Linear Sequence Alignment Travis Hillenbrand 1 Methods of

Linear Sequence Alignment Travis Hillenbrand 1

Methods of Comparison l Dot Matrix l Dynamic Programming Algorithm l Greedy X-drop Approach l Linear Alignment 2

Dot Matrix Method 3 http: //arbl. cvmbs. colostate. edu/molkit/dnadot/index. html

Sequence Alignment ATCGATACG, ATGGATTACG 3 possibilities Match …C… | …C… Mismatch …C… …G… Indel …C… …-… 4

Global Pairwise Alignment ATCGATACG, ATGGATTACG Matches: Mismatches: Gaps: ATCGAT-ACG || ||| ATGGATTACG +1 +1 +1 -1 -2 +1 +1 = +8 = -1 = -2 Total score = +5 5

Dynamic Programming Global alignment (Needleman-Wunsch) algorithm 6

Dynamic Programming Global alignment (Needleman-Wunsch) algorithm 7

Dynamic Programming Global alignment (Needleman-Wunsch) algorithm 8

Dynamic Programming Global alignment (Needleman-Wunsch) algorithm +1 Max= 1 9

Dynamic Programming Global alignment (Needleman-Wunsch) algorithm 10

Dynamic Programming Global alignment (Needleman-Wunsch algorithm) GATC || | GA-C 11

Greedy X-drop Alignment l Aligns sequences that differ by sequencing errors l Works with measure of difference l Restricts indel penalty 12 Zhang et al. 2000
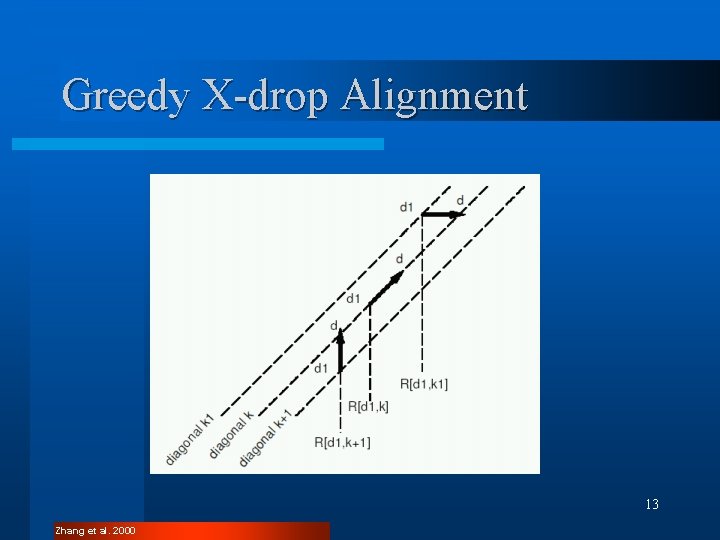
Greedy X-drop Alignment 13 Zhang et al. 2000

Greedy X-drop Alignment 14

Greedy X-drop Alignment l X-drop condition saves computation 15
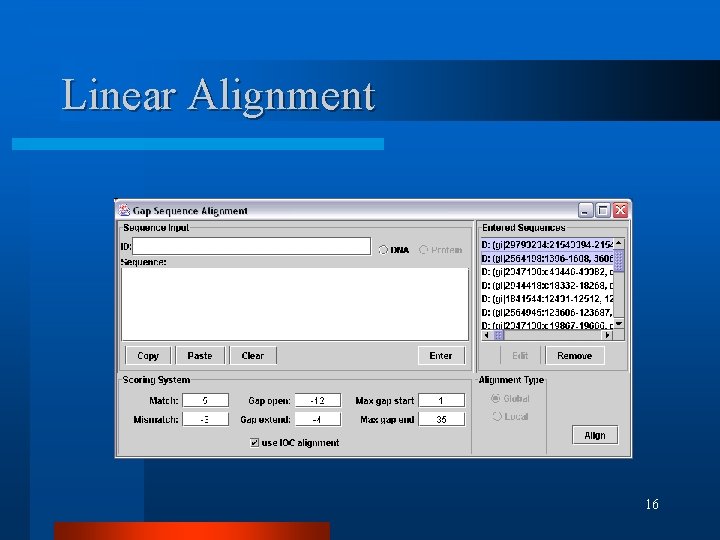
Linear Alignment 16

Linear Alignment l Index of coincidence – Maximum number of matches between two sequences – Ungapped alignment ATCGATACG ATGGATTACG ATCGATACG | ATGGATTACG … ATCGATACG ATGGATTACG ATCGATACG || ||| ATGGATTACG 17

Linear Alignment l Attempt ATCGATACG || ||| ATGGATTACG Window score: 2 to increase similarity -ATCGATACG |||| ATGGATTACG -3 ATCGATACG | -ATGGATTACG -3 ATCGATACG || ||| ATGGATTACG 18

Comparison of alignments l 9 human/mouse homologous gene cds pairs retrieved (Jareborg et al. 1999) l Greedy alignment run first mat=10, mis=-6, X=2200 (indel=-11) l Dynamic Programming and Linear alignment using truncated seqs 19
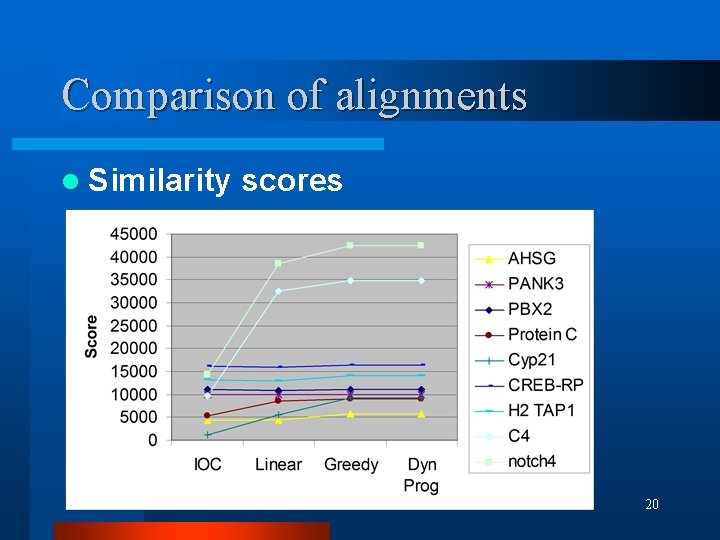
Comparison of alignments l Similarity scores 20
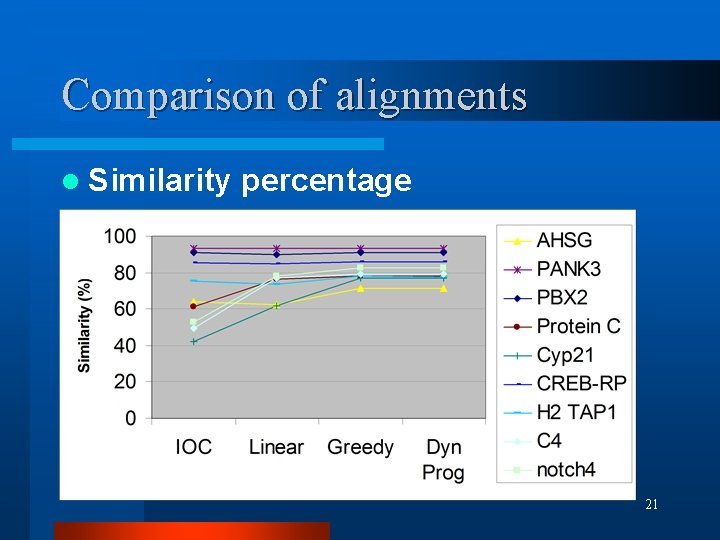
Comparison of alignments l Similarity percentage 21
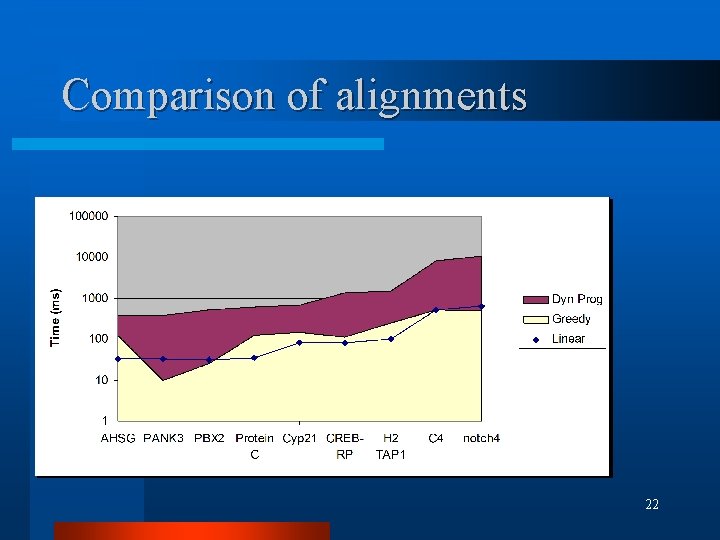
Comparison of alignments 22
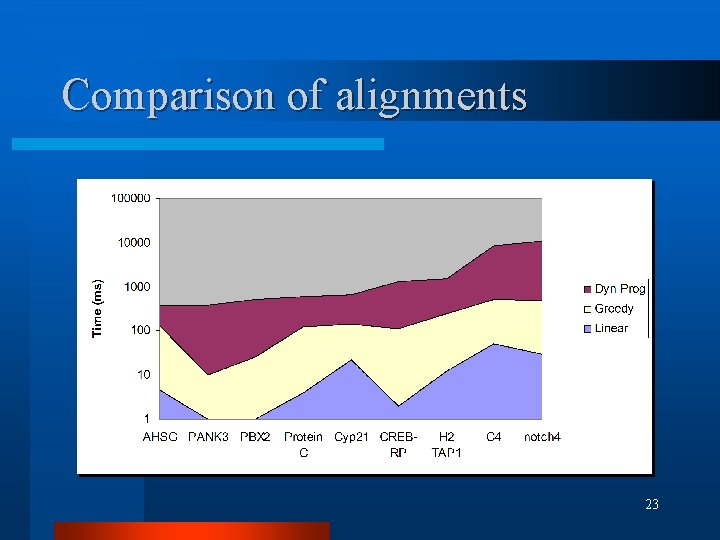
Comparison of alignments 23
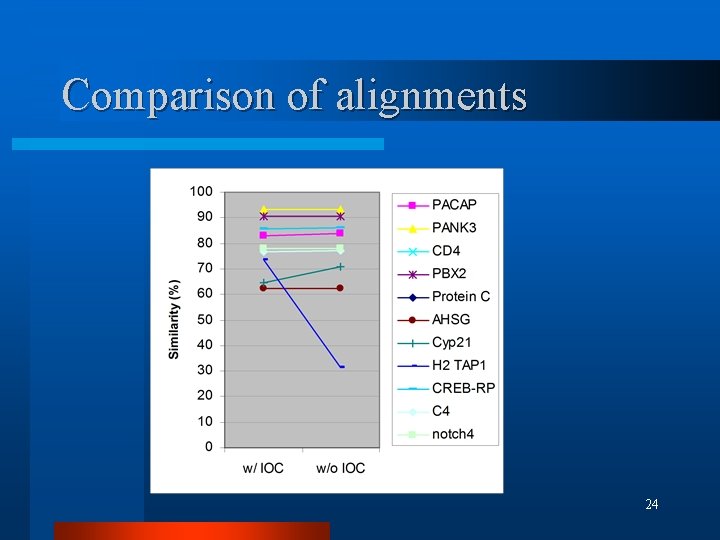
Comparison of alignments 24

Comparison of alignments Maximum coincidence alignment: Offset -72 yielded 1642 matches of 2175 possible (75. 4943% similarity), score 6611 ACAGTACTGCTACTTCTCGCCGACTGGGTGCTGCTCCGGACCGCGCTGCCCCGCATATTCTCCCTGCTGGTGCCCACCGCGCTGCCACTGCTCCGGGT | | | | ||||||| | | ||| | ATGGCTGCGCACGTCTGGCGGCCGCCCTGCTCCTTCTGGTGGACTGGCTGCTGCTGCGGCCCATGCTCCCGGGAATCTTCTCCCTGTTGGTTCC ACGGGCCGCCTCACTGGATTCTACAAGATGGCTCAGCCGATACCTTCACTCGAAACTTAACTCTCATGTCCATTCTCACCATAGCCAGTGCAGT |||||||||||||||| || |||||| || ||||||||||||| ||| ACGGGCCGCATCACTGGATTCTTCAGGATAAGACAGTTCCTAGCTTCACCCGCAACATATGGCTCATGTCCATTCTCACCATAGCCAGCACAGC Decreasing the gap penalty allows similar regions to be aligned without using IOC 25

Comparison of alignments References Needleman, S. B. & Wunsch, C. D. 1970. A general method applicable to the search for similarities in the amino acid sequence of two proteins. Journal of Molecular Biology. 48: 443 -453. Setubal, J. and Meidanis, J. 1997. Introduction to Computational Molecular Biology. Pacific Grove, California: Brooks/Cole. Zhang, Z. ; Schwartz, S. ; Wagner, L. ; and Miller, W. 2000. A greedy algorithm for aligning DNA sequences. Journal of Computational Biology 7: 203 -214. 26

Linear Sequence Alignment Travis Hillenbrand 27
- Slides: 27