Listeria monocytogenes Escape from the Gut Angela Harris




























- Slides: 35

Listeria monocytogenes: Escape from the Gut Angela Harris November 26 th, 2013

Listeria monocytogenes • Gram positive, rod-shaped foodborne pathogen that causes the bacterial infection listeriosis • Causes gastroenteritis, meningitis, encephalitis and mother-to-fetus infections usually by the consumption of contaminated food • Usually self limiting in healthy individuals, but may become severe in immunocompromised individuals, the elderly, pregnant woman, neonates, and young children http: //itthing. com/whats-in-your-fast-food/listeria-monocytogenes

Listeria monocytogenes • Can grow at temperatures ranging from 2 -45°C -ideal growth temperature 7°C • FDA has a zero-tolerance policy in food • Catalase positive, Oxidase negative, produces βhemolysin on blood agar • Infections are treated with ampicillin or gentamicin

Listeria monocytogenes • Enters the host primarily through the intestine • Can grow at p. H between 4. 6 and 9. 5 and can survive the acidic conditions of the stomach • Enters the small intestine, crosses the epithelial barrier, and spreads to other organs • Severe in immunocompromised individuals if crosses the blood-brain-barrier

Overview • Life Cycle of L. monocytogenes • Phagosomal Escape -Listeriolysin O -Type C Phospholipases -GILT + CFTR -Act. A • Autophagy • LC 3+ and SLAPs

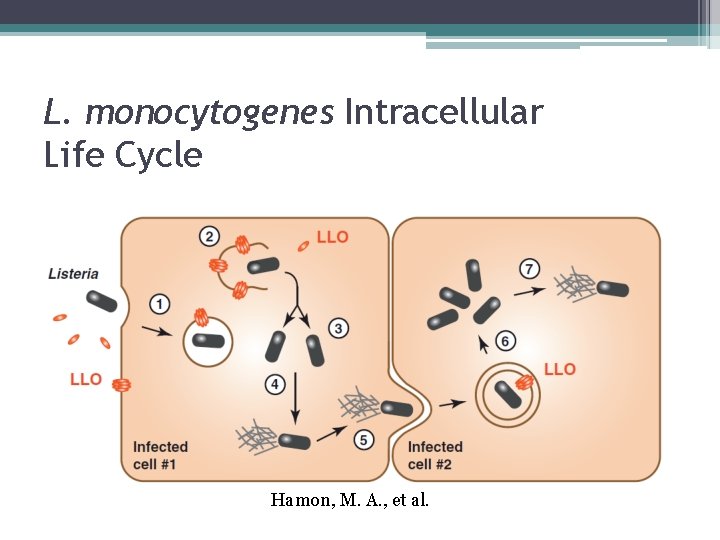
L. monocytogenes Intracellular Life Cycle Hamon, M. A. , et al.

L. monocytogenes Intracellular Life Cycle 1) Engulfed into a phagosome 2) Creation of listeriolysin O and phospholipase C causes the degradation of the phagosome and allows entry into the cytoplasm 3) Replicates in the cytoplasm 4) Expression of Act. A causes the polymerization of actin

L. monocytogenes Intracellular Life Cycle (continued) 5) Able to propel across the cytosol and spread to other cells where it becomes entrapped in a double membrane 6) The creation of listeriolysin O and phospholipase C helps in the degradation and escape from the double membrane phagosome 7) The intracellular life cycle repeats

Phagosomal Escape Bacterial Factors • Listeriolysin O • Two type C phospholipases: Phosphatidylinositol (PI-PLC) Phosphatidylcholine (PC-PLC) • Act. A Host Factors • GILT and CFTR

Listeriolysin O (LLO) • Pore-forming toxin • Belongs to the family of cholesterol-dependent cytolysins (CDCs) • Secreted as soluble monomers that bind to cholesterol of host membranes and oligomerize to form pores • Activated in acidic environment of phagosomes • Helps escape from primary or secondary phagosome

Listeriolysin O Pore Formation • 2 models for pore formation 1 -Monomers oligomerize to form a prepore ring Pores always the same size 2 -Pore is formed even if the ring is not complete. Size may vary and the pore will be permeable to ions Hamon, M. A. , et al.

Intracellular Listeriolysin O Regulation of NOX 2 NADPH Oxidation activation • Produces Reactive Oxygen Species (ROS) in macrophages -play a key role in limiting bacterial replication • LLO is able to suppress ROS production Regulation of Autophagy • In later slides

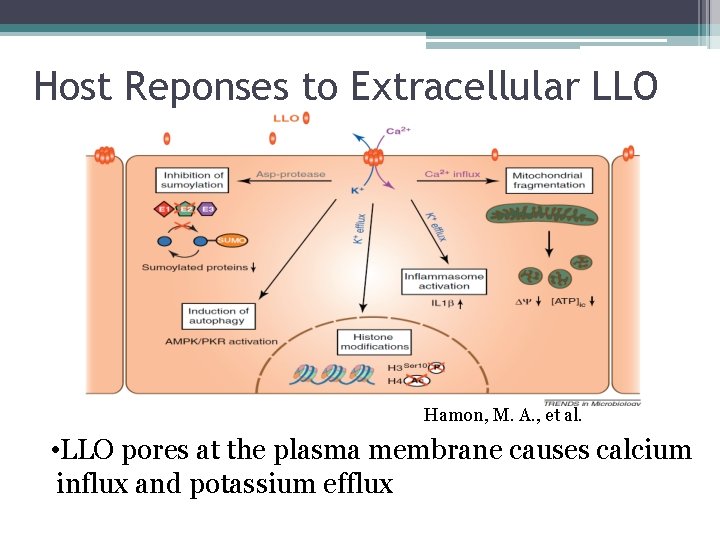
Host Reponses to Extracellular LLO Hamon, M. A. , et al. • LLO pores at the plasma membrane causes calcium influx and potassium efflux

Extracellular Listeriolysin O Deregulation of the SUMOylation machinery • SUMO: Small Ubiquitinated-like Modifier • Post-translational modification involved in transcription regulation, protein stability, etc. • Degrades E 2 SUMO -Blocks the host SUMOylation machinery -Results in de. SUMOylation of host proteins

Extracellular Listeriolysin O Immune activity • Adaptive immune response: Targets CD 4+ T-Cells (induces unresponsiveness) • Innate Immune Response: LLO sensed by TLR 4 Inflammasome activation • Recognition and containment of microbial pathogens • Secretes pro-inflammatory cytokines IL-Iβ and IL-18 Induction of Autophagy • AMPK/PKR activation • Host response

Extracellular Listeriolysin O LLO effect on host cell organelles • CDCs do not terminally damage organelles Mitochondria • Fragmentation of mitochondrial network by influx of calcium ions • Temporary shutdown of cellular bioenergetics resulting in a decrease in intracellular ATP • Interferes with host response to infection Endoplasmic Reticulum • Causes vesiculation

Extracellular Listeriolysin O Histone Modifications • Post translational modification of N-terminal histone tails causes changes in gene expression • LLO causes dephosphorylation of H 3 which downregulates several immune genes http: //www. nature. com/nrc/journal/v 1/n 3/fig_tab/nrc 1201 -194 a_F 1. html

Type C Phospholipases • Phosphatidylinositol-specific (PI-PLC) and Phosphatidylcholine (PC-PLC) • Activated by acidic p. H in the phagosome • Aid in phagosome and double-membrane phagosome escape by digesting the membrane • PI-PLC can produce the signalling molecule diacylglycerol (DAG) upon cleavage of phosphatidylinositol • L. monocytogenes can activate both host and bacterial phospholipases

Act. A • Mediates the addition of an actin tail on the end of the bacteria which allows it to spread across the cytosol and infect other cells • Act. A has been shown to help evade autophagy in the cytoplasm by preventing ubiquitination • Act. A mutants are targeted by adaptor proteins such as p 62 which leads to autophagy

Host Factors γ-interferon-inducible lysosomal thiol reductase (GILT) • Activates LLO via a thiol reductase mechanism • Active at low p. H (only in phagosomes) Cystic fibrosis trans membrane conductance regulator (CFTR) • Creates a channel that increases chloride concentration into the phagosome • Aids in LLO pore formation

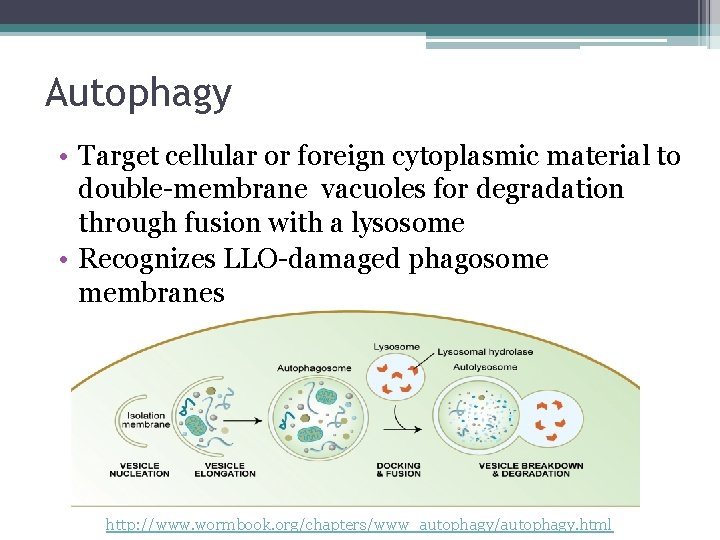
Autophagy • Target cellular or foreign cytoplasmic material to double-membrane vacuoles for degradation through fusion with a lysosome • Recognizes LLO-damaged phagosome membranes http: //www. wormbook. org/chapters/www_autophagy/autophagy. html

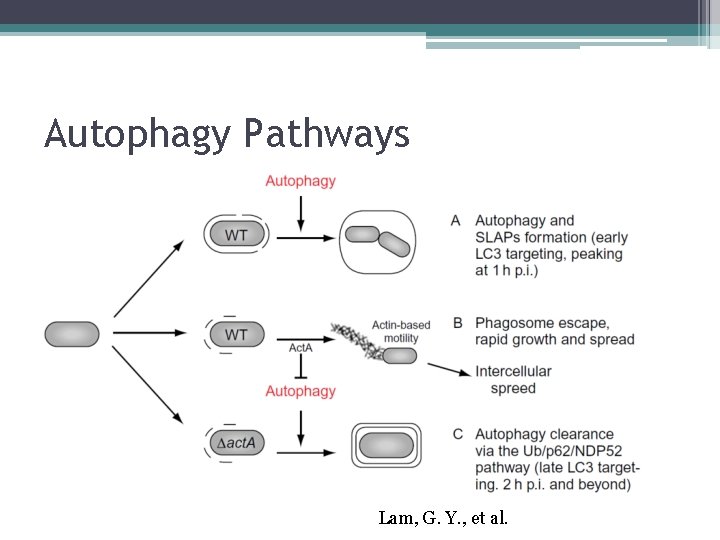
Autophagy Pathways Lam, G. Y. , et al.

LC 3 -Associated Phagocytosis (LAP) Pathway • Microtubule-associated protein 1 A/1 B-light chain 3 (LC 3) • Involved in promoting phagosome maturation, killing of microbes, and suppressing proinflammatory signals • Requires some components of the autophagy pathway • ROS production by the CYBB/NOX 2 NADPH oxidase is required

LC 3+ and SLAP Formation Study • L. monocytogenes co-localizes with LC 3 at 1 hour post-infection which is thought to give rise to spacious Listeria-containing phagosomes (SLAPs) • SLAPs: Membrane bound compartments that harbor slow-growing bacteria that is associated with persistent L. monocytogenes infection -“stalemate” between the host and the bacteria

LC 3+ and SLAP Formation Study Bacterial colocalize with LC 3 in the phagosome at 1 hour post infection • Creation of a probe: Cell wall binding domain fused to yellow fluorescent protein -can only bind to cytosolic bacteria • No accumulation of CBD-YFP to the population after one hour

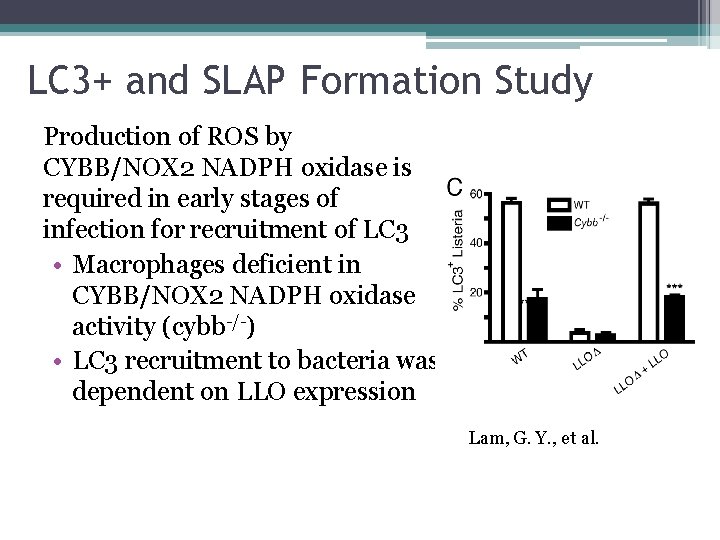
LC 3+ and SLAP Formation Study Production of ROS by CYBB/NOX 2 NADPH oxidase is required in early stages of infection for recruitment of LC 3 • Macrophages deficient in CYBB/NOX 2 NADPH oxidase activity (cybb-/-) • LC 3 recruitment to bacteria was dependent on LLO expression Lam, G. Y. , et al.

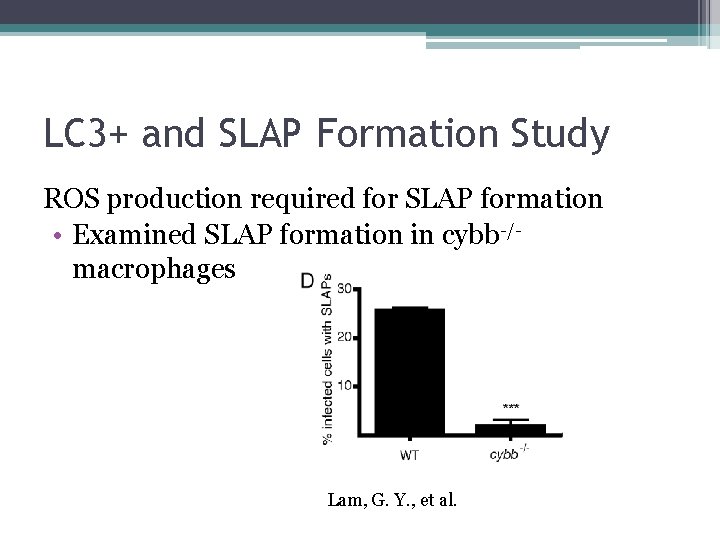
LC 3+ and SLAP Formation Study ROS production required for SLAP formation • Examined SLAP formation in cybb-/macrophages Lam, G. Y. , et al.

LC 3+ and SLAP Formation Study DAG produced by host and bacterial phospholipases • Generation of DAG is upstream of ROS production • Creation of a probe: DAG binding C 1 domain of PRKCD fused to green fluorescent protein • Used mutant strains lacking PI-PLC and PC-PLC (block bacteria) • Added inhibitors phospholipase D and phosphatidic acid phosphatase (PPAP 2 A) (block host)

LC 3+ and SLAP Formation Study Results • If both host and bacterial production of DAG blocked resulted in a decrease in DAG colocalization on phagosomes • However if only bacterial or host production of DAG blocked still accumulation of DAG on phagosomes • This suggests that both host and bacterial factors contribute to DAG accumulation on phagosomes

LC 3+ and SLAP Formation Study DAG Production is needed for LC 3 recruitment early in infection • LC 3 recruitment to both wild-type and PI-PLC and PC-PLC mutants upon treatment with PLD and PPAP 2 A inhibitors Results • No change in wild type when treated with PLD and PPAP 2 A inhibitors • Significant decrease in LC 3 recruitment in PC-PLC and PC-PLC mutants upon treatment with PLD and PPAP 2 A inhibitors

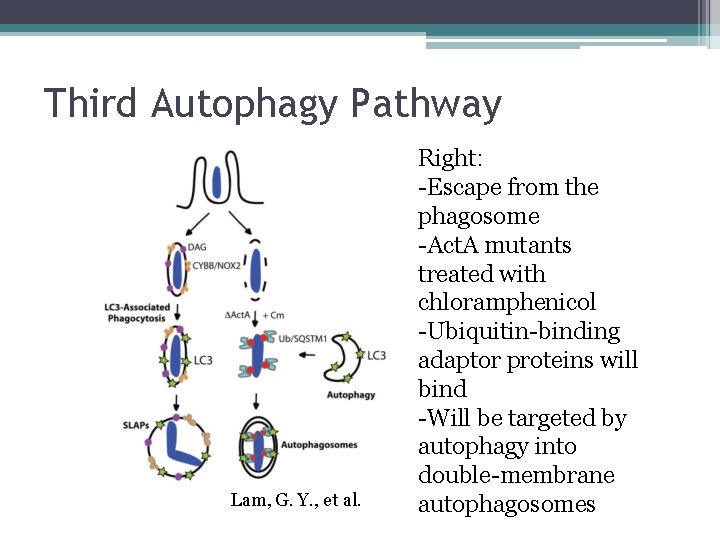
Third Autophagy Pathway Lam, G. Y. , et al. Right: -Escape from the phagosome -Act. A mutants treated with chloramphenicol -Ubiquitin-binding adaptor proteins will bind -Will be targeted by autophagy into double-membrane autophagosomes

Fourth Autophagy Pathway • Successful clearance of the bacteria before the escape from the phagosome or SLAP formation by fusion to a lysosome Birmingham, C. L. , et al.

Conclusions Four different pathways: • Escape from the phagosome via LLO and phospholipase C and spread from cell to cell • Recruitment of LC 3 and creation of SLAPs resulting in chronic L. monocytogenes infection • Act. A mutants are targeted in the cytoplasm by ubiquitin binding proteins which result in the creation of double membrane autophagosomes • Clearance by phagosome fusion to lysosome

Future Directions • Use of intracellular lifestyle of Listeria monocytogenes as a potential model for other intracellular pathogens -LAP Pathway -SLAPs • LLO questions

Questions?