Liquid biopsies for the early detection of PDAC

Liquid biopsies for the early detection of PDAC 2019년 5월 2일 소화기내과 목요세미나 R 3 박영수 / Pf. 황진혁
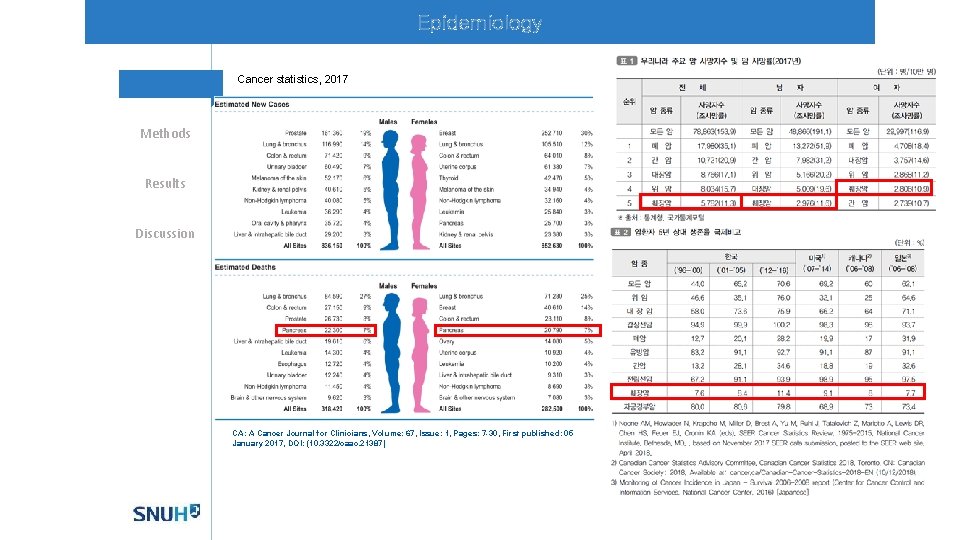
Epidemiology Cancer statistics, 2017 Methods Results Discussion CA: A Cancer Journal for Clinicians, Volume: 67, Issue: 1, Pages: 7 -30, First published: 05 January 2017, DOI: (10. 3322/caac. 21387)
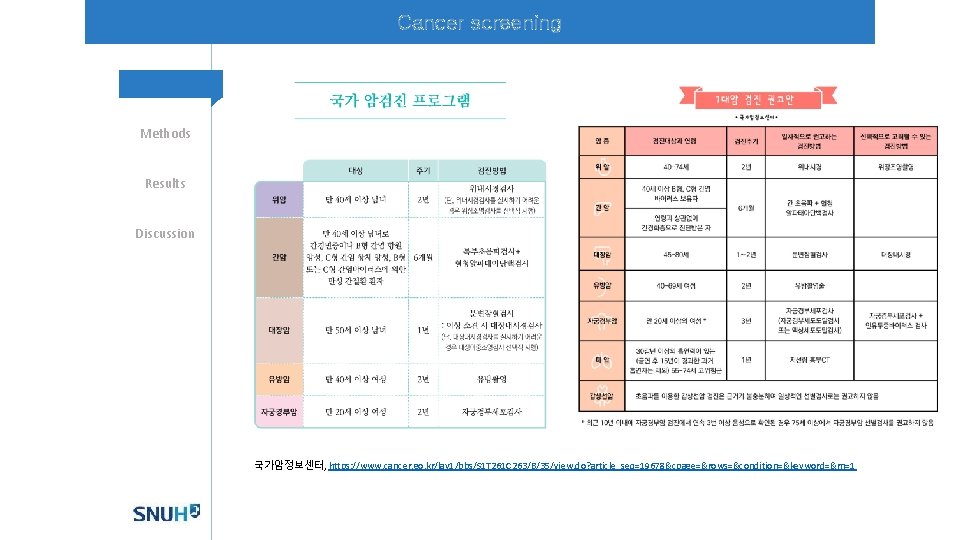
Cancer screening Methods Results Discussion 국가암정보센터, https: //www. cancer. go. kr/lay 1/bbs/S 1 T 261 C 263/B/35/view. do? article_seq=19678&cpage=&rows=&condition=&keyword=&rn=1
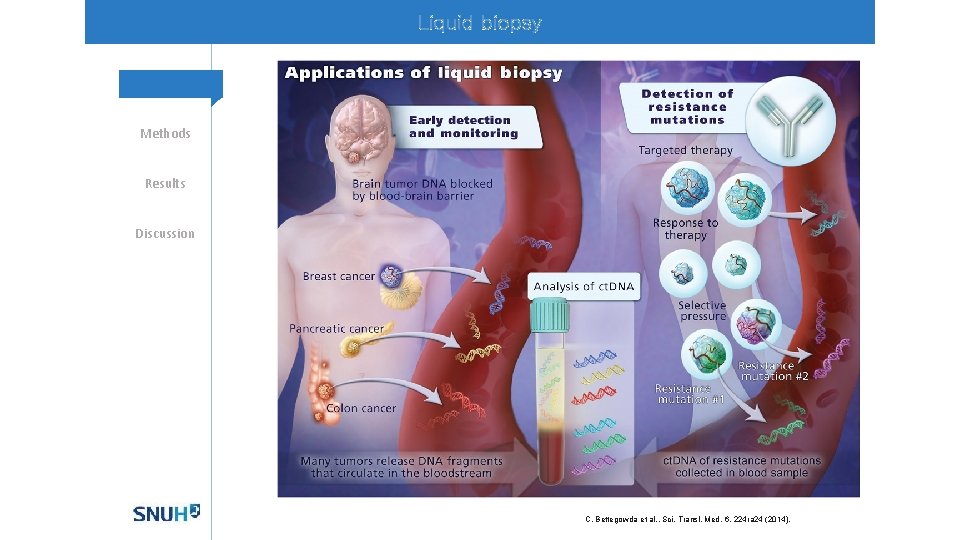
Liquid biopsy Methods Results Discussion C. Bettegowda et al. , Sci. Transl. Med. 6, 224 ra 24 (2014).

Initial studies Methods Results Discussion

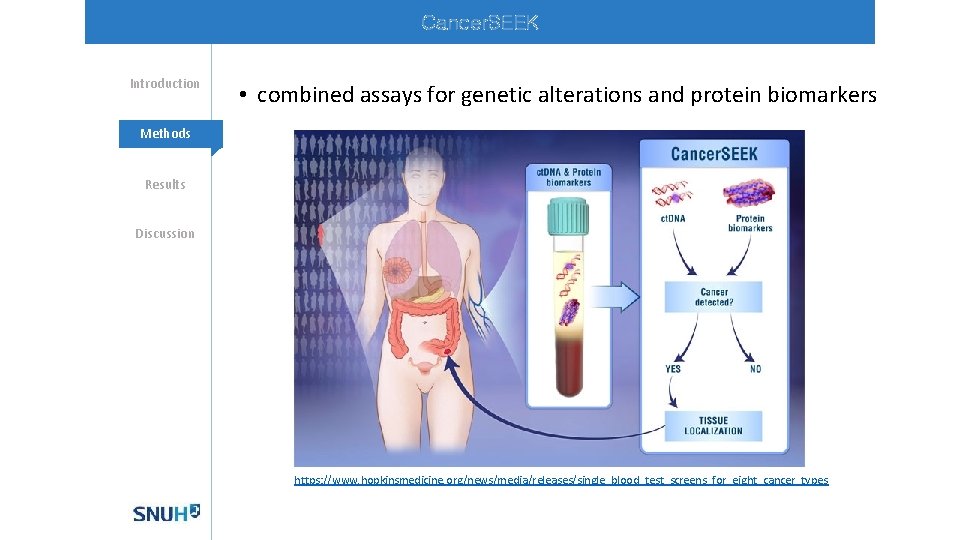
Cancer. SEEK Introduction • combined assays for genetic alterations and protein biomarkers Methods Results Discussion https: //www. hopkinsmedicine. org/news/media/releases/single_blood_test_screens_for_eight_cancer_types
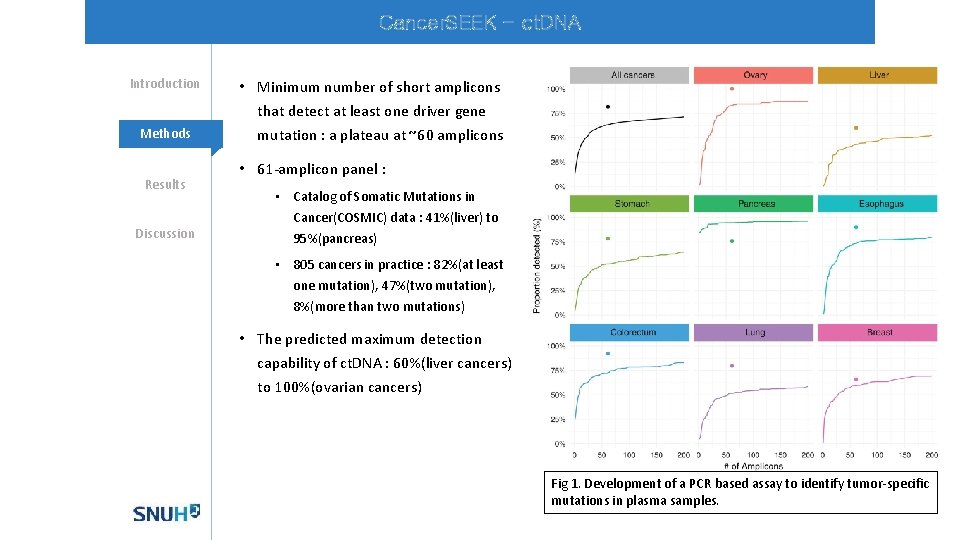
Cancer. SEEK - ct. DNA Introduction Methods Results Discussion • Minimum number of short amplicons that detect at least one driver gene mutation : a plateau at ~60 amplicons • 61 -amplicon panel : • Catalog of Somatic Mutations in Cancer(COSMIC) data : 41%(liver) to 95%(pancreas) • 805 cancers in practice : 82%(at least one mutation), 47%(two mutation), 8%(more than two mutations) • The predicted maximum detection capability of ct. DNA : 60%(liver cancers) to 100%(ovarian cancers) Fig 1. Development of a PCR based assay to identify tumor-specific mutations in plasma samples.
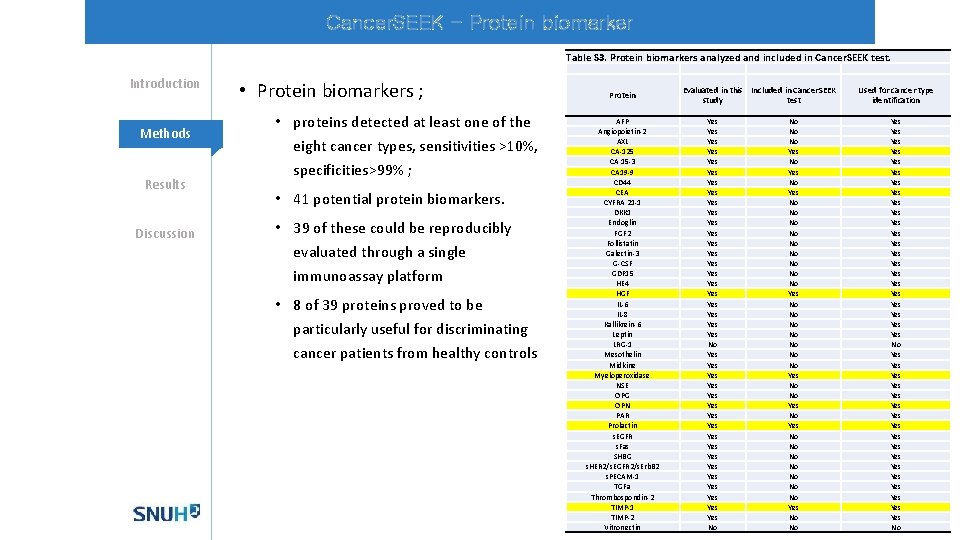
Cancer. SEEK - Protein biomarker Table S 3. Protein biomarkers analyzed and included in Cancer. SEEK test. Introduction Methods Results Discussion • Protein biomarkers ; • proteins detected at least one of the eight cancer types, sensitivities >10%, specificities>99% ; • 41 potential protein biomarkers. • 39 of these could be reproducibly evaluated through a single immunoassay platform • 8 of 39 proteins proved to be particularly useful for discriminating cancer patients from healthy controls Protein AFP Angiopoietin-2 AXL CA-125 CA 15 -3 CA 19 -9 CD 44 CEA CYFRA 21 -1 DKK 1 Endoglin FGF 2 Follistatin Galectin-3 G-CSF GDF 15 HE 4 HGF IL-6 IL-8 Kallikrein-6 Leptin LRG-1 Mesothelin Midkine Myeloperoxidase NSE OPG OPN PAR Prolactin s. EGFR s. Fas SHBG s. HER 2/s. EGFR 2/s. Erb. B 2 s. PECAM-1 TGFa Thrombospondin-2 TIMP-1 TIMP-2 Vitronectin Evaluated in this Included in Cancer. SEEK study test Yes Yes Yes Yes Yes Yes No Yes Yes Yes Yes Yes No No Yes No No No Yes No No No No Yes No No Used for cancer type identification Yes Yes Yes Yes Yes Yes No Yes Yes Yes Yes Yes No
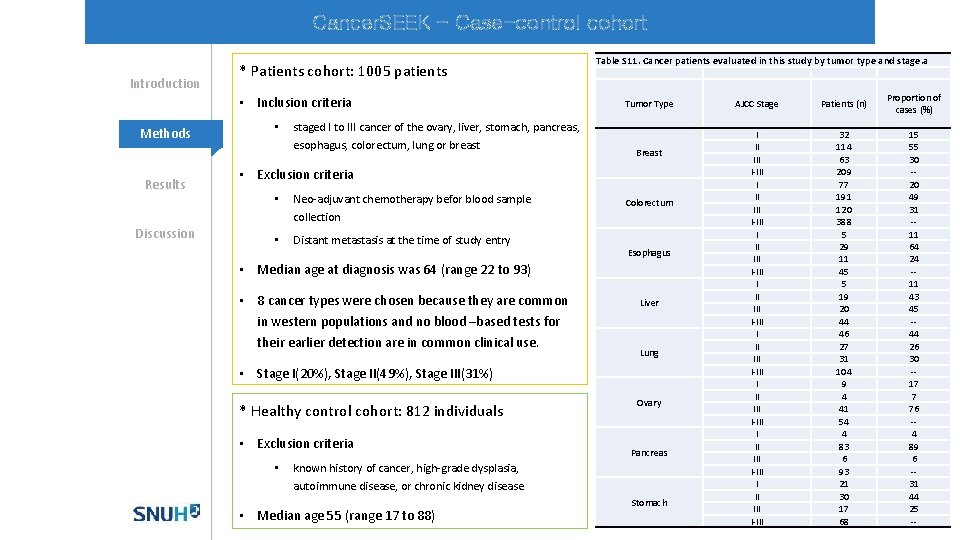
Cancer. SEEK - Case-control cohort Introduction * Patients cohort: 1005 patients • Inclusion criteria Methods Results Discussion • staged I to III cancer of the ovary, liver, stomach, pancreas, esophagus, colorectum, lung or breast Table S 11. Cancer patients evaluated in this study by tumor type and stage. a Tumor Type Breast • Exclusion criteria • Neo-adjuvant chemotherapy befor blood sample collection • Distant metastasis at the time of study entry Colorectum Esophagus • Median age at diagnosis was 64 (range 22 to 93) • 8 cancer types were chosen because they are common in western populations and no blood –based tests for their earlier detection are in common clinical use. Liver Lung • Stage I(20%), Stage II(49%), Stage III(31%) * Healthy control cohort: 812 individuals • Exclusion criteria • Ovary Pancreas known history of cancer, high-grade dysplasia, autoimmune disease, or chronic kidney disease • Median age 55 (range 17 to 88) Stomach AJCC Stage Patients (n) Proportion of cases (%) I II III I-III I II III I-III 32 114 63 209 77 191 120 388 5 29 11 45 5 19 20 44 46 27 31 104 9 4 41 54 4 83 6 93 21 30 17 68 15 55 30 -20 49 31 -11 64 24 -11 43 45 -44 26 30 -17 7 76 -4 89 6 -31 44 25 --

Cancer. SEEK - Probability of cancer Introduction Methods Results • Cancer. SEEK evaluates levels of eight proteins and the presence of mutations in 1933 distinct genomic positions (single base substitutions, insertions, or deletions) • The presence of a mutation in an assayed gene or an elevation in the level of any of these proteins would classify a patient as positive Discussion • Use log ratios to evaluate mutations and incorporated them into a logistic regression algorithm; mutation data and protein biomarker levels to score Cancer SEEK test results • The mean sensitivities and specificities were determined by 10 iterations of 10 -fold cross validations.
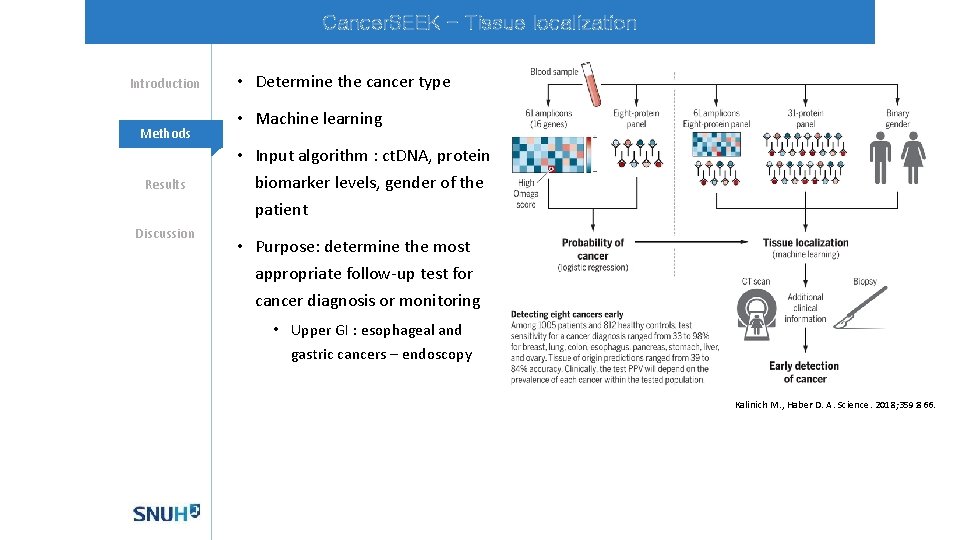
Cancer. SEEK - Tissue localization Introduction Methods Results Discussion • Determine the cancer type • Machine learning • Input algorithm : ct. DNA, protein biomarker levels, gender of the patient • Purpose: determine the most appropriate follow-up test for cancer diagnosis or monitoring • Upper GI : esophageal and gastric cancers – endoscopy Kalinich M. , Haber D. A. Science. 2018; 359: 866.
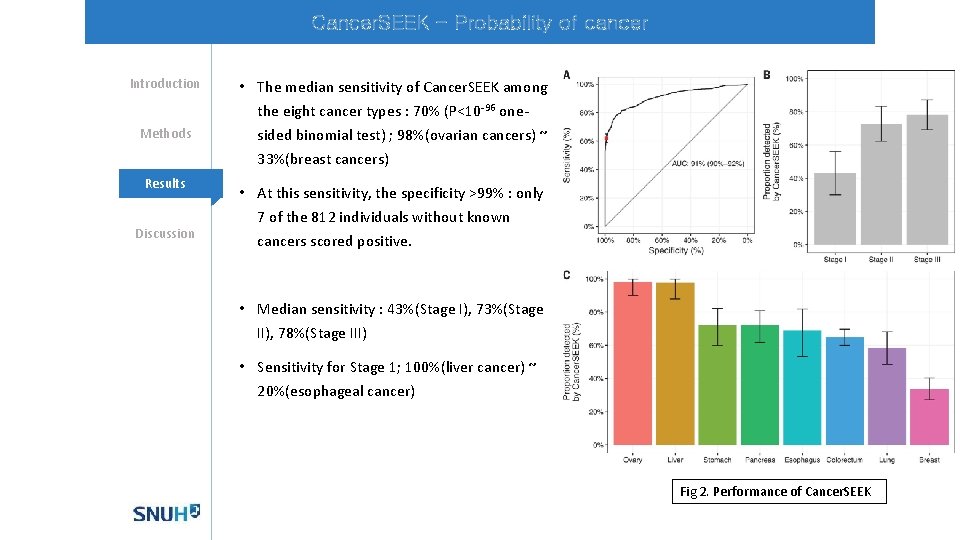
Cancer. SEEK - Probability of cancer Introduction Methods Mechanisms Results Discussion • The median sensitivity of Cancer. SEEK among the eight cancer types : 70% (P<10 -96 onesided binomial test) ; 98%(ovarian cancers) ~ 33%(breast cancers) • At this sensitivity, the specificity >99% : only 7 of the 812 individuals without known cancers scored positive. • Median sensitivity : 43%(Stage I), 73%(Stage II), 78%(Stage III) • Sensitivity for Stage 1; 100%(liver cancer) ~ 20%(esophageal cancer) Fig 2. Performance of Cancer. SEEK
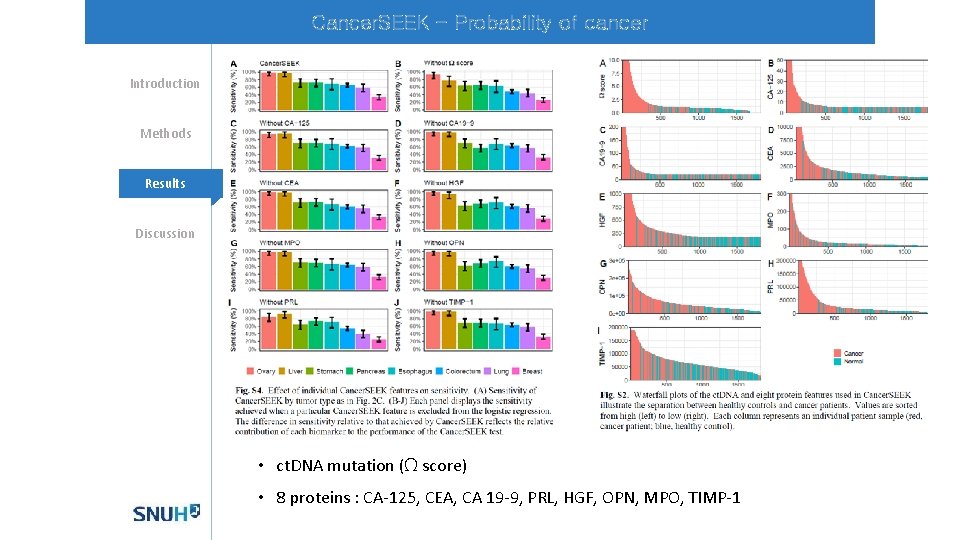
Cancer. SEEK - Probability of cancer Introduction Methods Mechanisms Results Discussion • ct. DNA mutation (Ω score) • 8 proteins : CA-125, CEA, CA 19 -9, PRL, HGF, OPN, MPO, TIMP-1

Cancer. SEEK - ct. DNA from cancer cells Introduction Methods Mechanisms Results Discussion • The basis of liquid biopsy : mutant DNA templates in plasma are derived from dying cancer cells • Evaluated tumor tissue from 153 patients in whom ct. DNA could be detected at statistically significant levels and for whom primary tumors were available • The mutation in the plasma was identical to a mutation found in the primary tumor of the same individual in 138(90%) of 153 • ; 100%(ovarian cancer, pancreatic cancer) ~ 82%(stomach cancer)
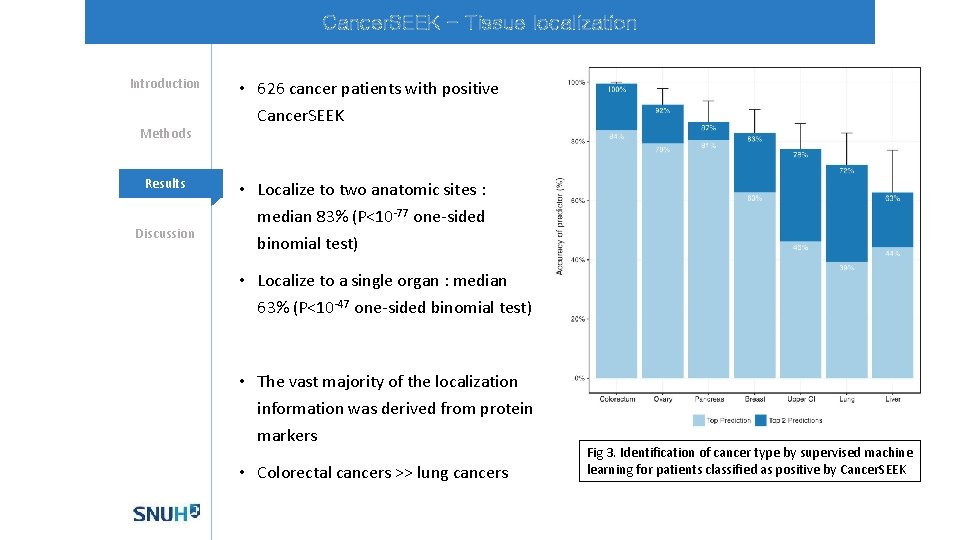
Cancer. SEEK - Tissue localization Introduction Methods Mechanisms Results Discussion • 626 cancer patients with positive Cancer. SEEK • Localize to two anatomic sites : median 83% (P<10 -77 one-sided binomial test) • Localize to a single organ : median 63% (P<10 -47 one-sided binomial test) • The vast majority of the localization information was derived from protein markers • Colorectal cancers >> lung cancers Fig 3. Identification of cancer type by supervised machine learning for patients classified as positive by Cancer. SEEK

Strength Introduction Methods Results Discussion • Combined protein biomarkers with genetic biomarkers to increase sensitivity without substantially decreasing specificity • Provide additional information that could help identify those patients most likely to harbor a malignancy • Cost would be less than $500 • The eight cancer types account for 360, 000(60%) of the estimated cancer deaths in the US in 2017 ; earlier detection could reduce deaths from these disease.

Limitations Introduction Methods Results Discussion • The patient cohort was composed of individuals with known cancers, most diagnosed on the basis of symptoms of disease. • Most individuals in a true screening setting would have less advanced disease, the sensitivity of detection is likely to be less than reported here. • The controls were limited to healthy individuals, whereas in a true cancer screening setting, some individuals might have inflammatory or other disease. • False positive results than observed in this study.

Limitations Introduction Methods Results Discussion • Not able to use a completely independent set of cases for testing, which would have been optimal. • The proportion of cancers of each type in the cohort was purposefully not representative of those in the US as a whole because they wanted to evaluate at least 50 examples of each cancer type with the resources available • when weighted for actual incidence in the US, estimated sensitivity of Cancer. SEEK to be 55% among all eight cancer types. • This weighting would not affect the high sensitivities of Cancer. SEEK (69 to 98%) to detect 5 cancer types (ovary, liver, stomach, pancreas, esophagus), no screening tests available for average –risk individuals.


Study Designs Introduction Methods • Two restrospective studies • The Scandinavian cohort : identification study • The US(United States) cohort : validation study Results Discussion • PDAC staging : AJCC 7 th edition 2010 • blood samples were collected and processed before resection or start of chemotherapy : • 5 u. L of the serum samples • a recombinant antibody microarray platform ; 349 human recombinant sc. Fvs(single-chain variable fragments) directed against 156 antigens
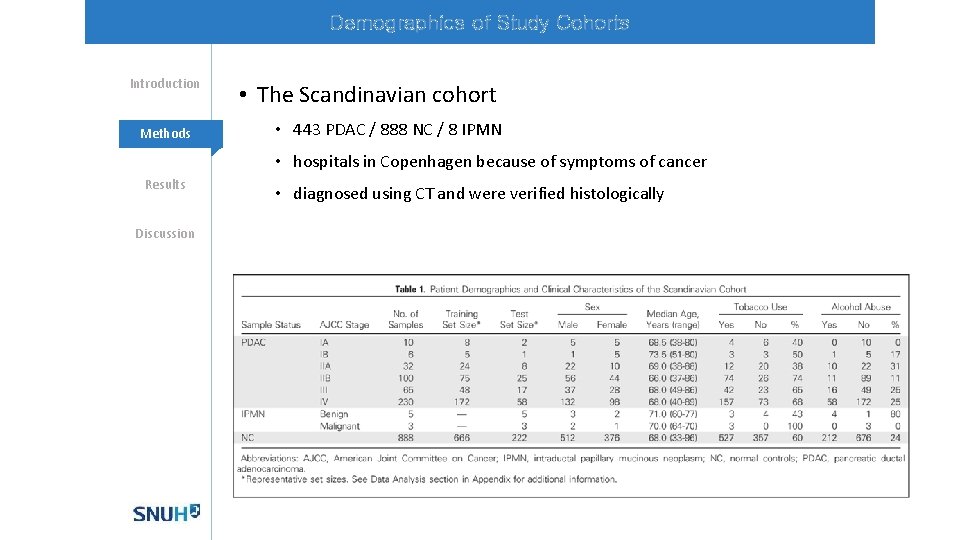
Demographics of Study Cohorts Introduction Methods • The Scandinavian cohort • 443 PDAC / 888 NC / 8 IPMN • hospitals in Copenhagen because of symptoms of cancer Results Discussion • diagnosed using CT and were verified histologically
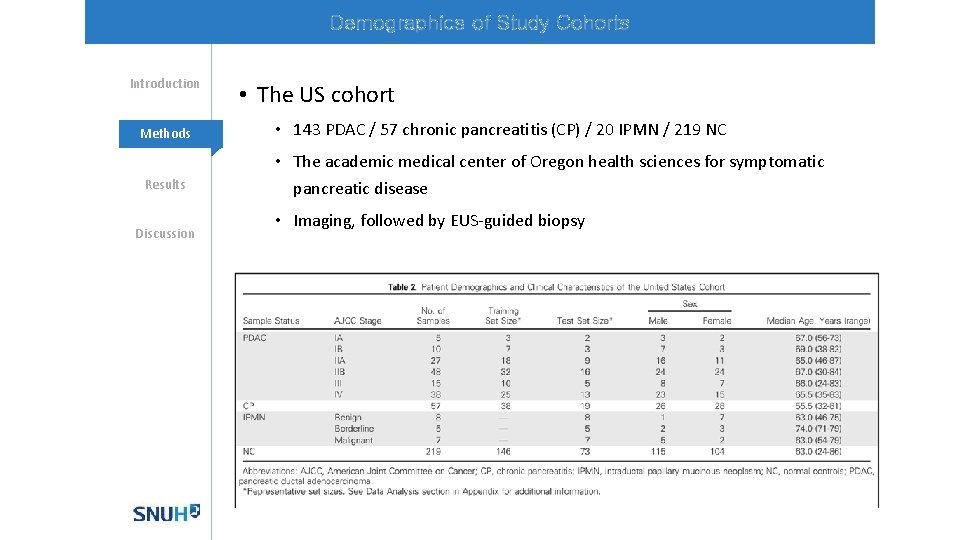
Demographics of Study Cohorts Introduction Methods Results Discussion • The US cohort • 143 PDAC / 57 chronic pancreatitis (CP) / 20 IPMN / 219 NC • The academic medical center of Oregon health sciences for symptomatic pancreatic disease • Imaging, followed by EUS-guided biopsy

Biomarker selection Introduction Methods Results Discussion • Training set : Test set = 3: 1 -> 4 unique test and training set • Backward elimination (BE) algorithm • For each BE iteration, the antibody with the highest Kullback-Leibler divergence value obtained in the classification analysis was eliminated • the antibody combinations expressing the lowest values were used to design the predictive biomarker signature. • The identified biomarker signature was then used to build a prediction model by frozen support vector machine (SVM) in R, using only the training data set. • To avoid overfitting, the model was tested on the corresponding test set ; ROC cureves and AUC values. This process was performed on all four training and test sets.
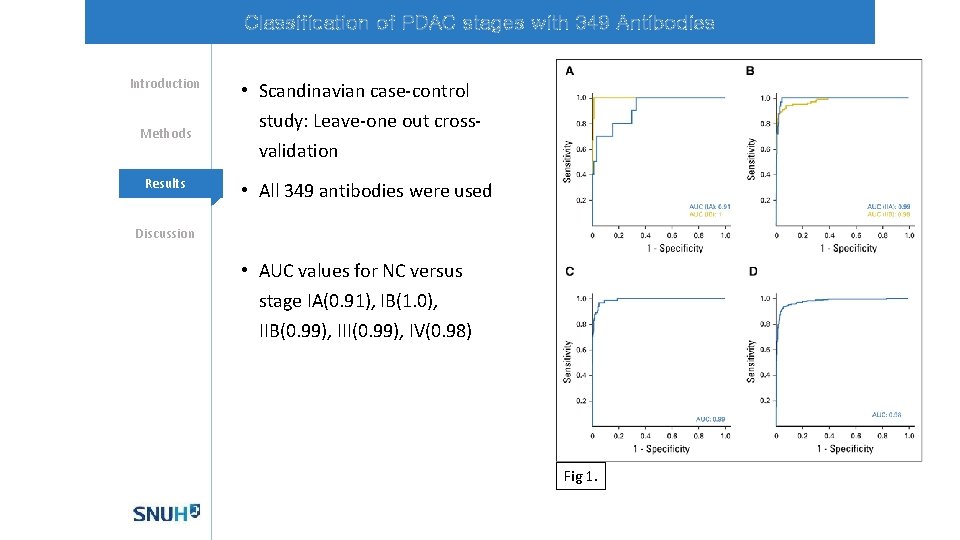
Classification of PDAC stages with 349 Antibodies Introduction Methods Mechanisms Results • Scandinavian case-control study: Leave-one out crossvalidation • All 349 antibodies were used Discussion • AUC values for NC versus stage IA(0. 91), IB(1. 0), IIB(0. 99), III(0. 99), IV(0. 98) Fig 1.

Classification PDAC stage I and II with a Biomarker Signature Introduction Methods Mechanisms Results Discussion Fig 2. • AUC value stage I and II vs NC was 0. 96 (Fig 2 A) • specificity/sensitivity 94%/95% • false positive rate 6% • AUC value stage III and IV vs NC was 0. 98 (Fig 2 B)
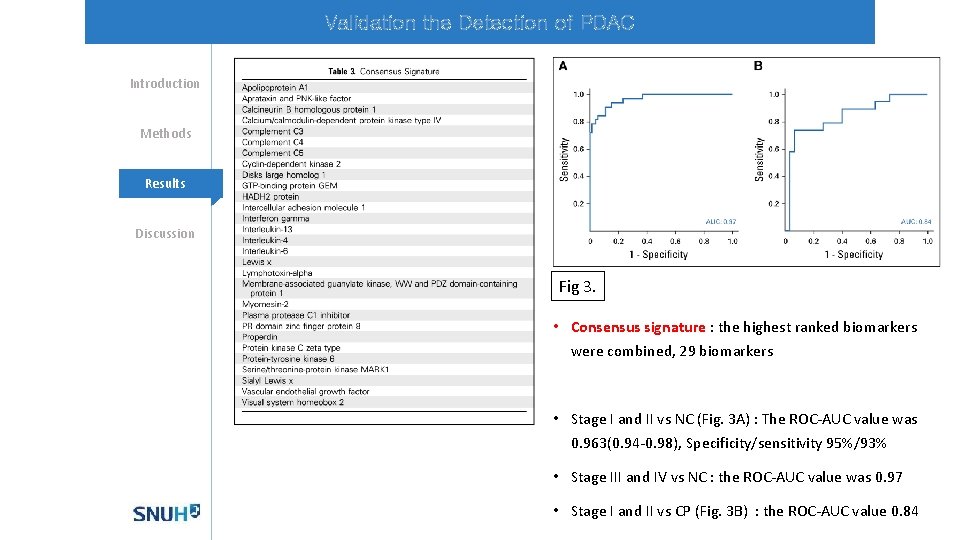
Validation the Detection of PDAC Introduction Methods Mechanisms Results Discussion Fig 3. • Consensus signature : the highest ranked biomarkers were combined, 29 biomarkers • Stage I and II vs NC (Fig. 3 A) : The ROC-AUC value was 0. 963(0. 94 -0. 98), Specificity/sensitivity 95%/93% • Stage III and IV vs NC : the ROC-AUC value was 0. 97 • Stage I and II vs CP (Fig. 3 B) : the ROC-AUC value 0. 84

Influence of DM and Jaundice Introduction Methods Mechanisms Results Discussion • In the Scandinavian cohort, 103(23. 3%) with PDAC were diabetic • The US cohort, 38(26. 6%) with PDAC had diabetes • New-onset diabetes (NOD) comprised 26. 2% of diabetes in both cohort • Diabetes, including NOD, is not a confounding factor in the classification of NC vs PDAC (P=0. 47 and 0. 96) • Jaundice is not a confounding factor (P=0. 21) Fig A 1.
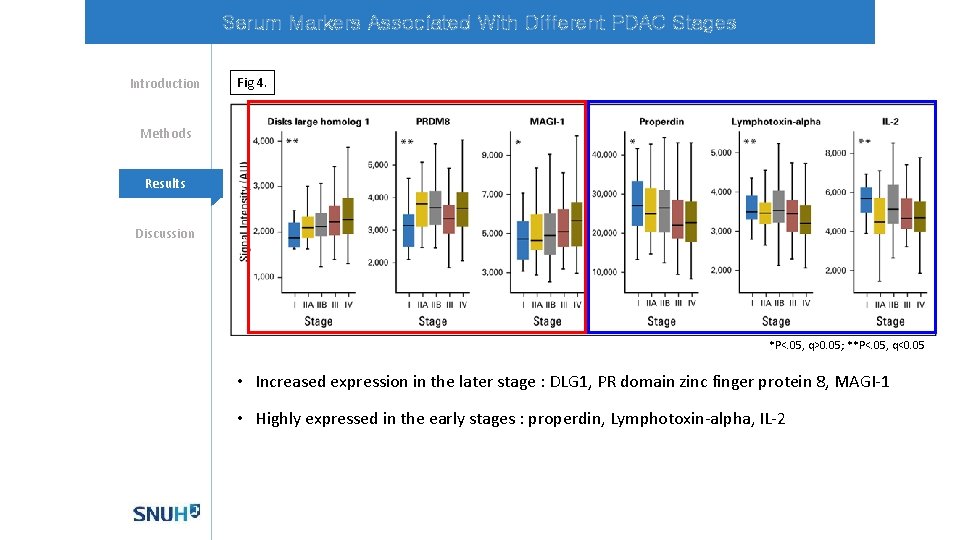
Serum Markers Associated With Different PDAC Stages Introduction Fig 4. Methods Mechanisms Results Discussion *P<. 05, q>0. 05; **P<. 05, q<0. 05 • Increased expression in the later stage : DLG 1, PR domain zinc finger protein 8, MAGI-1 • Highly expressed in the early stages : properdin, Lymphotoxin-alpha, IL-2
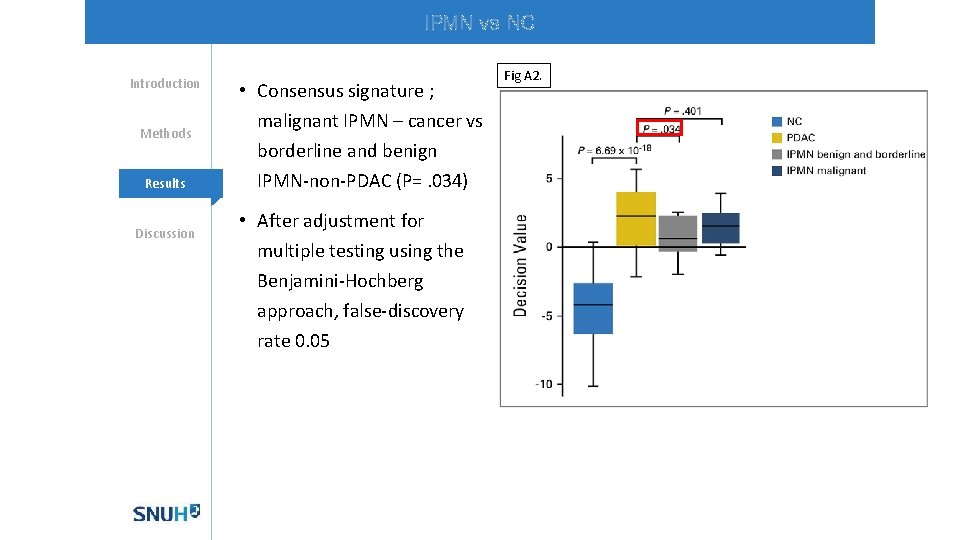
IPMN vs NC Introduction Methods Mechanisms Results Discussion • Consensus signature ; malignant IPMN – cancer vs borderline and benign IPMN-non-PDAC (P=. 034) • After adjustment for multiple testing using the Benjamini-Hochberg approach, false-discovery rate 0. 05 Fig A 2.

Proteomic analysis in surveillance of PDAC Introduction Methods Results Discussion • Proteomic multiparametric analysis could discriminate patients with early-stage I and II PDAC from controls • If supported by prospective validation studies, it would be beneficial in the surveillance of (1) high-risk patients (hereditary PDAC, CP, and peutz-jeghers syndrome) (2) patients with late-onset diabetes (3) patients with vague abdominal symptoms • Prevalance affects both PPV and NPV: • Higher risk of PDAC (eg. 1 -degree relatives) [prevalence 3. 75%]; PPV/NPV 0. 75/0. 99 • NOD older than 55 years [prevalence 1. 0%]; PPV/NPV 0. 46/1. 0 • CA 19 -9 was not included in biomarker signatures, because it did not contribute enough orthogonal information

Subgroup analysis Introduction Methods • No difference in decision value between PDAC with DM and PDAC without DM • The validated biomarker signature could potentially contribute to clinically rule out PDAC in patients with DM Results • Differential diagnosis of PDAC vs CP ; Discussion • Previous studies • Late stage PDAC could be distinguished from different pancreatic inflammatory indications • Different pancreatitis subtypes (acute, chronic, autoimmune) could be identified and distinguished from PDAC • In this study, CP could be discriminated from early-stage I and II PDAC, ROC-AUC 0. 84 • The consensus signature could discriminate Benign and borderline IPMN from stage I and II PDAC • Limitation is a fairly low number of clinical samples (n=20)

Interpretation of different expression of biomarkers Introduction Methods Results Discussion • DLG 1 interacts with APC, β-catenin, PTEN to regulate cell proliferation, cytokinesis, migration, and adhesion. • MAGI-1 both inhibit apoptosis and stimulate cell proliferation in HPV-induced malignancy. • PR domain zinc finger protein 8(BLIMP-1), DNA-binding protein, regulates neural and steroidrelated transcription and is a regulator of tumorigenesis in pituitary adenomas, where it most likely contributes to increased tumor invasiveness • Lymphotoxin-alpha is produced by type 1 T helper cells to induce phagocyte binding to endothelial cells. Some polymorphisms of this protein contribute to an increased risk of developing adenocarcinoma. • Properdin, positive complement regulator, not only does inhibition of complement activation typically promote cancer cell immune evasion, it has also been shown to hamper the efficacy of cancer immunotherapy. • IL-2 treatment in combination with conventional therapy can attenuate pancreatic cancer progression

Limitations Introduction Methods Results Discussion • Case-control studies ; Prospective validation studies need for clinical surveillance or therapeutic setting. • With known disease status, the signature has limited role in prediagnostic status. • Samples were collected at diagnosis, we cannot predict the performance of the signature after surgical removal of the tumor. • ct. DNA, not yet shown evidence of clinical usefulness for the detection of early-stage cancer ; a low sensitivity and the requirement of large volumes of plasma. • Protein-based approaches ; high sensitivity and microliter sample volumes ; although well-designed, validation studies are required.
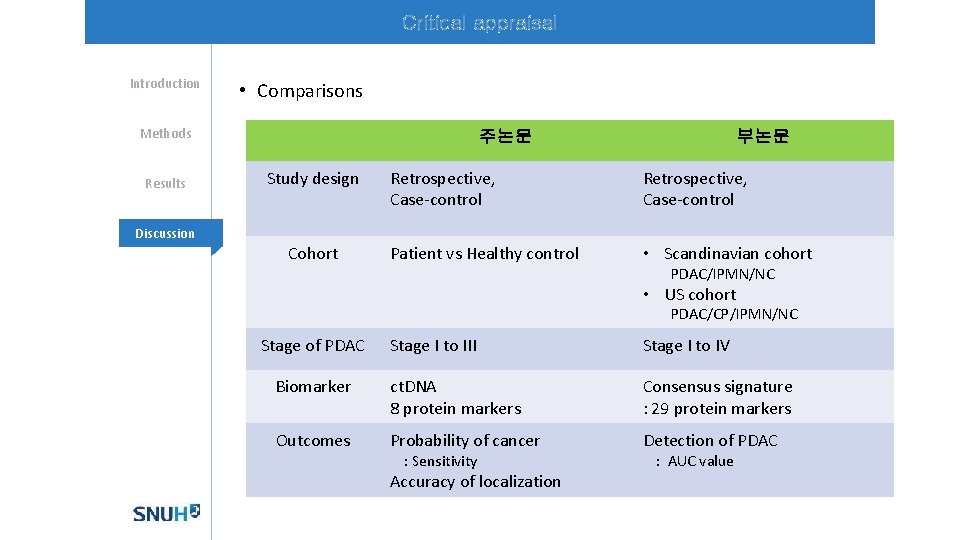
Critical appraisal Introduction • Comparisons Methods Results 주논문 Study design 부논문 Retrospective, Case-control Patient vs Healthy control • Scandinavian cohort Discussion Cohort PDAC/IPMN/NC • US cohort PDAC/CP/IPMN/NC Stage of PDAC Stage I to III Stage I to IV Biomarker ct. DNA 8 protein markers Consensus signature : 29 protein markers Outcomes Probability of cancer Detection of PDAC : Sensitivity Accuracy of localization : AUC value

Critical appraisal Introduction Methods • Retrospective study : • Cancer patients were already diagnosed by their symptoms • Asymptomatic patients? Prediagnostic or surveillance? Results Discussion • Other confounding factor? • ex) CA 19 -9 ; obstructive jaundice • Apolipoprotein A 1 (major component of HDL particles) ; hyperlipidemia (statin) • Complement C 3, C 4, C 5 ; antiphospholipid syndrome, mixed cryoglobulinemia, Sjögren syndrome, and membranoproliferative glomerulonephritis • Vascular endothelial growth factor(VEGF) ; rheumatoid arthritis, age-related macular degeneration

Critical appraisal Introduction Methods • Further study • Prospective study? RCT? • High risk group of PDAC Results Discussion • hereditary PDAC, peutz-jeghers syndrome, FAP. . . • late-onset DM, chronic pancreatitis, Pancreatic cyst • Environment : Cigarette smoking, Obesity, physical inactivity. . . • Liquid biopsy • Ct. DNA • Protein • mi. RNA, Tumor-educated platelets, Exosome. . . • Surveillance interval

- Slides: 38