Linkage Disequilibrium and Association Mapping Issues Opportunities for

Linkage Disequilibrium and Association Mapping: Issues & Opportunities for the Triticeae Mark E. Sorrells and Flavio Breseghello Department of Plant Breeding & Genetics Cornell University

Overview • Part I: A Genetic Model for Association Mapping in Plant Breeding Populations • Part II: Comparison of Different Plant Breeding Materials for Association Mapping • Part III: Association Mapping of Kernel Size and Milling Quality in Soft Winter Wheat Cultivars

A Definition of Association Mapping “Association analysis, also known as LD mapping or association mapping, is a population-based survey used to identify trait-marker relationships based on linkage disequilibrium” (Flint-Garcia et al. 2003)

Association Mapping as a Plant Breeding Strategy: AM versus QTL Mapping • Association Mapping can be conducted directly on the breeding material, therefore: • Direct inference from research to breeding is possible • Phenotypic variation is observed for most traits of interest • Marker polymorphism is higher than in biparental populations • Routine evaluations provide phenotypic data • Association Mapping provides other useful information about: • Organization of genetic variation • Polymorphism across the genome

Association Mapping as a Plant Breeding Strategy: AM versus QTL Mapping • Type I error (false positives) can be higher because of: • Unaccounted population structure • Simultaneous selection of combinations of alleles at different loci • High sampling variance of rare alleles • Type II error can be higher (low power) because of: • Lower LD than in mapping populations • Unbalanced design due to differences in allele frequencies • Serious multiple-testing problem
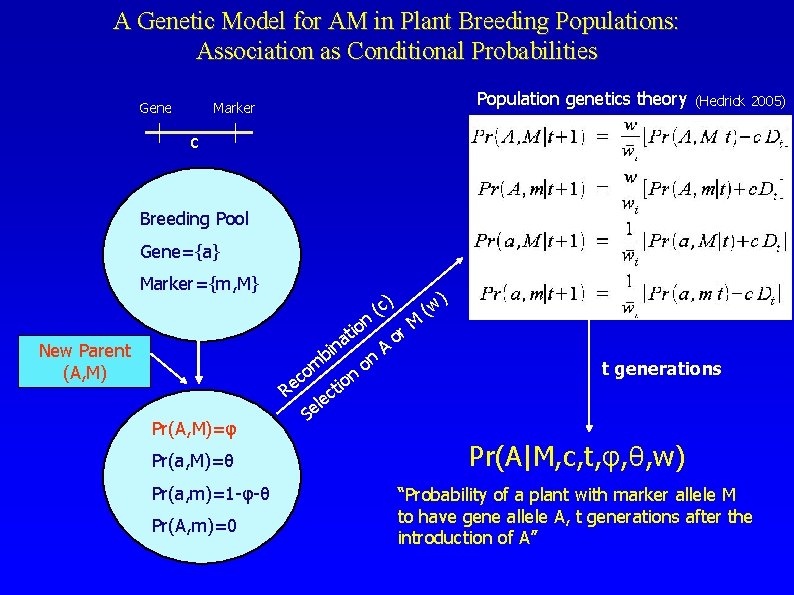
A Genetic Model for AM in Plant Breeding Populations: Association as Conditional Probabilities Gene Population genetics theory Marker (Hedrick 2005) c Breeding Pool Gene={a} Marker={m, M} n tio New Parent (A, M) na i b m o ec R Pr(A, M)=φ Pr(a, M)=θ Pr(a, m)=1 -φ-θ Pr(A, m)=0 t c e l Se i on ) (c on A M r o ) w ( t generations Pr(A|M, c, t, φ, θ, w) “Probability of a plant with marker allele M to have gene allele A, t generations after the introduction of A”
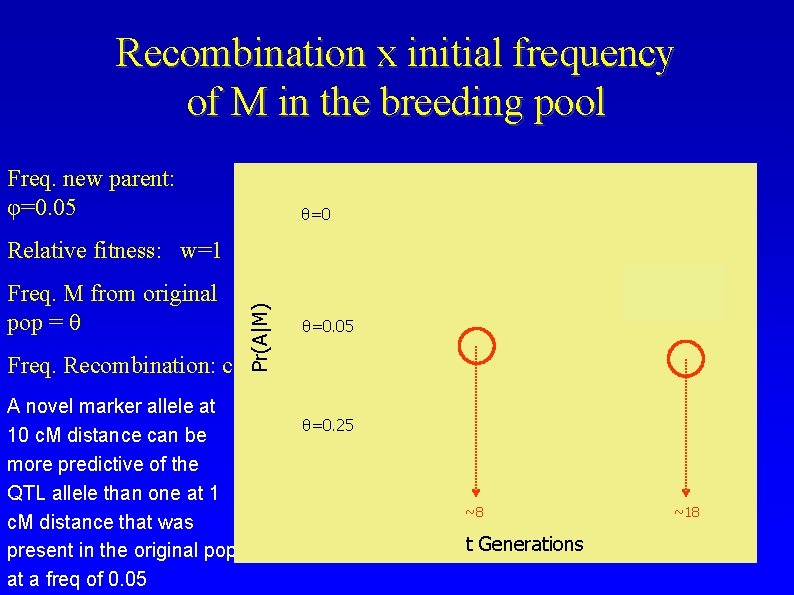
Recombination x initial frequency of M in the breeding pool Freq. new parent: φ=0. 05 θ=0 Freq. M from original pop = θ Freq. Recombination: c A novel marker allele at 10 c. M distance can be more predictive of the QTL allele than one at 1 c. M distance that was present in the original pop at a freq of 0. 05 Pr(A|M) Relative fitness: w=1 θ=0. 05 θ=0. 25 ~8 t Generations ~18
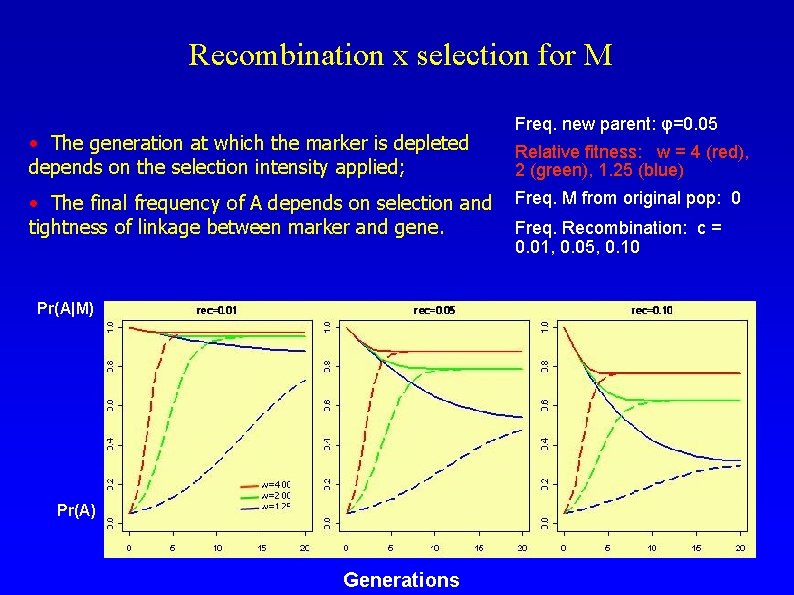
Recombination x selection for M • The generation at which the marker is depleted depends on the selection intensity applied; • The final frequency of A depends on selection and tightness of linkage between marker and gene. Pr(A|M) Pr(A) Generations Freq. new parent: φ=0. 05 Relative fitness: w = 4 (red), 2 (green), 1. 25 (blue) Freq. M from original pop: 0 Freq. Recombination: c = 0. 01, 0. 05, 0. 10

Summary Part I • In plant breeding populations, the locus most associated with the trait is not necessarily the closest locus; • Loosely linked markers can still be useful for MAS if high intensity of selection is applied.

Overview • Part I: A Genetic Model for Association Mapping in Plant Breeding Populations • Part II: Comparison of Different Plant Breeding Materials for Association Mapping • Part III: Association Mapping of Kernel Size and Milling Quality in Soft Winter Wheat Cultivars

Types of Populations • Germplasm Bank Collection • A collection of genetic resources including landraces, exotic material and wild relatives. • Synthetic Populations • Outcrossing populations (either male-sterile or manually crossed) synthesized from inbred lines. May be used for recurrent selection. • Elite Lines • Inbred lines (and checks) manipulated with the objective of releasing new varieties in the short term.

Characteristics Related to Association Mapping: Practical aspects Synthetic Populations Aspects of AM Germplasm bank Elite Germplasm Sample Core-collection Segregating progenies Elite lines and checks Sample turnover Static Ephemeral Gradually substituted Source of phenotypic data Screenings Progeny tests Yield trials Type of traits High heritability traits; Domestication traits Depends on the Low heritability traits: evaluation scheme yield, resistance to abiotic stresses Type of marker SNP SSR / SNP SSR

Characteristics Related to Association Mapping: Genetic Expectations Aspects of AM Germplasm bank Synthetic Populations Elite Germplasm Linkage Disequilibrium Low Intermediate and fast-decaying High Medium Low High Allele diversity among samples High Intermediate Low Allele diversity within samples Variable 1 or 2 alleles (diploid species) 1 allele (inbred lines) Population structure

Characteristics Related to Association Mapping: Potential Applications Aspects Germplasm bank Synthetic Populations Elite Germplasm Power Low Intermediate and decreasing High; could allow genome scan Resolution High; could allow fine Intermediate and mapping increasing Low Use of significant markers Transfer of new alleles Incorporation in by marker-assisted selection index backcross MAS in progenies (requires validation)

Summary Part II • Germplasm bank core-collections could be useful for allele-mining of candidate genes and fine-mapped QTLs; • Elite lines could be useful to detect genomic regions associated with traits of interest; • Synthetic populations might represent a balance between power and precision, and have the major advantage of being unstructured.

Overview • Part I: A Genetic Model for Association Mapping in Plant Breeding Populations • Part II: Comparison of Different Plant Breeding Materials for Association Mapping • Part III: Association Mapping of Kernel Size and Milling Quality in Soft Winter Wheat Cultivars
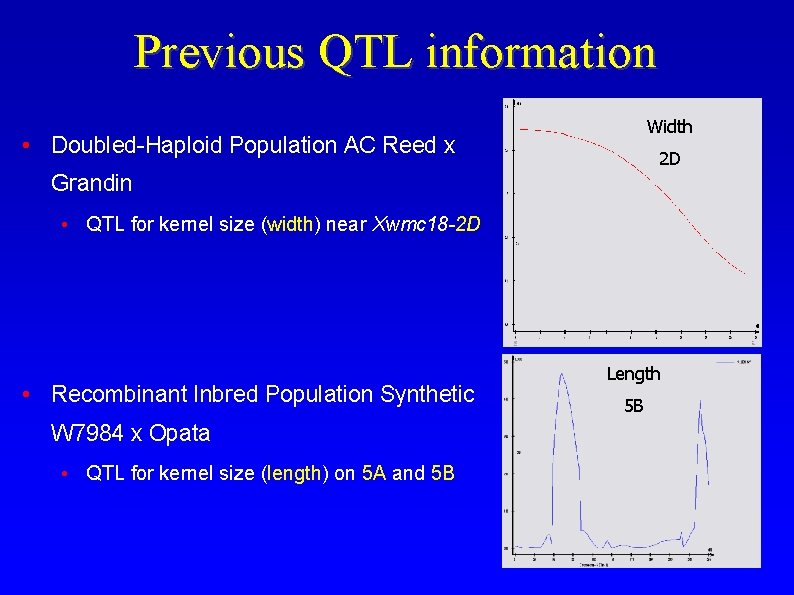
Previous QTL information Width • Doubled-Haploid Population AC Reed x 2 D Grandin • QTL for kernel size (width) near Xwmc 18 -2 D • Recombinant Inbred Population Synthetic W 7984 x Opata • QTL for kernel size (length) on 5 A and 5 B Length 5 B

Plant Material • 95 cultivars of soft winter wheat from the Northeast of USA • Mostly recent releases: 92>1990; 39>2000 • Representing 35 seed companies / institutions • selected from 149 cultivars based on 18 unlinked SSR markers

Genotypic Data • Marker distribution: 93 SSR loci • 33 on chromosome 2 D • 20 on chromosome 5 A • 9 on chromosome 5 B • 31 on 16 other chromosomes • Data trimming • rare alleles (freq<5%) were pooled with missing data, and • considered as missing for LD and population structure analysis • considered as allele for AM analysis

Methods: Population Structure • Data: 36 “unlinked” SSR markers • Program: Structure (Pritchard et al. , 2000, Genetics 155: 945) • Model: without admixture (cultivars discretely assigned to subpopulations) • Validated subpopulations: Resampled subsets of 12, 18, 24 and 30 unlinked loci • Visualization: Factorial Correspondence Analysis (Benzecri, 1973 L' Analyse des correspondances. Dunod)

Methods: Linkage Disequilibrium • Statistics: r 2 , with p-values from 1000 permutations • Program: Tassel (maizegenetics. net) • LD among linked loci: • Scan of entire chromosome 2 D • Scan of pericentromeric region of chromosome 5 A • LD among unlinked loci: • Computed among 36 unlinked loci

Methods: Association Mapping • Statistical Model: Linear mixed-effects model • marker as fixed effects • subpopulations as random effects • Program: R package lme (Pinheiro & Bates, 2000 Mixed-Effects Models in S and S-PLUS. Springer) • Multiple testing correction: 1000 permutations chromosome-wise • Two-marker models: tested by likelihood ratio test

Population Structure: Sample Subdivisions Subpopulation No. of Varieties Fst 1 19 0. 337 2 32 0. 111 3 13 0. 295 4 31 0. 064 Total 95 0. 188 Moderate Population Subdivision
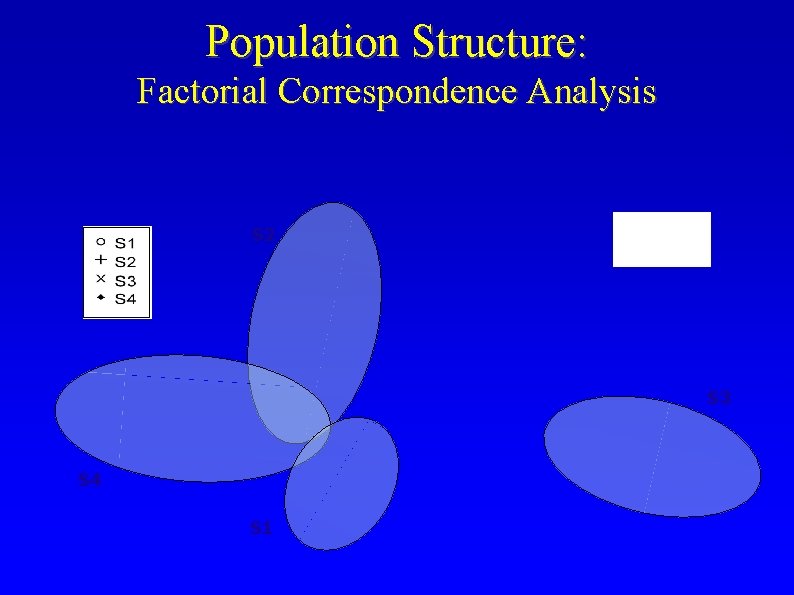
Population Structure: Factorial Correspondence Analysis S 2 S 3 S 4 S 1
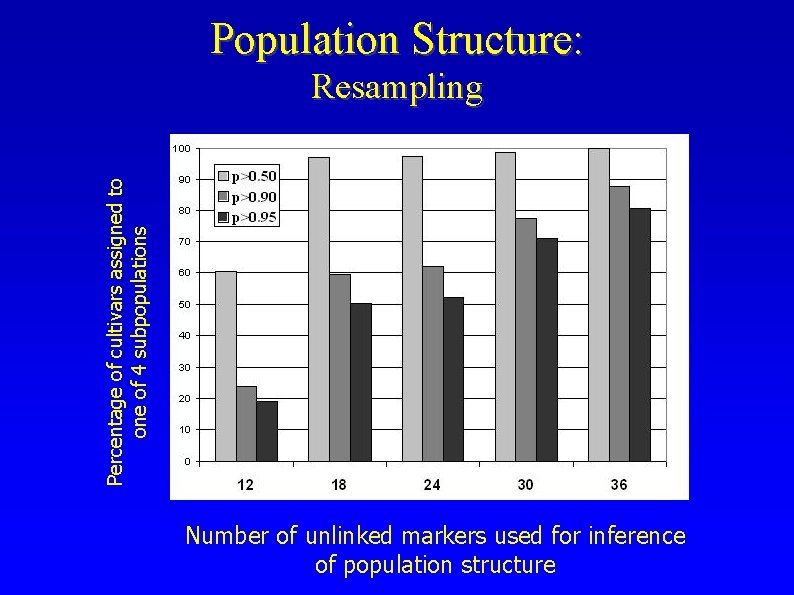
Population Structure: Percentage of cultivars assigned to one of 4 subpopulations Resampling Number of unlinked markers used for inference of population structure
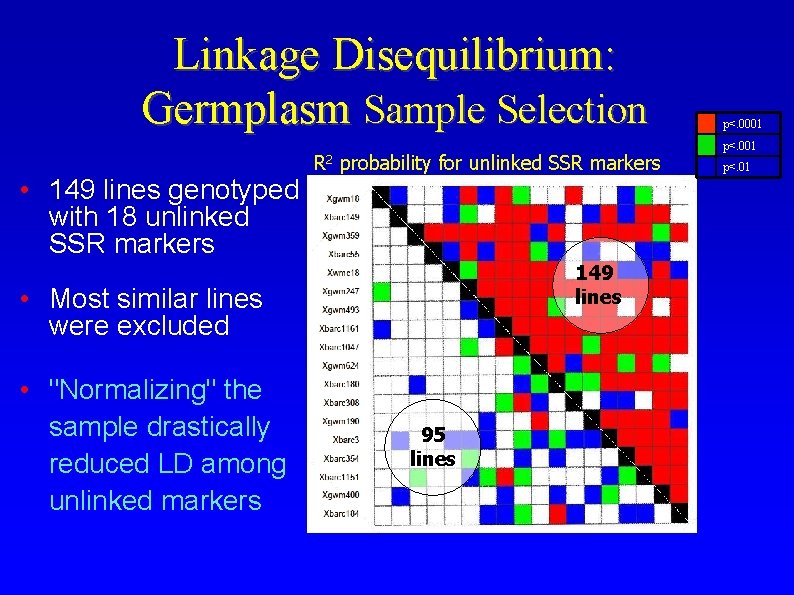
Linkage Disequilibrium: Germplasm Sample Selection • 149 lines genotyped with 18 unlinked SSR markers R 2 probability for unlinked SSR markers 149 lines • Most similar lines were excluded • "Normalizing" the sample drastically reduced LD among unlinked markers 95 lines p<. 0001 p<. 01

Definition of a baseline-LD specific for our sample Defined as the 95 th percentile of the distribution of r 2 among unlinked loci r 2 estimates above this value are probably due to genetic linkage Baseline LD for this sample: r 2 = 0. 0654 Normal curve LD baseline Normal Distr. 95 th percentile

Linkage Disequilibrium: Chromosome 2 D Consistent LD was below 1 c. M

Linkage Disequilibrium: Chromosome 5 A LD extended for 5 c. M ~5 c. M
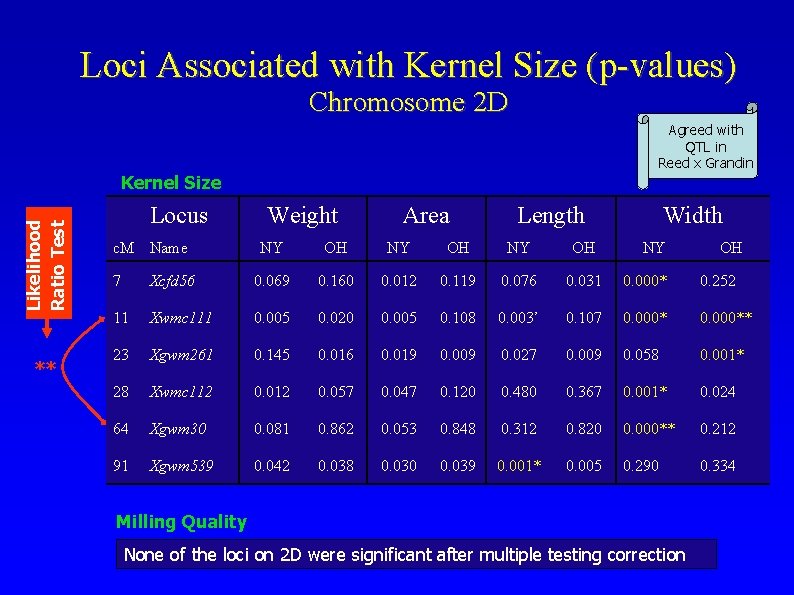
Loci Associated with Kernel Size (p-values) Chromosome 2 D Agreed with QTL in Reed x Grandin Likelihood Ratio Test Kernel Size ** Locus c. M Name Weight Area Length NY OH Width NY OH 7 Xcfd 56 0. 069 0. 160 0. 012 0. 119 0. 076 0. 031 0. 000* 0. 252 11 Xwmc 111 0. 005 0. 020 0. 005 0. 108 0. 003’ 0. 107 0. 000** 23 Xgwm 261 0. 145 0. 016 0. 019 0. 009 0. 027 0. 009 0. 058 0. 001* 28 Xwmc 112 0. 057 0. 047 0. 120 0. 480 0. 367 0. 001* 0. 024 64 Xgwm 30 0. 081 0. 862 0. 053 0. 848 0. 312 0. 820 0. 000** 0. 212 91 Xgwm 539 0. 042 0. 038 0. 030 0. 039 0. 001* 0. 005 0. 290 0. 334 Milling Quality None of the loci on 2 D were significant after multiple testing correction
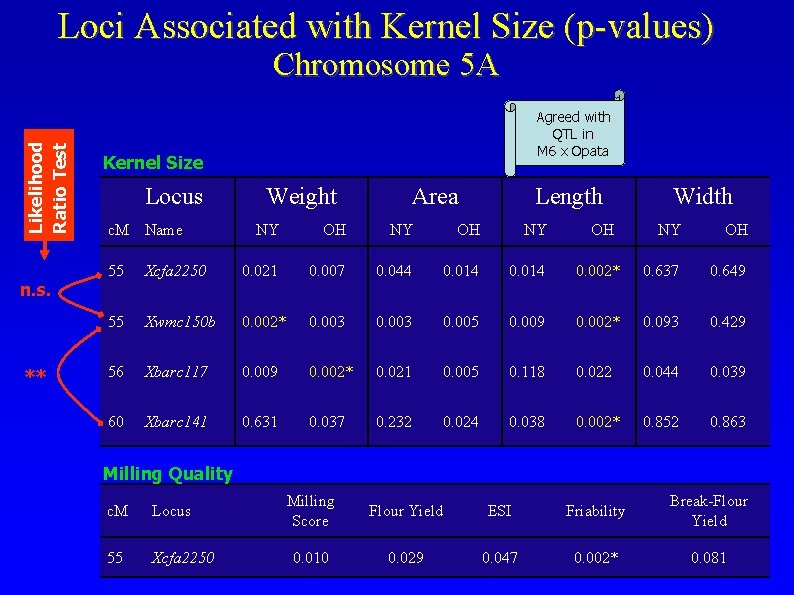
Loci Associated with Kernel Size (p-values) Likelihood Ratio Test Chromosome 5 A n. s. ** Agreed with QTL in M 6 x Opata Kernel Size Locus c. M Name Weight Area Length NY OH Width NY OH 55 Xcfa 2250 0. 021 0. 007 0. 044 0. 014 0. 002* 0. 637 0. 649 55 Xwmc 150 b 0. 002* 0. 003 0. 005 0. 009 0. 002* 0. 093 0. 429 56 Xbarc 117 0. 009 0. 002* 0. 021 0. 005 0. 118 0. 022 0. 044 0. 039 60 Xbarc 141 0. 631 0. 037 0. 232 0. 024 0. 038 0. 002* 0. 852 0. 863 Milling Quality c. M Locus 55 Xcfa 2250 Milling Score Flour Yield ESI Friability Break-Flour Yield 0. 010 0. 029 0. 047 0. 002* 0. 081
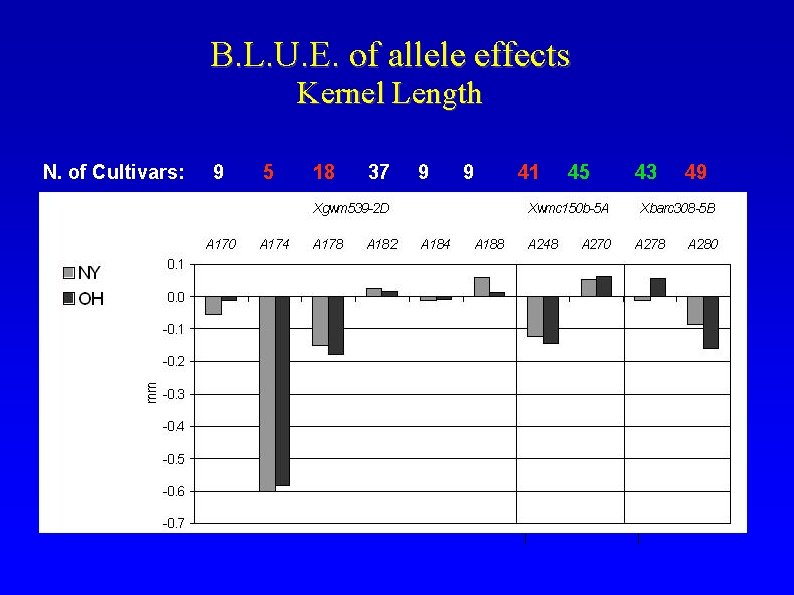
B. L. U. E. of allele effects Kernel Length N. of Cultivars: 9 5 18 37 9 9 41 45 43 49
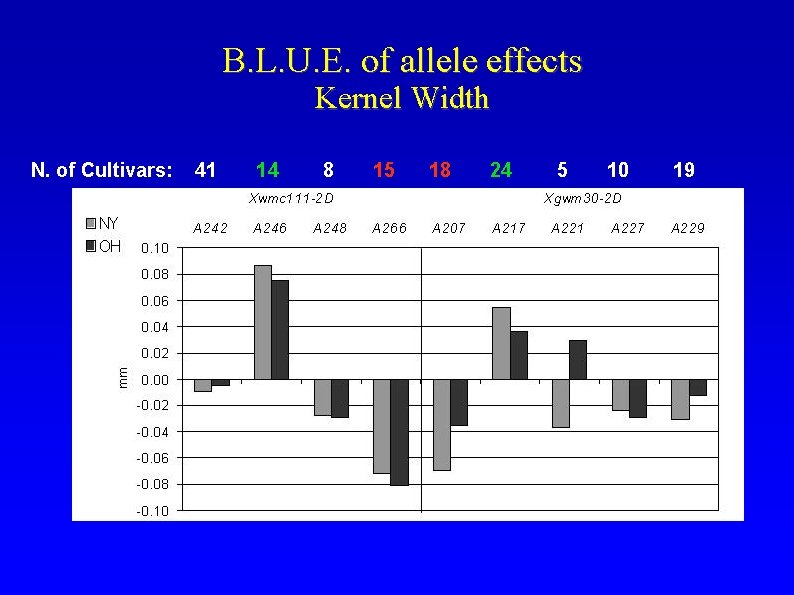
B. L. U. E. of allele effects Kernel Width N. of Cultivars: 41 14 8 15 18 24 5 10 19
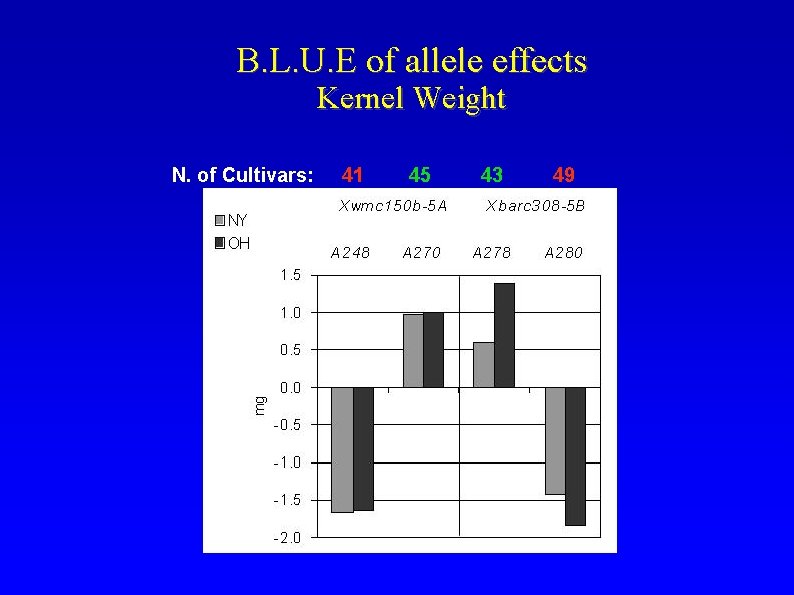
B. L. U. E of allele effects Kernel Weight N. of Cultivars: 41 45 43 49

Summary Part III • Linkage Disequilibrium • LD on chromosome 2 D was in the subcentimorgan scale • LD on chromosome 5 A extended for 5 c. M, forming an LD block • Association Mapping • Loci on chromosome 2 D were associated with kernel width • Loci on chromosome 5 A were associated with kernel length and friability • Favorable and unfavorable marker alleles were identified • In recurrent selection, markers could be used to carry information from a “good year” to a “bad year” • In pedigree breeding, markers could carry information about yield potential from the phase of replicated field trials to the phase of singleplant selection

Acknowledgements • USDA Soft Wheat Quality Lab, Wooster, OH • Embrapa • Technical Support: • David Benscher • James Tanaka • Gretchen Salm

Cornell Small Grains Breeding & Genetics Project James Tanaka Dani Grechen Satwayan Salm Mike Gifford Rob Elshire Jesse Munkvold David Benscher Abigail Losh
- Slides: 37