Linkage Mapping Part 2 PBG 430 Making Linkage

Linkage Mapping – Part 2 PBG 430

Making Linkage Maps § Linkage analysis is now an "automated" procedure! § Data can be obtained on thousands and thousands of loci § The likelihood odds score (LOD) is a test statistic that is used to test the hypothesis that there is no linkage § LOD of 3. 00 is approximately equal to P ~ 0. 001 § LOD > 3, then one concludes that the two loci are indeed linked § The goal is to accurately determine locus order and distance § A linkage map is a genetic map – a reference for many useful analyses

Oregon Wolfe Barley Population § Doubled haploids made from the F 1 of OWB-D and OWB-R § International, publicly available resource OSU double haploid lab Dr. Robert Wolfe OWB-R and OWB-D (parents)

Why chose the Oregon Wolfe Barley Population for linkage mapping? § Many polymorphisms § Many markers available: morphological, isozyme, RFLP, SSR, AFLP, DAr. T, SNP Chutimanitsakun et al. (2011)

Adding new markers into an existing map § Be sure to check: § Number of linkage groups should remain the same (equal to n) § Order and distance between old markers should remain the same § Length (c. M) of linkage groups should remain the same (maybe shorter but never expand) Existing markers New markers added Chutimanitsakun et al. (2011)
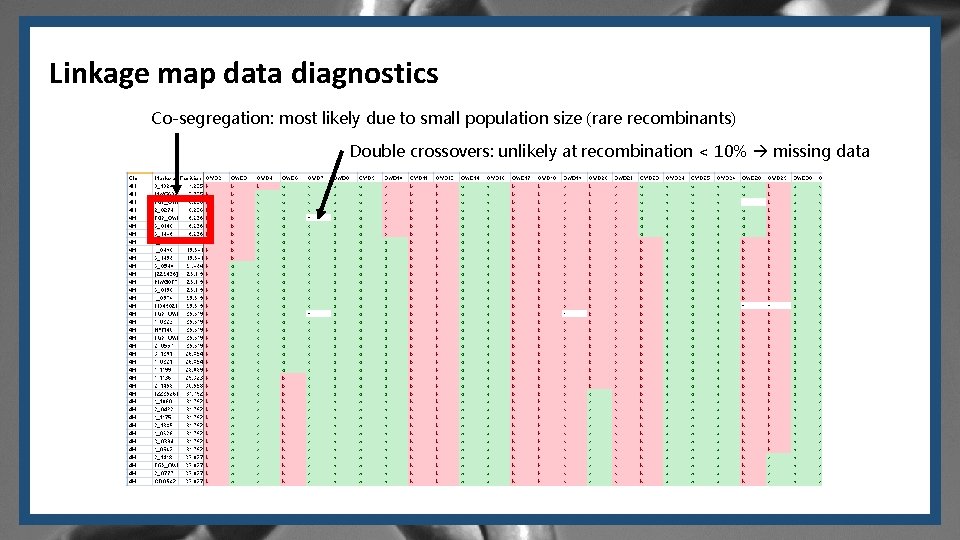
Linkage map data diagnostics Co-segregation: most likely due to small population size (rare recombinants) Double crossovers: unlikely at recombination < 10% missing data
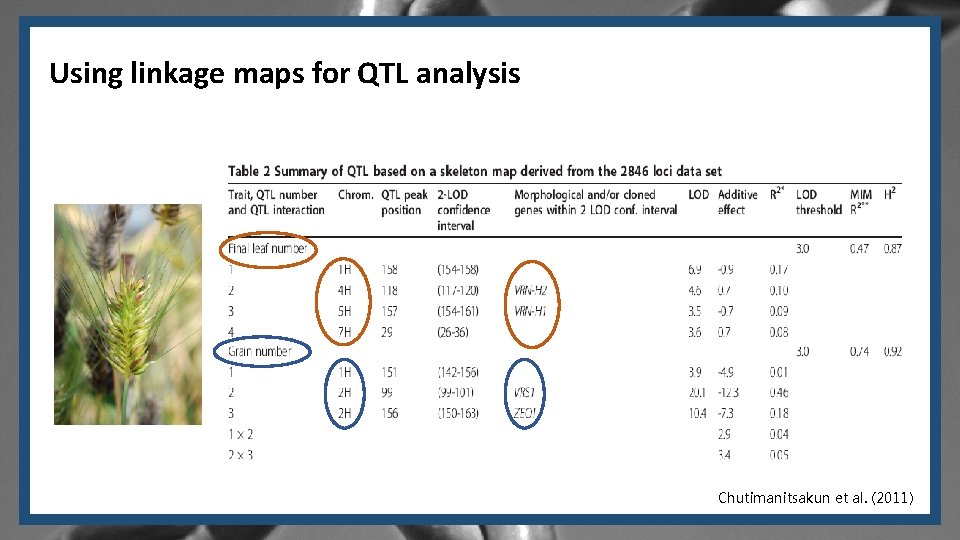
Using linkage maps for QTL analysis Chutimanitsakun et al. (2011)
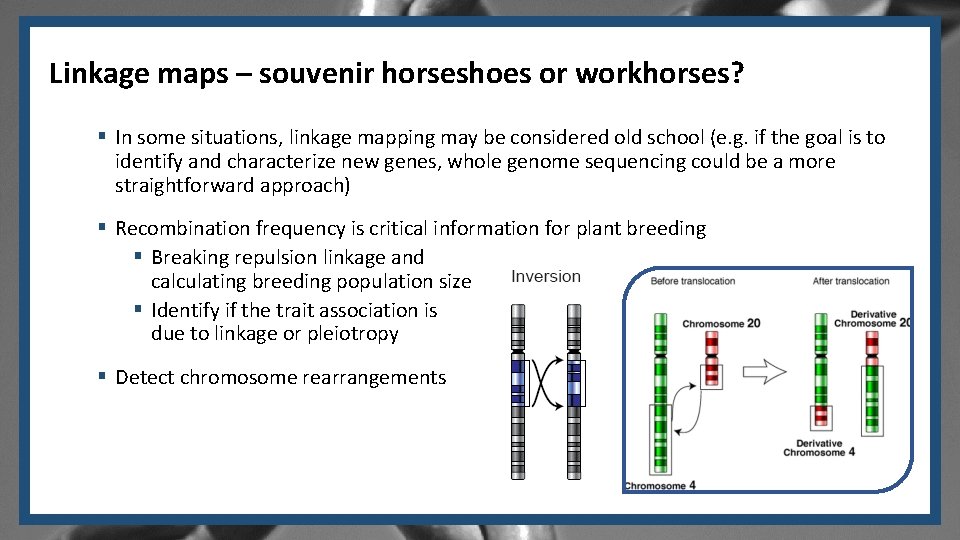
Linkage maps – souvenir horseshoes or workhorses? § In some situations, linkage mapping may be considered old school (e. g. if the goal is to identify and characterize new genes, whole genome sequencing could be a more straightforward approach) § Recombination frequency is critical information for plant breeding § Breaking repulsion linkage and calculating breeding population size § Identify if the trait association is due to linkage or pleiotropy § Detect chromosome rearrangements

By now, you should be able to… § Explain why the OWB population is a good one to use to build a linkage map. § Base your explanation on the recombination frequency formula. § Looking at a linkage map built by adding new markers into an existing map, identify if there any quality issues related to the number of linkage groups, the order and distance between loci, and the length of linkage groups. § Interpret a linkage map dataset (like the one shown in slide #6). § What are two possible reasons for co-segregation? § Why were double crossovers treated as missing data points? § Defend the statement: “With the availability of genome sequencing, linkage maps may have become an unnecessary step in certain situations. However, there are still many situations in which linkage maps are necessary. ”
- Slides: 9