires gttcgcaaag agtcagccat gactgagtgc acgcatatgc gtgtgtacacac acactcatat ttccagagca gacataatag acataatggg attagttcca tgggattcaa ggcagaagtt caagtctgtt cactgttgtc aggaaatagg gagtagctgg tgtctagcta tcattttttg ctcctgtcta gaatttctag atctcggtcc cagtttgtgg accctgcttg ttgtctgatg atgtaggtac aaagcctgat ggaggggaaa ggtgatgatc aagttctcag gagttcaggt tcactcttgg ctctgttcgt ggaggggcac atttacacca gatgcggagt ccgctttgat acctgctgca ggagctggga tttctcagga aaccaggggt tggtctgagg ttagagatcc © 2019. Hagit Shatkay, CIS 5
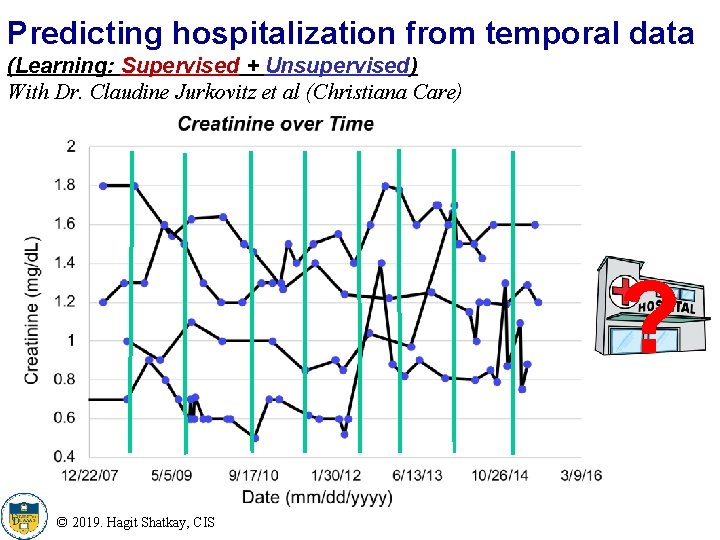
Predicting hospitalization from temporal data (Learning: Supervised + Unsupervised) With Dr. Claudine Jurkovitz et al (Christiana Care) ? © 2019. Hagit Shatkay, CIS

Computationally Glancing at (Biomedical) Images (Learning: Supervised + Unsupervised) Using Text and Image to find EVIDENCE: Relevant Info about genes proteins or drug-interactions… Martin Ringwald, Cynthia Smith, Judy Blake (Jackson Labs) Xiangying Jiang, Pengyuan Li, Gongbo Zhang (Udel) Prof. Liz Marai and her group (University of Ill. , Chicago) Prof. Cahndra Kambhamettu and Vinit Singh (UDel) Profs. Luis Rocha (Indiana University) and Lang Li (Ohio State) © 2019. Hagit Shatkay, CIS 7

New Science-enhanced Machine Learning for Astro-physics Event Detection With Christopher Tunnell & Waheed Bajwa
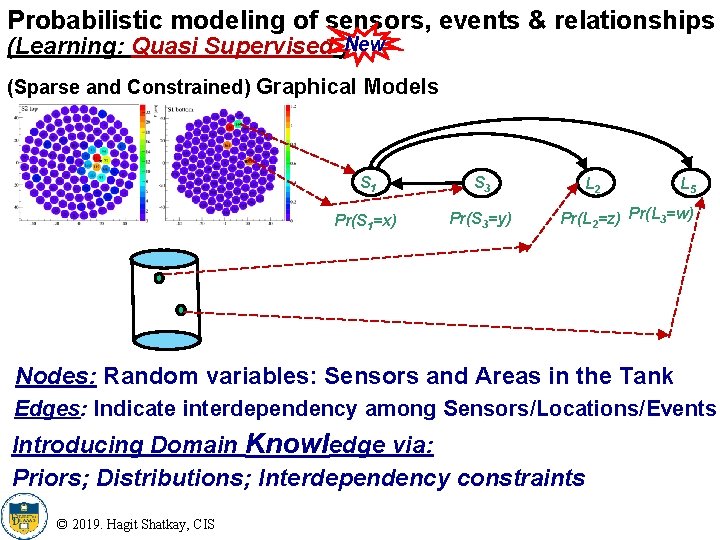
Probabilistic modeling of sensors, events & relationships (Learning: Quasi Supervised )New (Sparse and Constrained) Graphical Models S 1 S 3 Pr(S 1=x) Pr(S 3=y) L 2 L 5 Pr(L 2=z) Pr(L 3=w) Nodes: Random variables: Sensors and Areas in the Tank Edges: Indicate interdependency among Sensors/Locations/Events Introducing Domain Knowledge via: Priors; Distributions; Interdependency constraints © 2019. Hagit Shatkay, CIS

Thank You shatkay@cis. udel. edu http: //www. cis. udel. edu/~shatkay © 2019. Hagit Shatkay, CIS
- Slides: 10