Ligand binding Cyanide binding to the respiratory enzyme
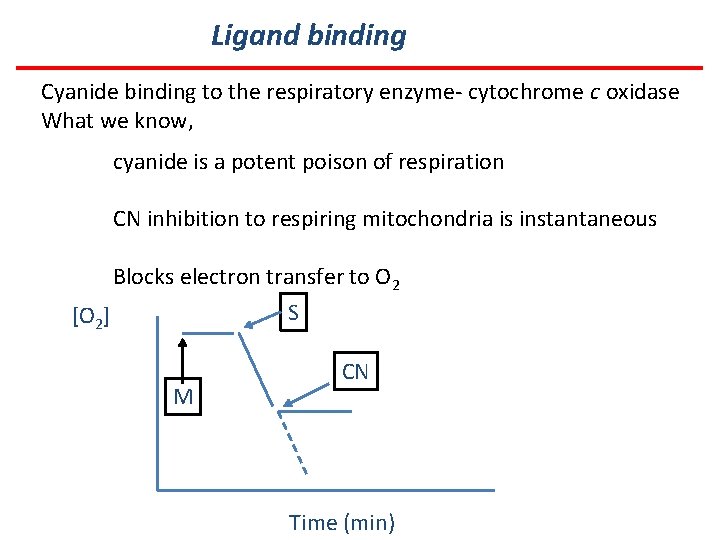
Ligand binding Cyanide binding to the respiratory enzyme- cytochrome c oxidase What we know, cyanide is a potent poison of respiration CN inhibition to respiring mitochondria is instantaneous Blocks electron transfer to O 2 S [O 2] M CN Time (min)

Cyanide reaction with cytochrome c oxidase A ACN=εCNc. CNl 1 A=εcl 0. 5 A 430 nm E-CN E +CN 0 400 420 440 460 Wavelength (nm) 0 1 10 100 Time (m) 1000
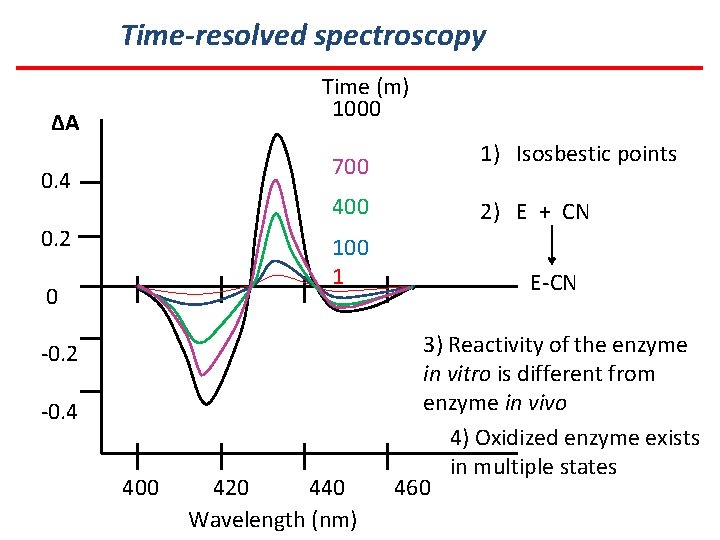
Time-resolved spectroscopy Time (m) 1000 ΔA 700 0. 4 400 0. 2 100 1 0 -0. 2 -0. 4 400 420 440 Wavelength (nm) 1) Isosbestic points 2) E + CN E-CN 3) Reactivity of the enzyme in vitro is different from enzyme in vivo 4) Oxidized enzyme exists in multiple states 460
![Ligand binding-equilibrium case ΔA KD=. 011 m. M ΔA =. 105 [ligand] (m. M) Ligand binding-equilibrium case ΔA KD=. 011 m. M ΔA =. 105 [ligand] (m. M)](http://slidetodoc.com/presentation_image_h2/13577805c274da7df555d430f9c32227/image-4.jpg)
Ligand binding-equilibrium case ΔA KD=. 011 m. M ΔA =. 105 [ligand] (m. M)
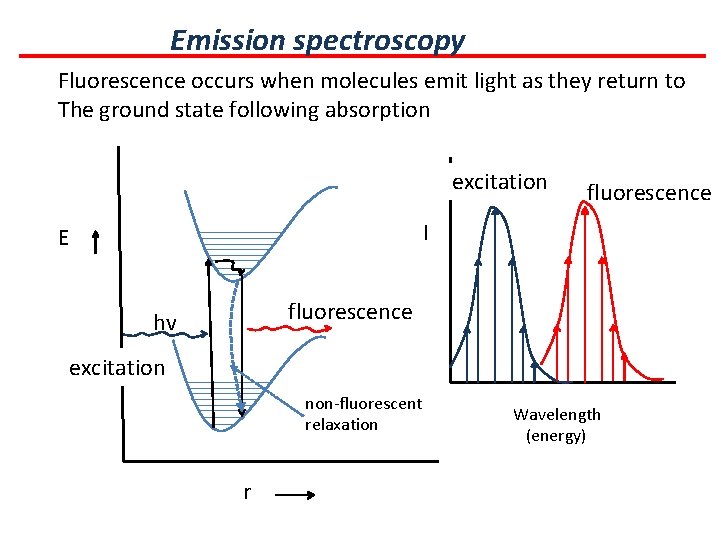
Emission spectroscopy Fluorescence occurs when molecules emit light as they return to The ground state following absorption excitation fluorescence I E fluorescence hv excitation non-fluorescent relaxation r Wavelength (energy)

The timescales of fluorescence 1) Excitation from ground state to excited state ~ 10 -15 s 2) Vibrational relaxation within excited state ~10 -12 s 3) Spontaneous emission 10 -6 -10 -9 s Other processes 4) Photochemistry – reactions from the excited state 5) Phosphorescence (ms to minutes) 6) Non-radiative relaxation to ground state

Fluorescence measurables 1) Excitation spectra (λmax, ex) 2) Emission spectra (λmax, em) 3) Intensity (Quantum yield) 4) Lifetime 5) Polarization

Lifetime, quantum yield and fluorescence intensity Radiative decay to the ground state is a first order process and lifetime is the inverse of decay rate, E τF = 1/k. F hv There are other processes (non-radiative) that excitation lead to depopulation of the excited state, k = k. F + ∑ki (overall lifetime τ=1/k) fluorescence r The quantum yield (ΦF)is the fraction of molecules that return to the ground state via a fluorescence pathway, ΦF = k. F / (k. F + ∑ki) = τ / τF where τ is the observed lifetime of the overall process Fluorescence intensity (F) will then depend on the initial population of the excited state (IA) and the quantum yield, F = IA ΦF
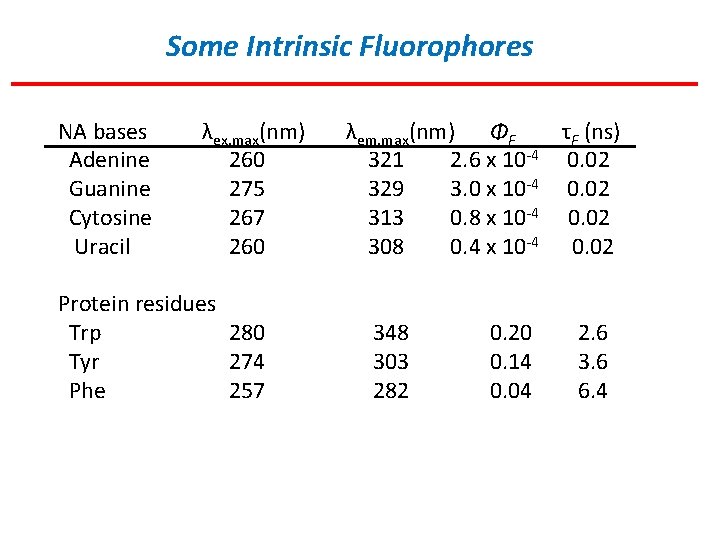
Some Intrinsic Fluorophores NA bases Adenine Guanine Cytosine Uracil λex, max(nm) 260 275 267 260 Protein residues Trp 280 Tyr 274 Phe 257 λem, max(nm) ΦF τF (ns) 321 2. 6 x 10 -4 0. 02 329 3. 0 x 10 -4 0. 02 313 0. 8 x 10 -4 0. 02 308 0. 4 x 10 -4 0. 02 348 303 282 0. 20 0. 14 0. 04 2. 6 3. 6 6. 4
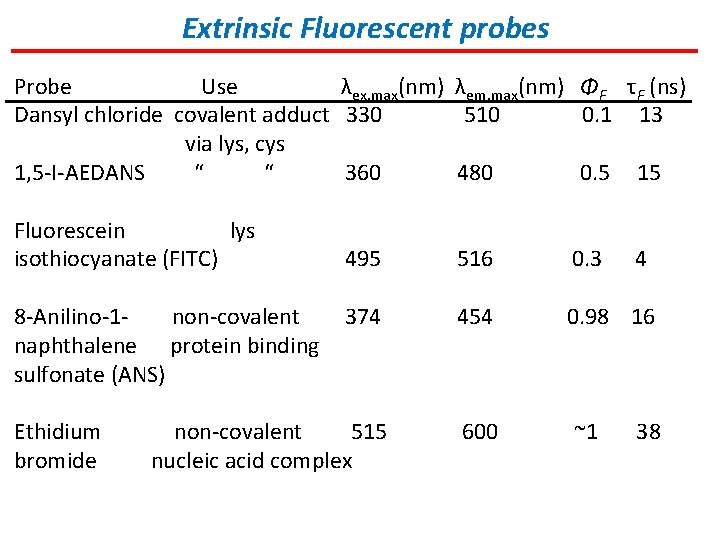
Extrinsic Fluorescent probes Probe Use λex, max(nm) λem, max(nm) ΦF τF (ns) Dansyl chloride covalent adduct 330 510 0. 1 13 via lys, cys 1, 5 -I-AEDANS “ “ 360 480 0. 5 15 Fluorescein lys isothiocyanate (FITC) 495 516 0. 3 374 454 0. 98 16 non-covalent 515 nucleic acid complex 600 8 -Anilino-1 non-covalent naphthalene protein binding sulfonate (ANS) Ethidium bromide ~1 4 38
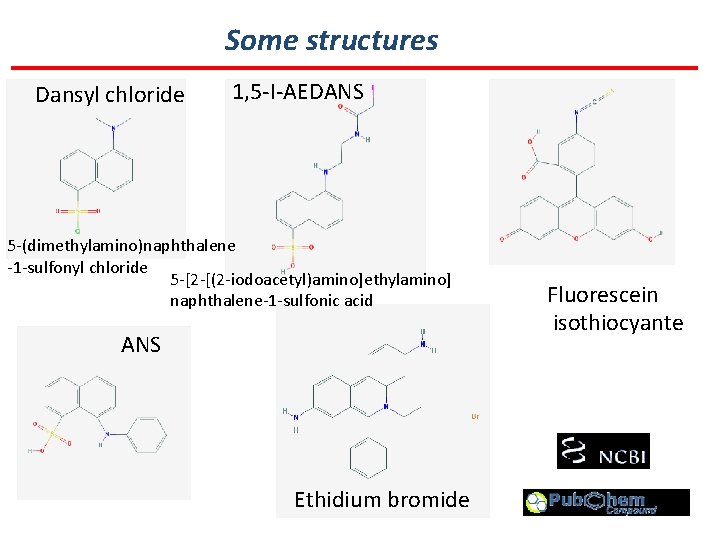
Some structures Dansyl chloride 1, 5 -I-AEDANS 5 -(dimethylamino)naphthalene -1 -sulfonyl chloride 5 -[2 -[(2 -iodoacetyl)amino]ethylamino] naphthalene-1 -sulfonic acid ANS Ethidium bromide Fluorescein isothiocyante
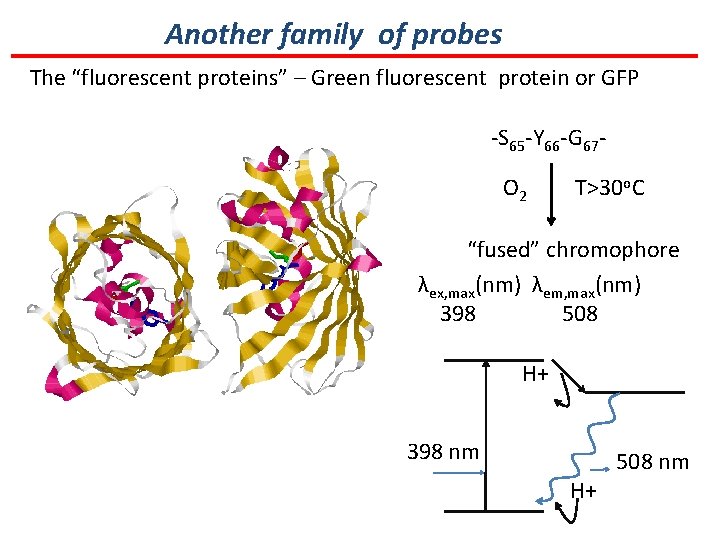
Another family of probes The “fluorescent proteins” – Green fluorescent protein or GFP -S 65 -Y 66 -G 67 O 2 T>30 o. C “fused” chromophore λex, max(nm) λem, max(nm) 398 508 H+ 398 nm H+ 508 nm
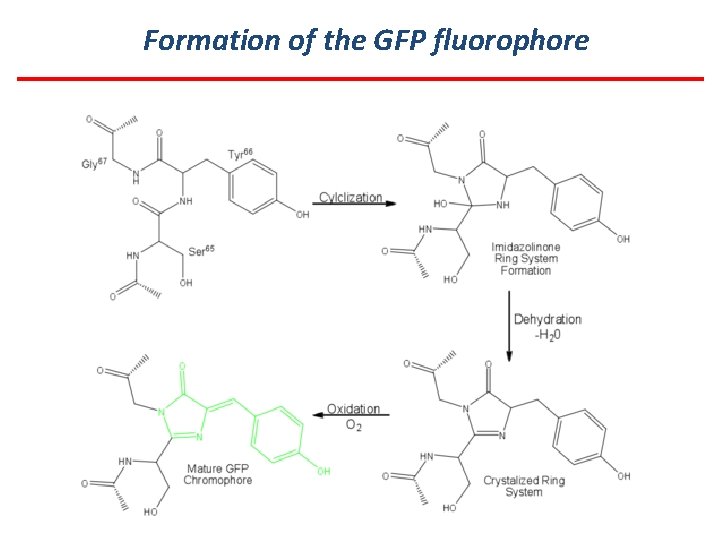
Formation of the GFP fluorophore
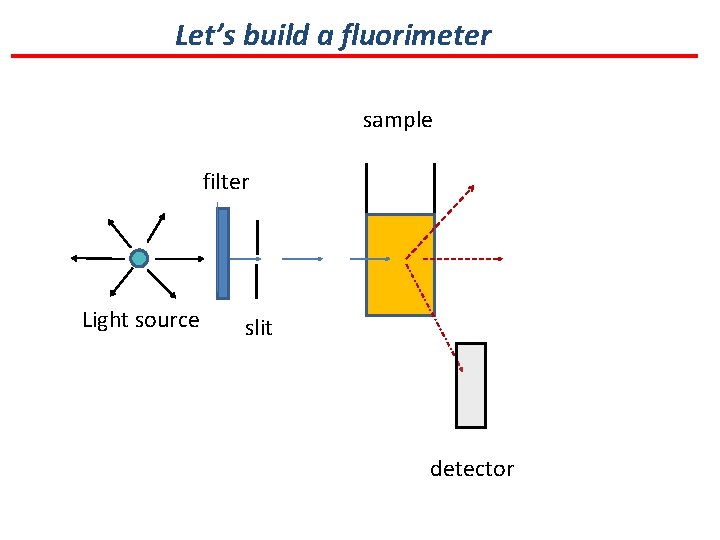
Let’s build a fluorimeter sample filter Light source slit detector
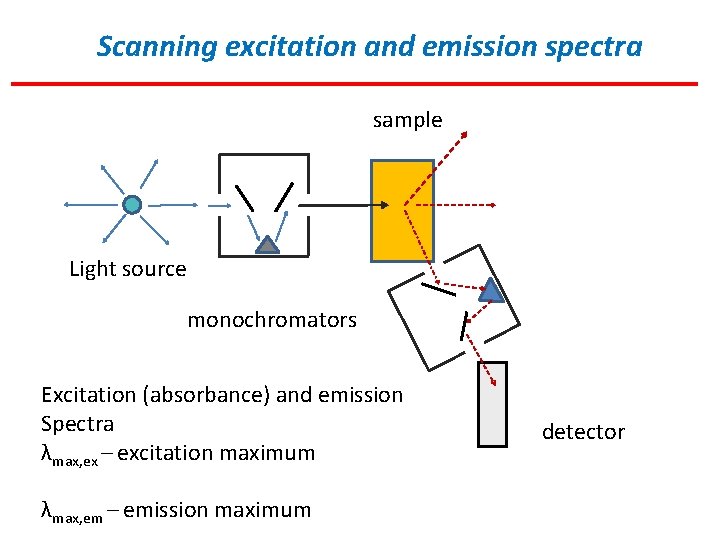
Scanning excitation and emission spectra sample Light source monochromators Excitation (absorbance) and emission Spectra λmax, ex – excitation maximum λmax, em – emission maximum detector

Scanning excitation and emission spectra
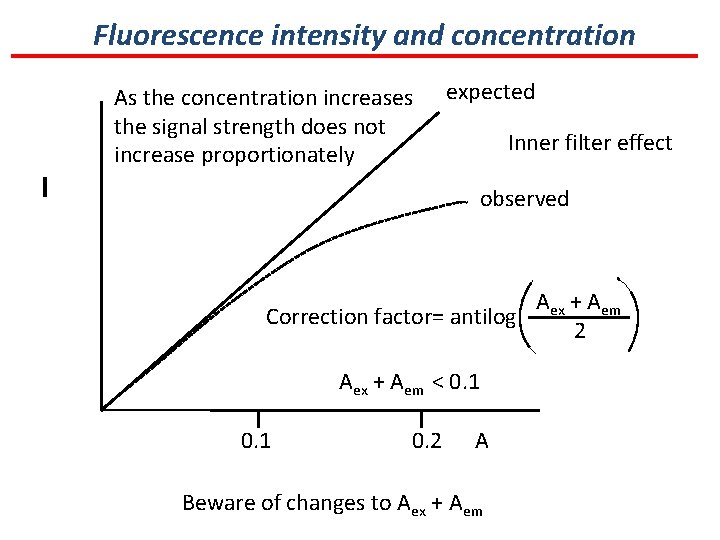
Fluorescence intensity and concentration I As the concentration increases the signal strength does not increase proportionately expected Inner filter effect observed Aex + Aem Correction factor= antilog 2 Aex + Aem < 0. 1 0. 2 Concentration A Beware of changes to Aex + Aem (α A)
- Slides: 17