Ligand Building with ARPw ARP Automated Model Building
Ligand Building with ARP/w. ARP
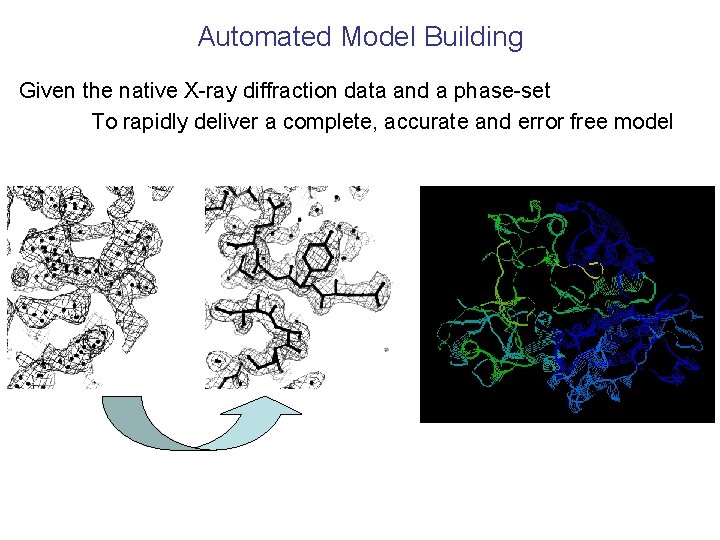
Automated Model Building Given the native X-ray diffraction data and a phase-set To rapidly deliver a complete, accurate and error free model
Building Ligands from Dummy Atoms / Seed Points Back to about 2000: a side project for a Ph. D student

Nearest Neighbour Distance Distribution Given a coordinate error, the inter-atomic distances in a protein model change:
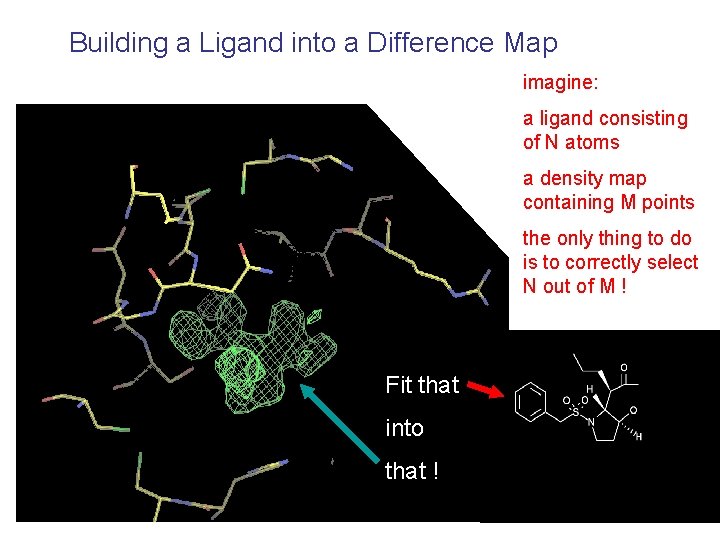
Building a Ligand into a Difference Map imagine: a ligand consisting of N atoms a density map containing M points the only thing to do is to correctly select N out of M ! Fit that into that !
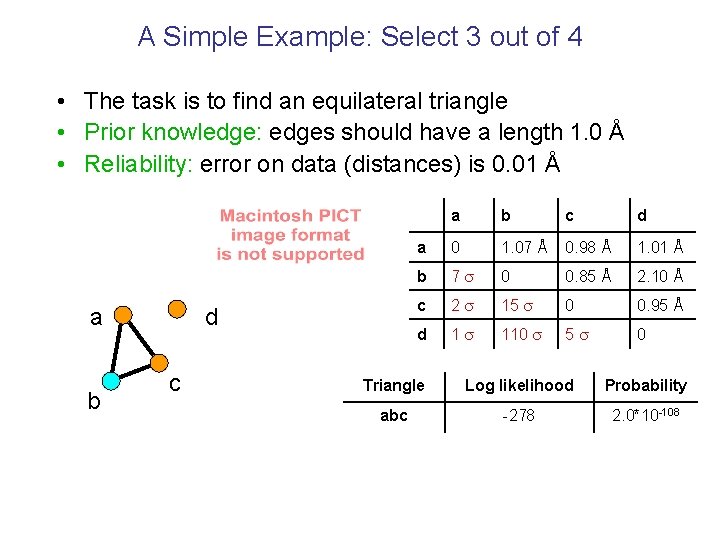
A Simple Example: Select 3 out of 4 • The task is to find an equilateral triangle • Prior knowledge: edges should have a length 1. 0 Å • Reliability: error on data (distances) is 0. 01 Å a b d c a b c d a 0 1. 07 Å 0. 98 Å 1. 01 Å b 7 0 0. 85 Å 2. 10 Å c 2 15 0 0. 95 Å d 1 110 5 0 Triangle Log likelihood Probability abc -278 2. 0*10 -108
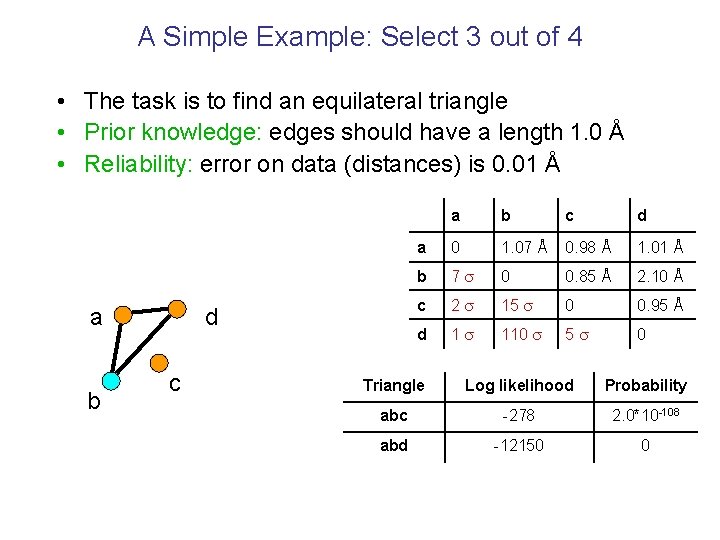
A Simple Example: Select 3 out of 4 • The task is to find an equilateral triangle • Prior knowledge: edges should have a length 1. 0 Å • Reliability: error on data (distances) is 0. 01 Å a b d c a b c d a 0 1. 07 Å 0. 98 Å 1. 01 Å b 7 0 0. 85 Å 2. 10 Å c 2 15 0 0. 95 Å d 1 110 5 0 Triangle Log likelihood Probability abc -278 2. 0*10 -108 abd -12150 0

A Simple Example: Select 3 out of 4 • The task is to find an equilateral triangle • Prior knowledge: edges should have a length 1. 0 Å • Reliability: error on data (distances) is 0. 01 Å a b d c a b c d a 0 1. 07 Å 0. 98 Å 1. 01 Å b 7 0 0. 85 Å 2. 10 Å c 2 15 0 0. 95 Å d 1 110 5 0 Triangle Log likelihood Probability abc -278 2. 0*10 -108 abd -12150 0 bcd -12350 0

A Simple Example: Select 3 out of 4 • The task is to find an equilateral triangle • Prior knowledge: edges should have a length 1. 0 Å • Reliability: error on data (distances) is 0. 01 Å a b d c a b c d a 0 1. 07 Å 0. 98 Å 1. 01 Å b 7 0 0. 85 Å 2. 10 Å c 2 15 0 0. 95 Å d 1 110 5 0 Triangle Log likelihood Probability abc -278 2. 0*10 -108 abd -12150 0 bcd -12350 0 acd -30 0. 9999
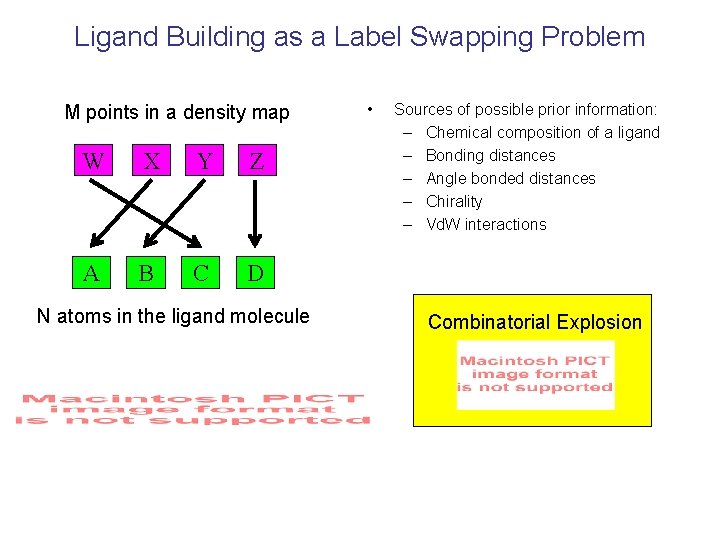
Ligand Building as a Label Swapping Problem M points in a density map W X Y Z A B C D N atoms in the ligand molecule • Sources of possible prior information: – Chemical composition of a ligand – Bonding distances – Angle bonded distances – Chirality – Vd. W interactions Combinatorial Explosion
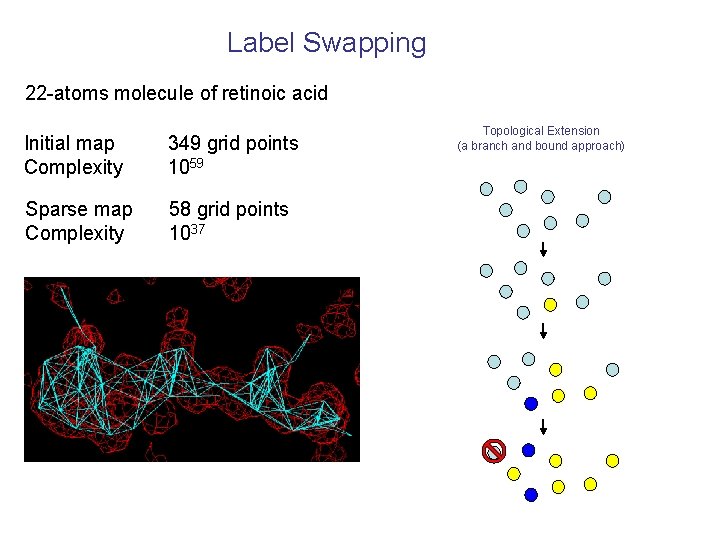
Label Swapping 22 -atoms molecule of retinoic acid Initial map Complexity 349 grid points 1059 Sparse map Complexity 58 grid points 1037 Topological Extension (a branch and bound approach)
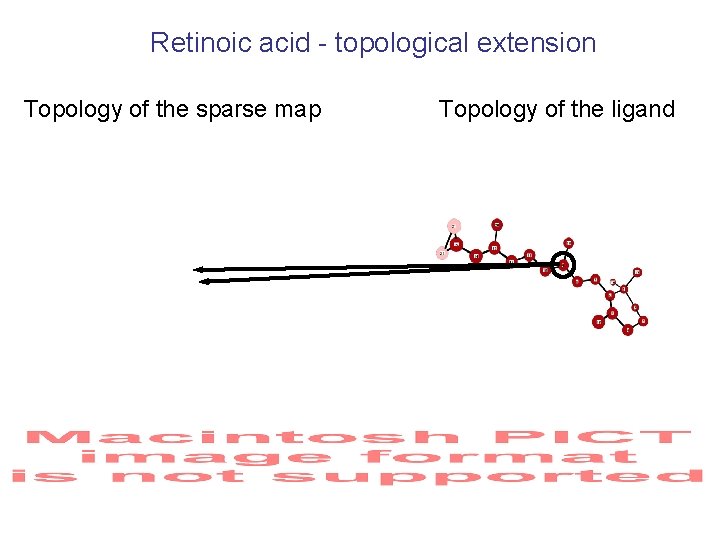
Retinoic acid - topological extension Topology of the sparse map Topology of the ligand

Real Space Fit for Final Selection of the Model 22 atoms molecule of retinoic acid: among 100 “top” models: 21 are less than 0. 5 Å r. m. s. d. from the final model the “best” model is 0. 14 Å r. m. s. d. from the final model
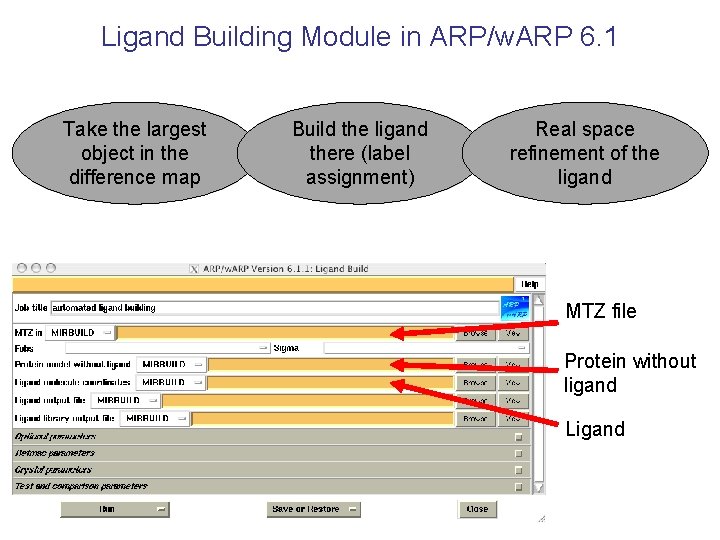
Ligand Building Module in ARP/w. ARP 6. 1 Take the largest object in the difference map Build the ligand there (label assignment) Real space refinement of the ligand MTZ file Protein without ligand Ligand

Ligand Building Module in ARP/w. ARP 6. 1 Location unknown Location known Single known ligand Yes (if the largest) No A ligand out of the list of expected ligands No No Partially ordered ligand No No
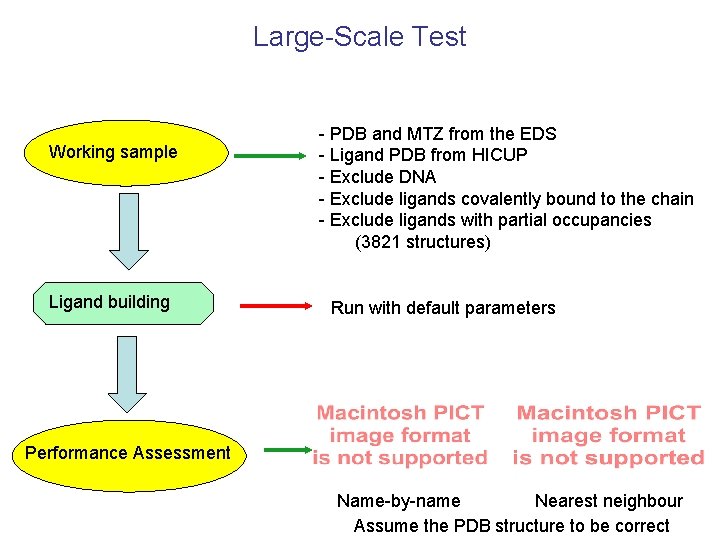
Large-Scale Test Working sample Ligand building - PDB and MTZ from the EDS - Ligand PDB from HICUP - Exclude DNA - Exclude ligands covalently bound to the chain - Exclude ligands with partial occupancies (3821 structures) Run with default parameters Performance Assessment Name-by-name Nearest neighbour Assume the PDB structure to be correct
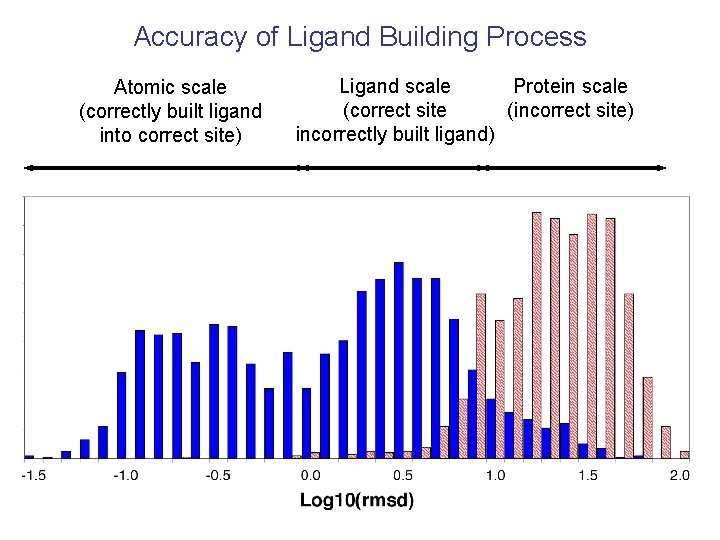
Accuracy of Ligand Building Process Atomic scale (correctly built ligand into correct site) Ligand scale Protein scale (correct site (incorrect site) incorrectly built ligand)
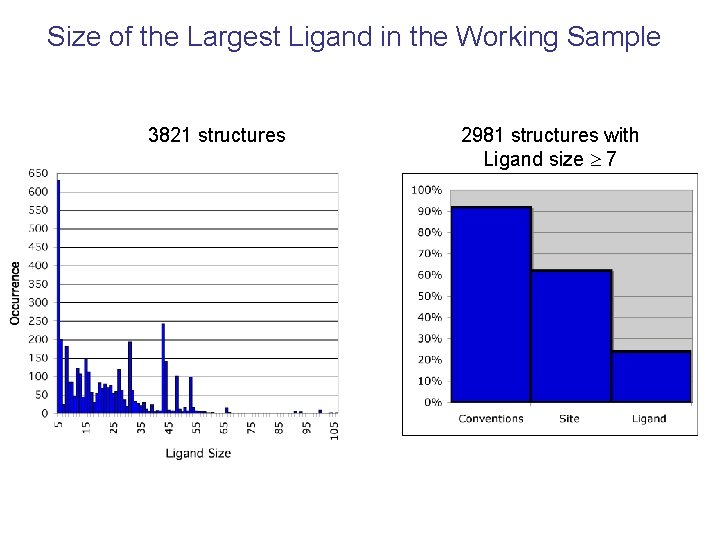
Size of the Largest Ligand in the Working Sample 3821 structures 2981 structures with Ligand size 7

Dependence on Resolution of the Data

Dependence on Ligand Disorder B factors
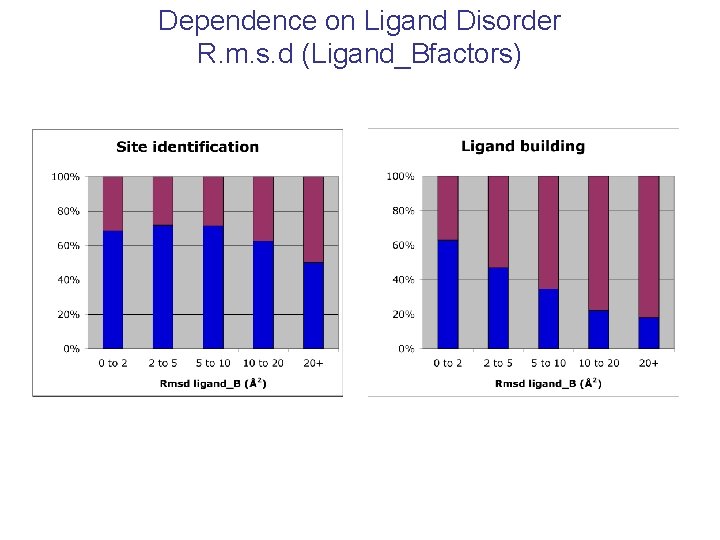
Dependence on Ligand Disorder R. m. s. d (Ligand_Bfactors)
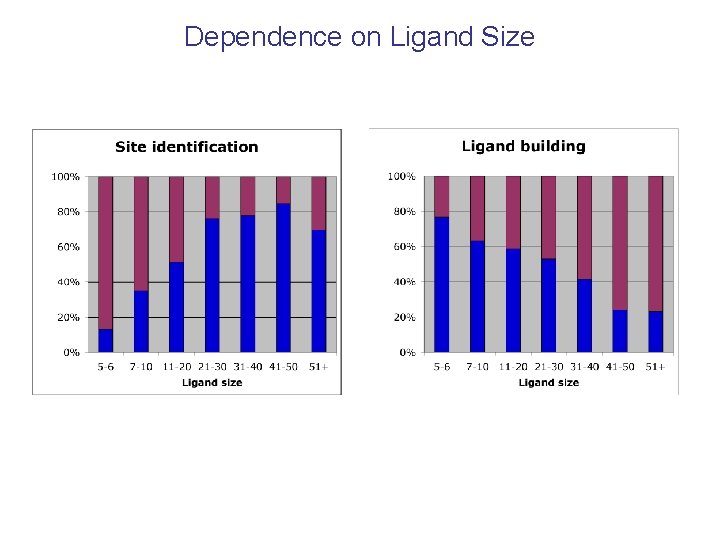
Dependence on Ligand Size
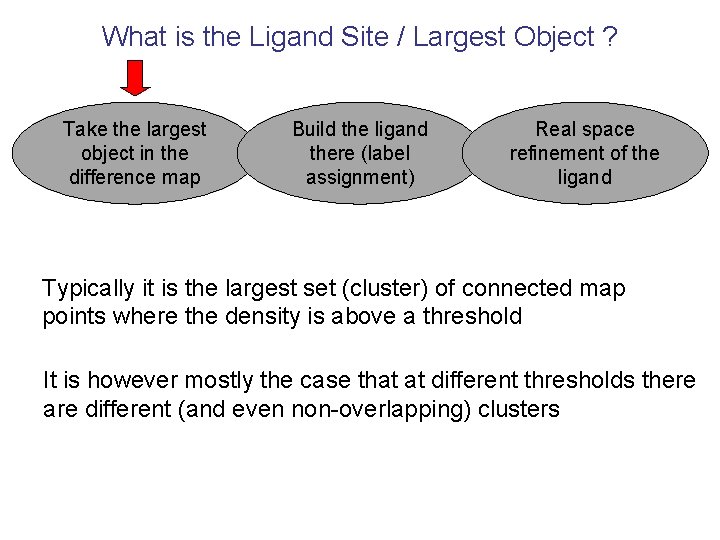
What is the Ligand Site / Largest Object ? Take the largest object in the difference map Build the ligand there (label assignment) Real space refinement of the ligand Typically it is the largest set (cluster) of connected map points where the density is above a threshold It is however mostly the case that at different thresholds there are different (and even non-overlapping) clusters

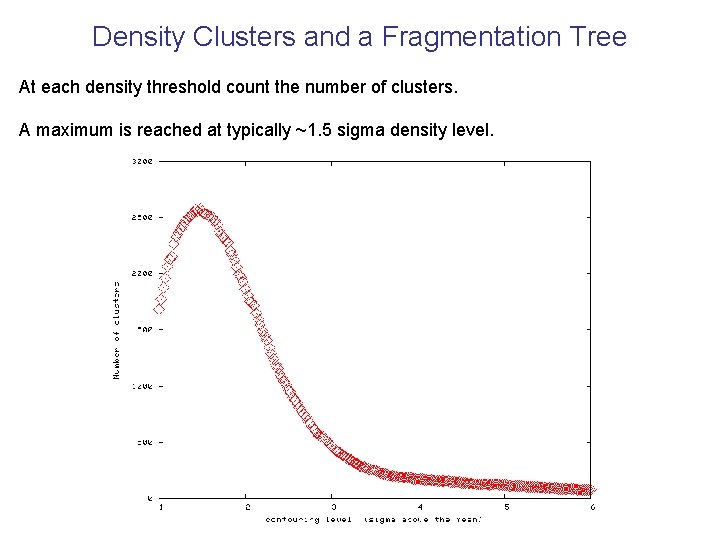
Density Clusters and a Fragmentation Tree At each density threshold count the number of clusters. A maximum is reached at typically ~1. 5 sigma density level.
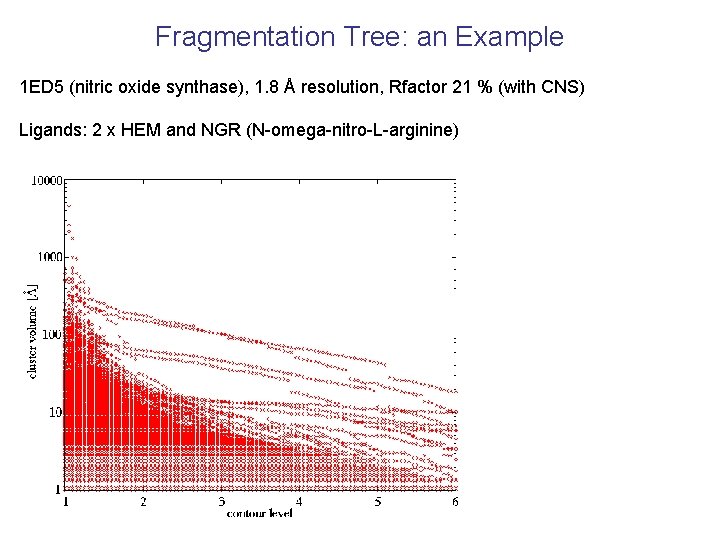
Fragmentation Tree: an Example 1 ED 5 (nitric oxide synthase), 1. 8 Å resolution, Rfactor 21 % (with CNS) Ligands: 2 x HEM and NGR (N-omega-nitro-L-arginine)
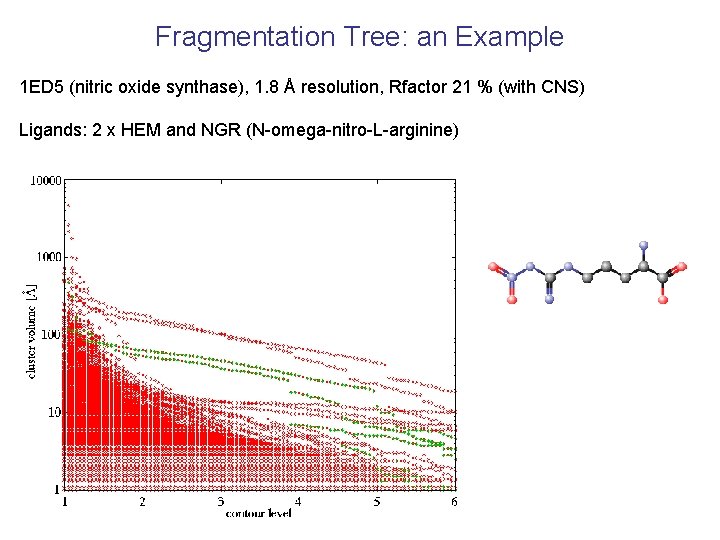
Fragmentation Tree: an Example 1 ED 5 (nitric oxide synthase), 1. 8 Å resolution, Rfactor 21 % (with CNS) Ligands: 2 x HEM and NGR (N-omega-nitro-L-arginine)
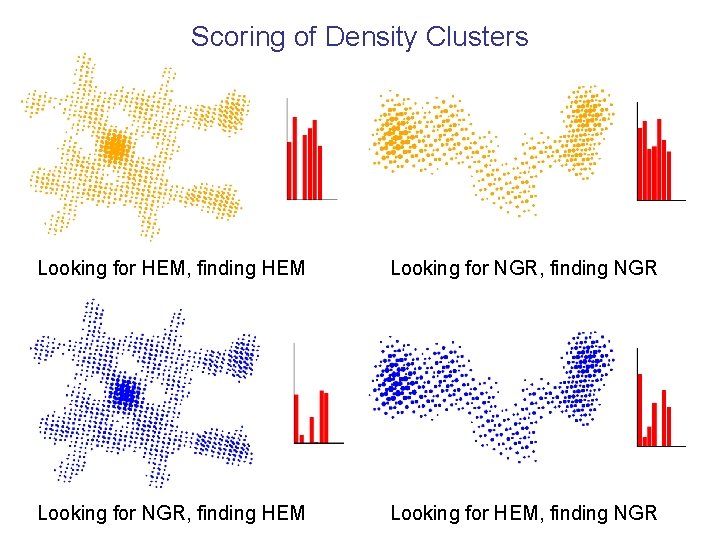
Scoring of Density Clusters Looking for HEM, finding HEM Looking for NGR, finding NGR Looking for NGR, finding HEM Looking for HEM, finding NGR
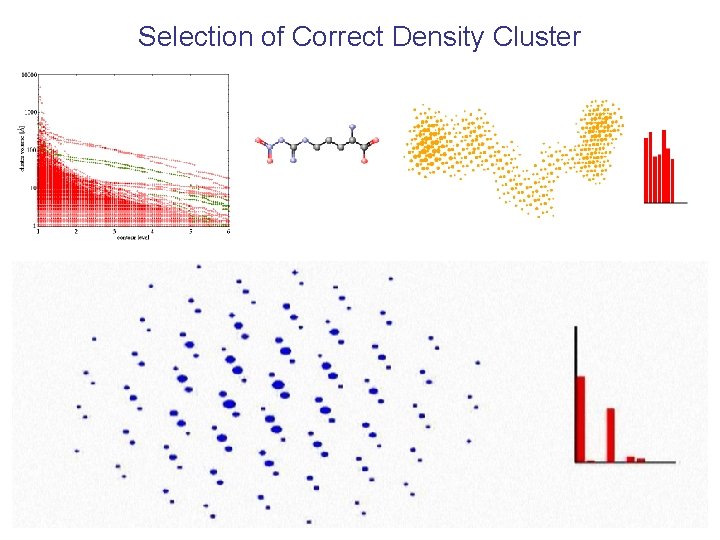
Selection of Correct Density Cluster
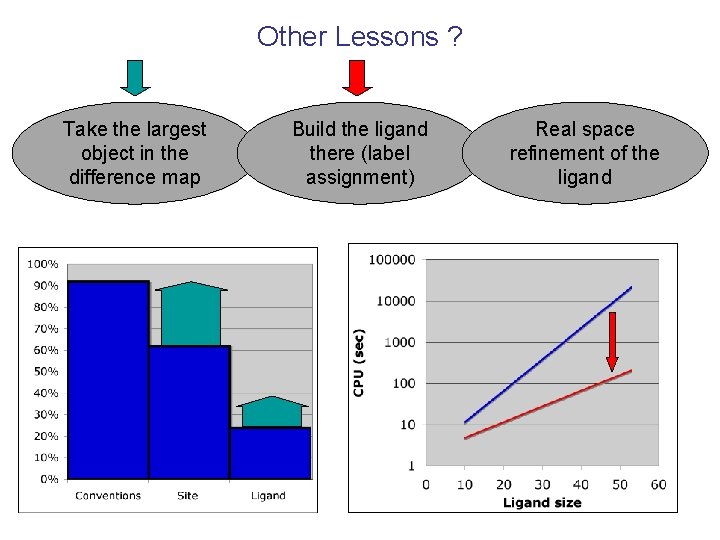
Other Lessons ? Take the largest object in the difference map Build the ligand there (label assignment) Real space refinement of the ligand
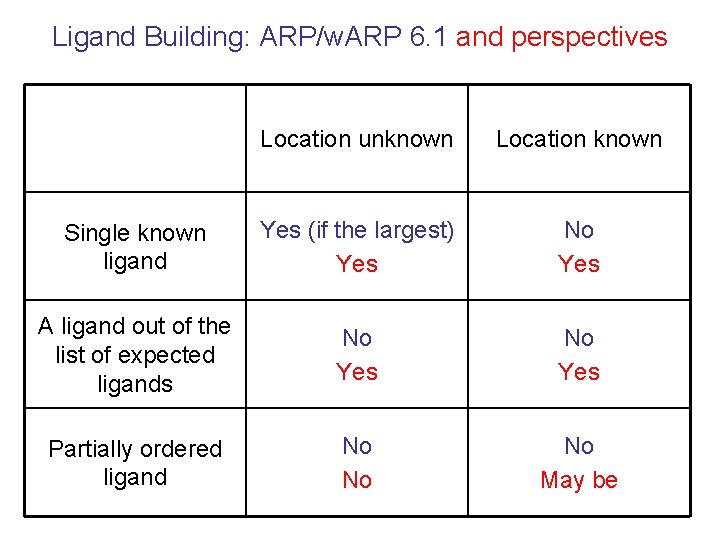
Ligand Building: ARP/w. ARP 6. 1 and perspectives Location unknown Location known Single known ligand Yes (if the largest) Yes No Yes A ligand out of the list of expected ligands No Yes Partially ordered ligand No No No May be

ARP/w. ARP - the people Developers EMBL Hamburg: Guillaume Evrard, Johan Hattne, Gerrit Langer, Venkat Parthasarathy, Tilo Strutz, Victor Lamzin and many in-house friends NKI Amsterdam: Serge Cohen, Diederick De Vries, Marouane Jelloul, Krista Joosten, Tassos Perrakis Former members and collaborators Richard Morris, Peter Zwart, Francisco Fernandez, Olga Kirillova, Matheos Kakaris, Gleb Bourenkov, Garib Murshudov, Alexei Vagin, Andrey Lebedev, Peter Briggs, Eleanor Dodson, Keith Wilson, Zbyszek Dauter, Gerard Klejwegt
- Slides: 32