Ligands dictionary and refinement Garib N Murshudov York

Ligands, dictionary and refinement Garib N Murshudov York Structural Biology Laboratory University of York

Outline 1. 2. 3. 4. 5. 6. Introduction Dictionary of ligands Sources of dictionary and idealised coordinates Tools for ligand description in ccp 4 How to use dictionary in refinement (REFMAC) Conclusions

The need for prior chemical knowledge • Refinement • Atomic model description Graphics Simulations ………. .

Atomic model description Default pointers in PDB file. . . ATOM 7 ATOM 8 ATOM 9 ATOM 10 ATOM 11. . . C O N CA CB LEU ILE ILE A A A 5 5 6 6 6 37. 584 36. 548 37. 887 37. 032 37. 835 Pointer to link description Pointer to monomer description Pointer to atom description 4. 085 3. 447 5. 098 5. 447 6. 276

Refmac 5 Dictionary • • • Describes all amino acids All nucleic acids Common sugars Many organic and inorganic compounds Links and modifications There are tools to deal with dictionary Dictionary format is mm. CIF


General category data_comp_list loop_ _chem_comp. id _chem_comp. three_letter_code _chem_comp. name _chem_comp. group _chem_comp. number_atoms_all _chem_comp. number_atoms_nh _chem_comp. desc_level. . GLC-b-D GLC 'beta_D_glucose ' D-pyranose Group: peptide, DNA/RNA, pyranose, non-polymer Level: C or M – complete or minimal description 24 12 .

Atom category loop_ _chem_comp_atom. comp_id _chem_comp_atom_id _chem_comp_atom. type_symbol _chem_comp_atom. type_energy _chem_comp_atom. partial_charge _chem_comp_atom. x _chem_comp_atom. y _chem_comp_atom. z GLC-b-D C 1 C CH 1 0 GLC-b-D H 1 H HCH 1 0. . . 0. 0 0. 522 -0. 087 0. 801

Bond category loop_ _chem_comp_bond. comp_id _chem_comp_bond. atom_id_1 _chem_comp_bond. atom_id_2 _chem_comp_bond. type _chem_comp_bond. value_dist_esd GLC-b-D O 1 C 1 single GLC-b-D C 2 C 1 single. . . 1. 410 1. 524 Type: single, double, triple, aromatic, metal 0. 020

Angle category loop_ _chem_comp_angle. comp_id _chem_comp_angle. atom_id_1 _chem_comp_angle. atom_id_2 _chem_comp_angle. atom_id_3 _chem_comp_angle. value_angle_esd GLC-b-D H 1 C 1 GLC-b-D O 1 C 1. . . O 1 C 2 109. 470 3. 000

Torsion angles category loop_ _chem_comp_tor. comp_id _chem_comp_tor. atom_id_1 _chem_comp_tor. atom_id_2 _chem_comp_tor. atom_id_3 _chem_comp_tor. atom_id_4 _chem_comp_tor. value_angle_esd _chem_comp_tor. period GLC-b-D var_1 C 2 GLC-b-D var_2 C 1 C 2. . . O 2 C 3 Period: number of energetic minima HO 2 0. 000 C 4 -50. 095 1 20. 000 3 4 2 3
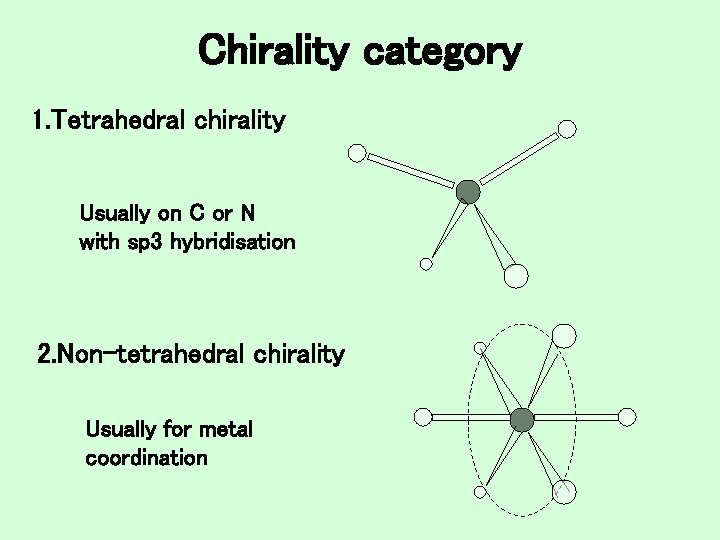
Chirality category 1. Tetrahedral chirality Usually on C or N with sp 3 hybridisation 2. Non-tetrahedral chirality Usually for metal coordination

Chirality category loop_ _chem_comp_chir. comp_id _chem_comp_chir. atom_id_centre _chem_comp_chir. atom_id_1 _chem_comp_chir. atom_id_2 _chem_comp_chir. atom_id_3 _chem_comp_chir. volume_sign GLC-b-D chir_01 C 5 C 4 GLC-b-D chir_02 C 4 C 3 GLC-b-D chir_03 C 2 GLC-b-D chir_04 C 2 C 1. . . O 5 O 4 O 3 O 2 C 6 C 5 C 4 C 3 positive negative positive 1 C Sign: positive, negative, both, anomer _ 3 +

Metal chirality is only used to create coordinates loop_ _chem_comp_chir. comp_id _chem_comp_chir. atom_id_centre _chem_comp_chir. atom_id_1 _chem_comp_chir. atom_id_2. . _chem_comp_chir. atom_id_8 _chem_comp_chir. volume_sign MONid chir_id Ac Ab Af A 1 A 2 A 3 A 4 A 5 A 6 cross 6 Where: Ac - chiral centre atom Ab - back atom, Af - forward atom A 1, A 2, . . . , AN - atoms in the same plane, N can be = 0, 1, 2, 3, 4, 5, 6 these atoms form the point group. cross. N - cross chirality specification
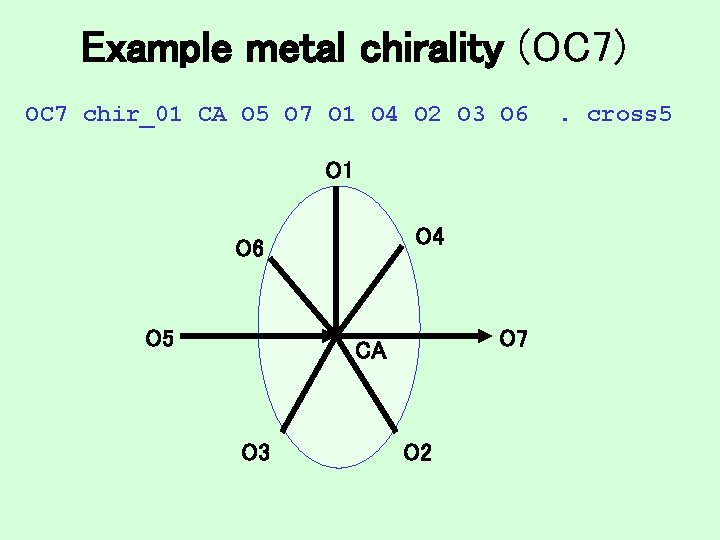
Example metal chirality (OC 7) OC 7 chir_01 CA O 5 O 7 O 1 O 4 O 2 O 3 O 6 O 1 O 4 O 6 O 5 O 7 CA O 3 O 2 . cross 5

Plane category loop_ _chem_comp_plane_atom. comp_id _chem_comp_plane_atom. plane_id _chem_comp_plane_atom_id _chem_comp_plane_atom. dist_esd PHE plan CB PHE plan CG PHE plan CD 1. . . 0. 020

Example of a modification Modification formalism allows to change a monomer Modification describes in details the result of chemical reaction

Modification: general category data_mod_list loop_ _chem_mod. id _chem_mod. name _chem_mod. comp_id _chem_mod. group_id. . . O 1 MET O 1_metyl_of_sugar group_id: means only for sugars . pyranose

Modification: atom category loop_ _chem_mod_atom. mod_id _chem_mod_atom. function _chem_mod_atom_id _chem_mod_atom. new_type_symbol _chem_mod_atom. new_type_energy _chem_mod_atom. new_partial_charge O 1 MET change O 1 MET delete HO 1. O 1 MET add. CM O 1 MET add. HM 1. . . . C H function: only - change, delete or add O 2. CH 3 HCH 0. 000

Modification: bond category loop_ _chem_mod_bond. mod_id _chem_mod_bond. function _chem_mod_bond. atom_id_1 _chem_mod_bond. atom_id_2 _chem_mod_bond. new_type _chem_mod_bond. new_value_dist_esd O 1 MET add O 1 CM O 1 MET add CM HM 1 O 1 MET add CM HM 2 O 1 MET add CM HM 3 single 1. 420 0. 960 0. 020

Example of peptide link Link formalism allows to join monomers together Link describes in details the result of chemical reaction

Link: general category data_link_list loop_ _chem_link. id _chem_link. comp_id_1 _chem_link. mod_id_1 _chem_link. group_comp_1 _chem_link. comp_id_2 _chem_link. mod_id_2 _chem_link. group_comp_2 _chem_link. name ALPHA 1 -4. DEL-HO 4 pyranose. DEL-O 1 glycosidic_bond_alpha 1 -4 pyranose mod_id _1: modification of first monomer before the linkage mod_id_2 : modification of second monomer before the linkage

Link: bond category loop_ _chem_link_bond. link_id _chem_link_bond. atom_1_comp_id _chem_link_bond. atom_id_1 _chem_link_bond. atom_2_comp_id _chem_link_bond. atom_id_2 _chem_link_bond. type _chem_link_bond. value_dist_esd ALPHA 1 -4 1 O 4 2 C 1 single 1. 439 atom_1_comp_id: means first monomer atom_2_comp_id: means second monomer 0. 020

Source of dictionary and coordinates • • • MSDchem PRODRG RELIBASE CORINA QM or other energy minimsation programs CSD
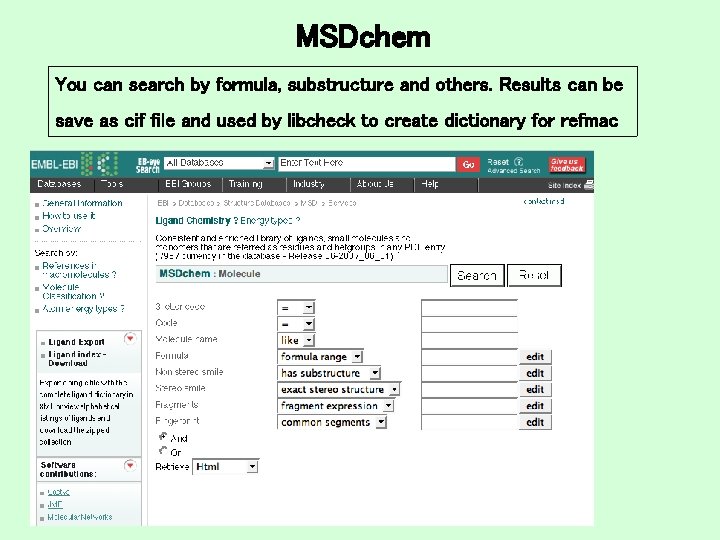
MSDchem You can search by formula, substructure and others. Results can be save as cif file and used by libcheck to create dictionary for refmac

MSDchem: JME 1) Draw substructure, write a smile file or load SDF, MOL, mm. CIF, 2) Search
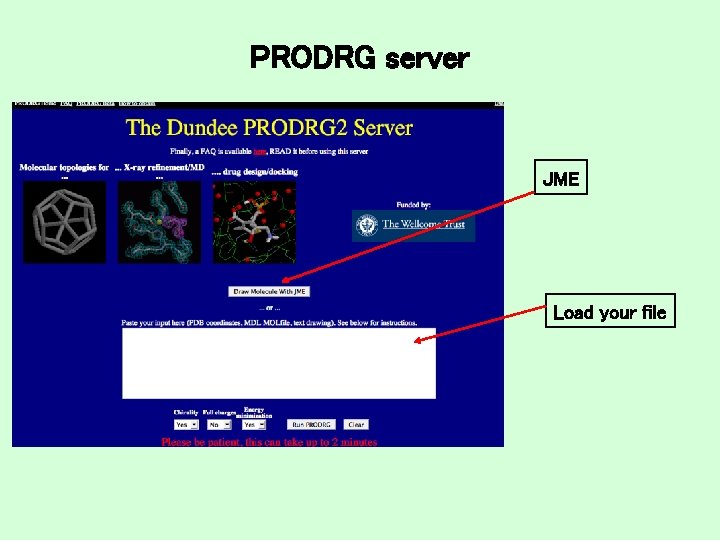
PRODRG server JME Load your file
PRODRG: JME Draw your ligand, transfer to PRODRG window and run

PRODRG output It can write out dictionaries for CNS REFMAC 5, SHELX and others

Tools in CCP 4 LIBCHECK - creates the complete monomer description from minimal - creates coordinates from complete monomer description SKETCHER - graphical program that creates the minimal monomer description for LIBCHECK MAKECIF - creates restraints

Ways to create dictionary 1. From chemical structure Using SKETCHER: monomer is drawn specifying atoms and bonds From SMILE strings, sdf file, mol 2 file 2. From Cartesian coordinates Coordinates from CSD Energetically optimised coordinates MOL 2 file SDF file
C(=O)O 3 D representation: For description of Smile strings: An example SMILE for ALA: N[C@@H](C)C(=O)O 3 D representation: For description of](http://slidetodoc.com/presentation_image_h/954640152019a2967024866089a24ccb/image-32.jpg)
Smile strings: An example SMILE for ALA: N[C@@H](C)C(=O)O 3 D representation: For description of smile: http: //www. daylight. com/dayhtml_tutorials/languages/smiles/index. html
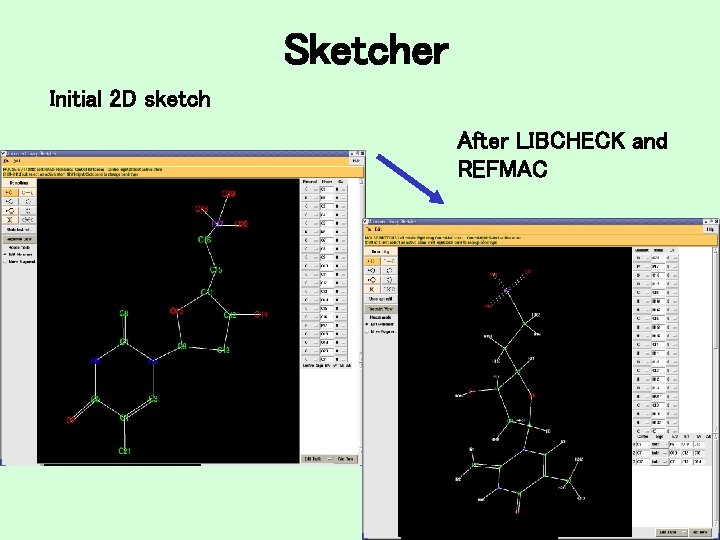
Sketcher Initial 2 D sketch After LIBCHECK and REFMAC

Restraints: monomer linkage 1. 2. 3. 4. 5. 6. Chain links (trans/cis, DNA/RNA, sugar links, gap) Standard links (SS bridges, sugar-protein links) Potential links Links between alternative conformations Symmetry links User links

Modifications and links in PDB file Link ID SSBOND LINK LINK 1 CYS L 88 SG CYS H TYR L GLY H MAG Y O LEU B OE 1 GLU A CYS L 23 195 2. 031 SG BCYS H 140 139 PRO L 140 127 GLY H 133 1 GAL Y 2 61 NA NA X 6 139 NA NA X 1 Standard name Name in PDB file MODRES GAL Y 2 GAL-b-D 12555 SS PCIS gap BETA 1 -4 LEU-NA symmetry Modification ID RENAME

Conclusions • Ligand dictionaries should designed with care. Interpetation of chemistry may depend on that • Such resources as MSDchem, PRODRG can help to create an accurate dictionary • Links and modifications are important component for understanding protein chemistry • Unfortunately no automatic link generation programs available yet (we are working on that)

Acknowledgments • • • Alexei Vagin Roberto Steiner Andrey Lebedev Liz Potterton Fei Long – YSBL, York – Kings coll. – YSBL, York • Wellcome Trust, BBSRC, BIOXHIT, CCP 4 – money
- Slides: 37