Lesson starter 1 Name the four bases found

Lesson starter 1. Name the four bases found in DNA 2. Which DNA bases pair together in complementary base pairing? 3. What determines the order of amino acids found in protein? 4. Name is the name of the three adjacent bases in an m. RNA molecule called? 5. Where in the cell is m. RNA made?

Translation Learning objectives: Describe, with the aid of diagrams, the way in which a nucleotide sequence codes for the amino acid sequence in a polypeptide, including the role of messenger RNA, transfer RNA and ribosomes. State that cyclic AMP activates proteins by altering their three-dimensional structure
Summary So Far… • Use the pictures to help you describe what you have learned so far.
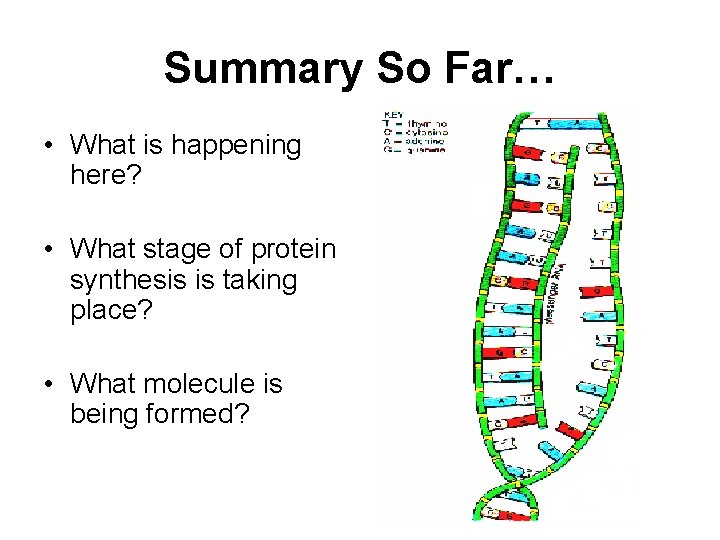
Summary So Far… • What is happening here? • What stage of protein synthesis is taking place? • What molecule is being formed?
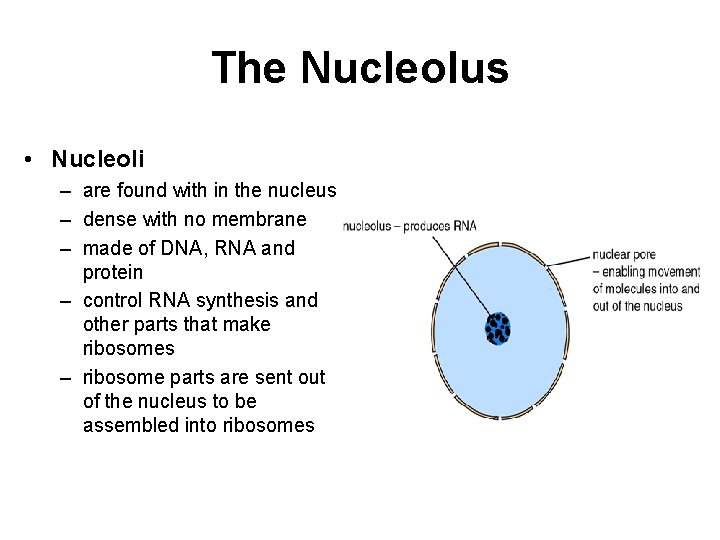
The Nucleolus • Nucleoli – are found with in the nucleus – dense with no membrane – made of DNA, RNA and protein – control RNA synthesis and other parts that make ribosomes – ribosome parts are sent out of the nucleus to be assembled into ribosomes
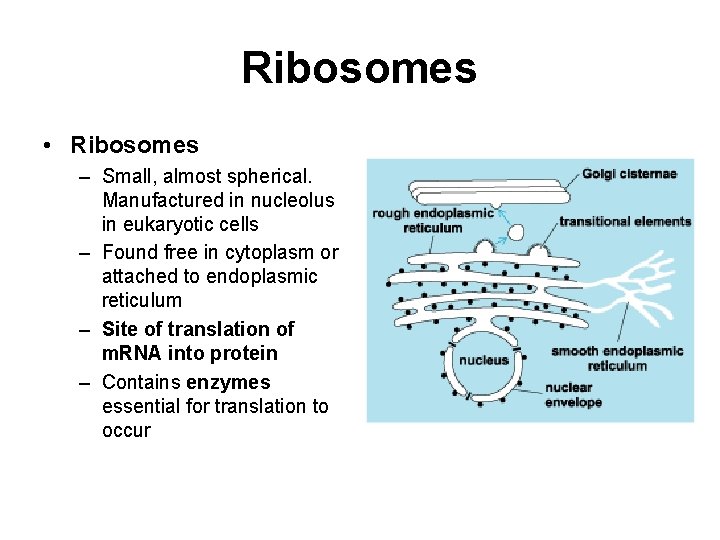
Ribosomes • Ribosomes – Small, almost spherical. Manufactured in nucleolus in eukaryotic cells – Found free in cytoplasm or attached to endoplasmic reticulum – Site of translation of m. RNA into protein – Contains enzymes essential for translation to occur
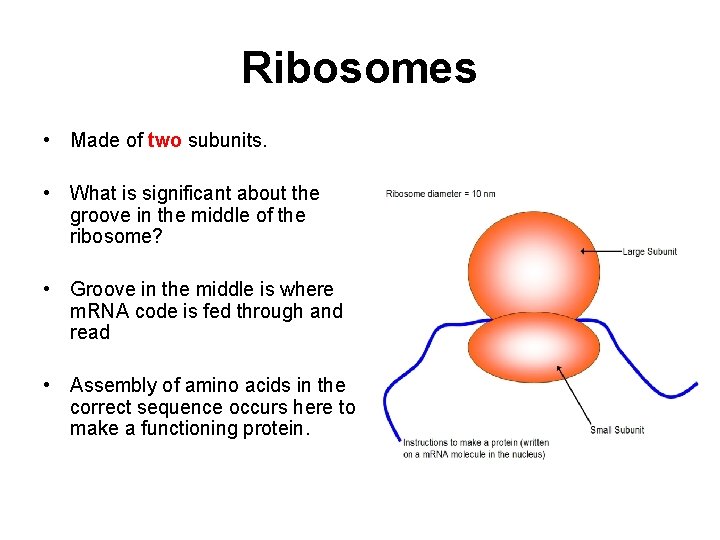
Ribosomes • Made of two subunits. • What is significant about the groove in the middle of the ribosome? • Groove in the middle is where m. RNA code is fed through and read • Assembly of amino acids in the correct sequence occurs here to make a functioning protein.

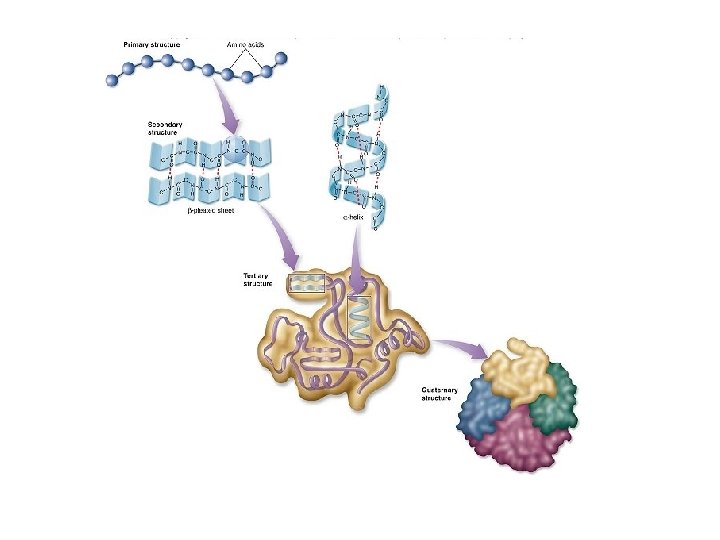
Making the correct proteins • Amino acids joined by peptide bonds make the primary structure of protein • Explain how the primary structure of a protein determines the tertiary structure • The primary structure determines the tertiary structure – How protein folds to made 3 D structure – 3 D structure is held in place by hydrogen/ionic bonds and hydrophobic interactions between R groups of amino acids – Tertiary structure is the functional protein. Therefore if the genetic sequence to make the primary structure is incorrect, the final protein will not function correctly.
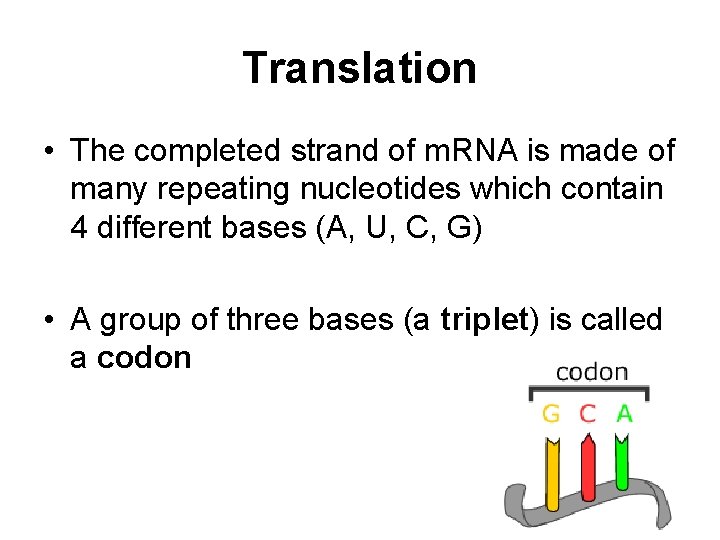
Translation • The completed strand of m. RNA is made of many repeating nucleotides which contain 4 different bases (A, U, C, G) • A group of three bases (a triplet) is called a codon
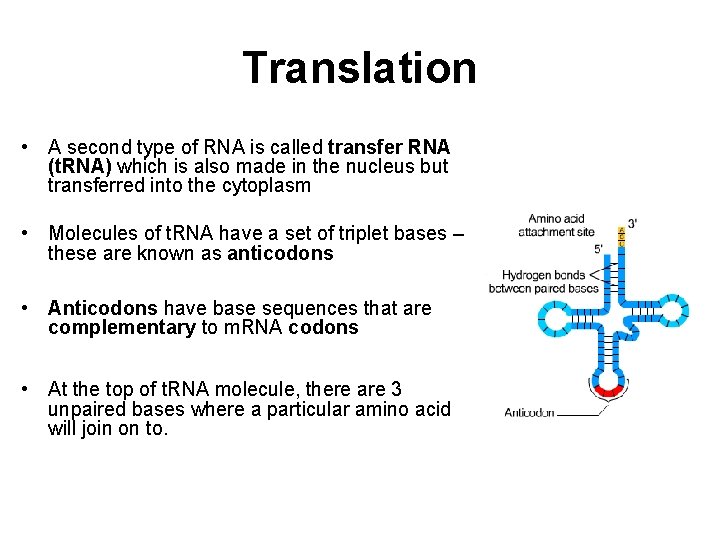
Translation • A second type of RNA is called transfer RNA (t. RNA) which is also made in the nucleus but transferred into the cytoplasm • Molecules of t. RNA have a set of triplet bases – these are known as anticodons • Anticodons have base sequences that are complementary to m. RNA codons • At the top of t. RNA molecule, there are 3 unpaired bases where a particular amino acid will join on to.
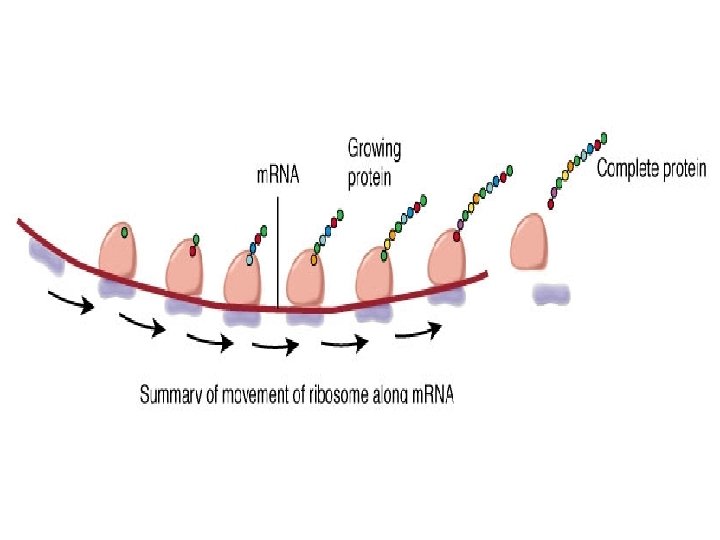
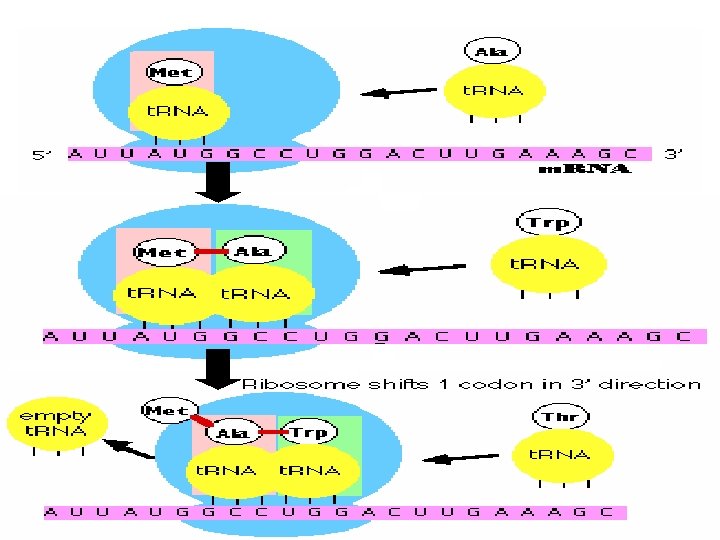

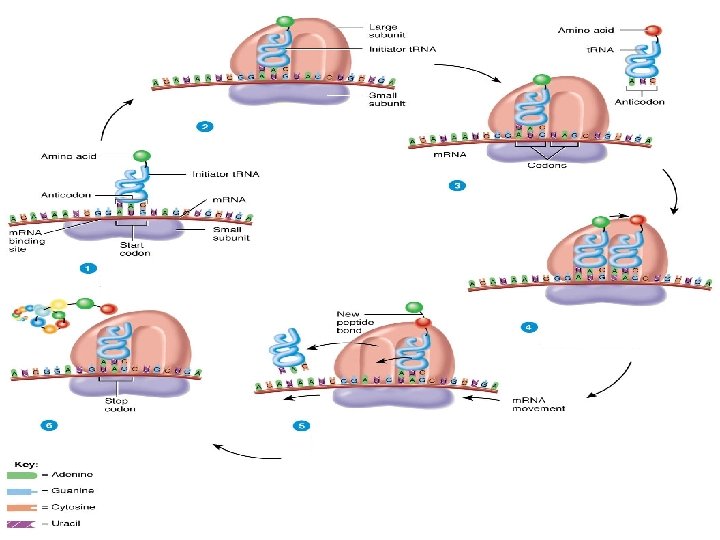
Translation 1. The m. RNA attaches itself to a ribosome and transfer RNA (t. RNA) molecules carry amino acids to the ribosome. 2. The first amino acid in any polypeptide is usually methionine. The codon for this is AUG, which has come to be known as the initiation codon as a result. 3. The t. RNA anticodon is therefore UAC. These attach together by complementary base pairing 4. The second t. RNA molecule attaches itself to the next codon on the m. RNA in the same way. 5. The two amino acids attached to the t. RNA molecules are joined by a peptide bond. The first t. RNA moves away, leaving the amino acid behind. 6. A third t. RNA molecule binds to the next codon on the m. RNA. Its amino acid binds to the first two and the second t. RNA molecule moves away. 7. This process continues, producing a chain of linked amino acids (a polypeptide chain) until there is a stop codon on the m. RNA molecule. 8. The polypeptide chain moves away from the ribosome and translation is complete.
U U A G C A U C G U A
U U A G C A U C G U A GCG A
U U A G C A U C G U A UGCG A

the m. RNA leaves the nucleus via a pore in the nuclear membrane U U A G C A U C G U A UGCG A
U U A G C UGCG A

U U UGCG A

UGCG
UG
U U U C G A U G C A U C G C A A C U C G C
U U U C G A U G C A U C G C A A C U C G C

U U U C G A U G C A U C G C A A C U C G C
U U U C G A U G C A U C G C A A C U C G C
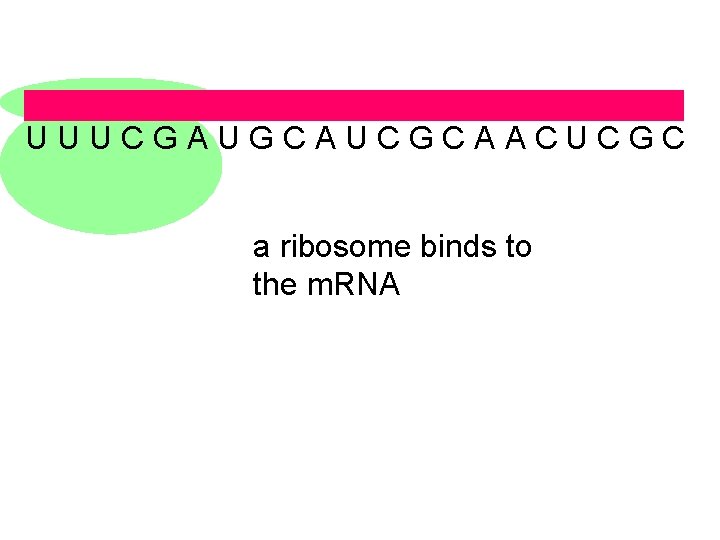
U U U C G A U G C A U C G C A A C U C G C a ribosome binds to the m. RNA

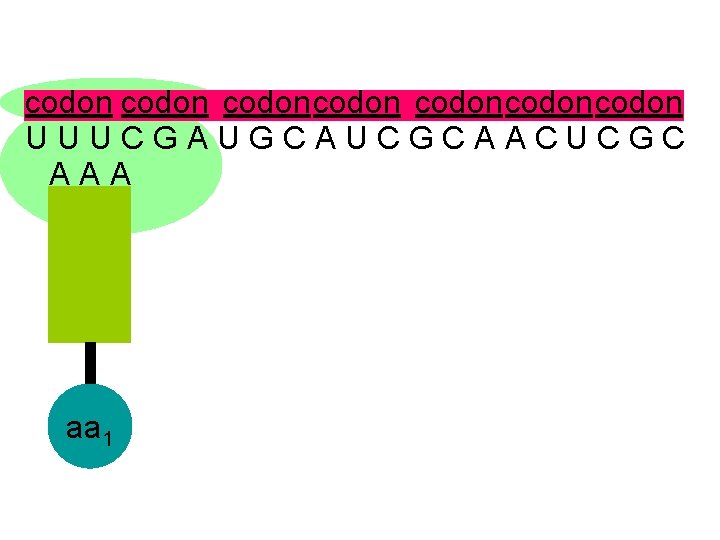
codon U U U C G A U G C A U C G C A A C U C G C the genetic code is read in groups of 3 letters called codons

codon codoncodon U U U C G A U G C A U C G C A A C U C G C
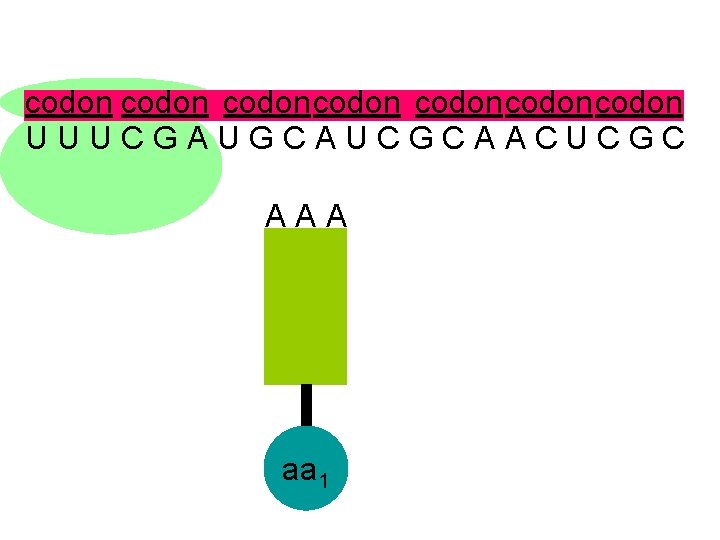
codon codoncodon U U U C G A U G C A U C G C A A C U C G C A A A aa 1
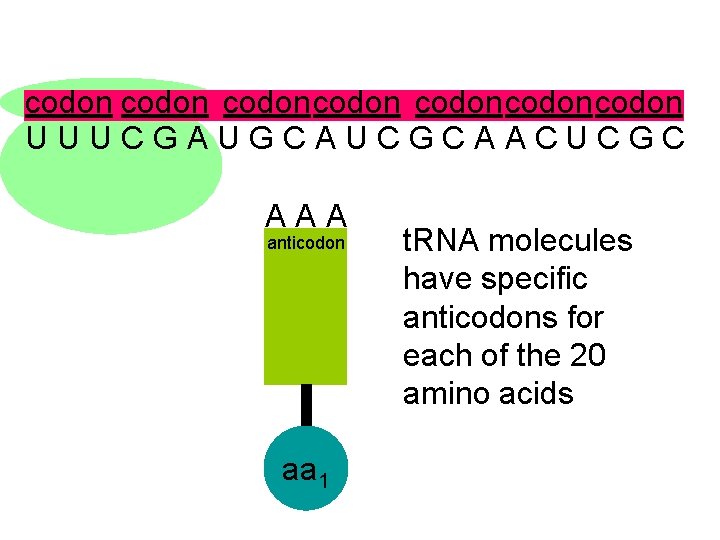
codon codoncodon U U U C G A U G C A U C G C A A C U C G C A A A anticodon aa 1 t. RNA molecules have specific anticodons for each of the 20 amino acids
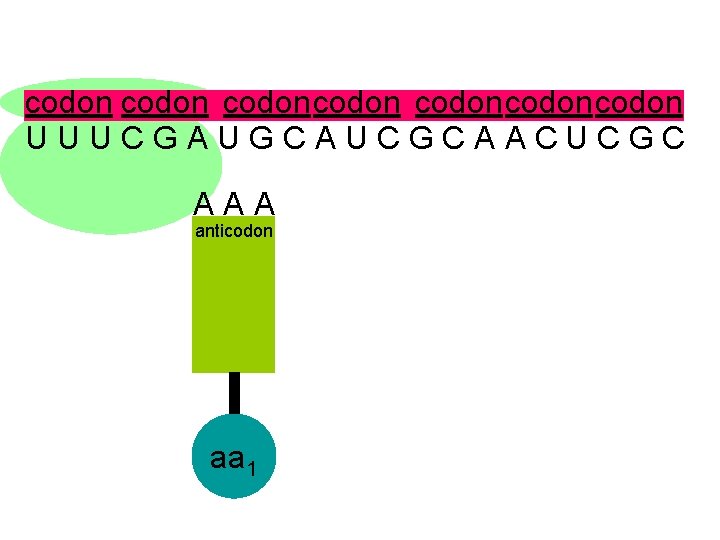
codon codoncodon U U U C G A U G C A U C G C A A C U C G C A A A anticodon aa 1
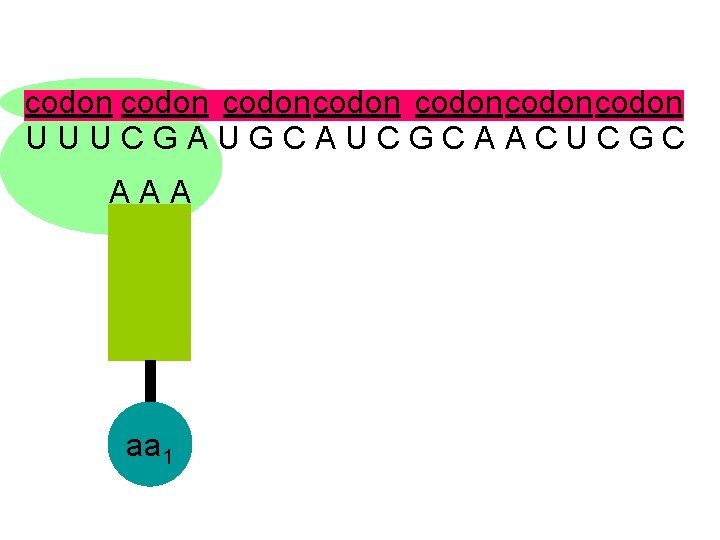
codon codoncodon U U U C G A U G C A U C G C A A C U C G C A A A aa 1
codon codoncodon U U U C G A U G C A U C G C A A C U C G C A A A aa 1

codon codoncodon U U U C G A U G C A U C G C A A C U C G C A A A the complementary anticodon is attracted to the first codon on the m. RNA aa 1
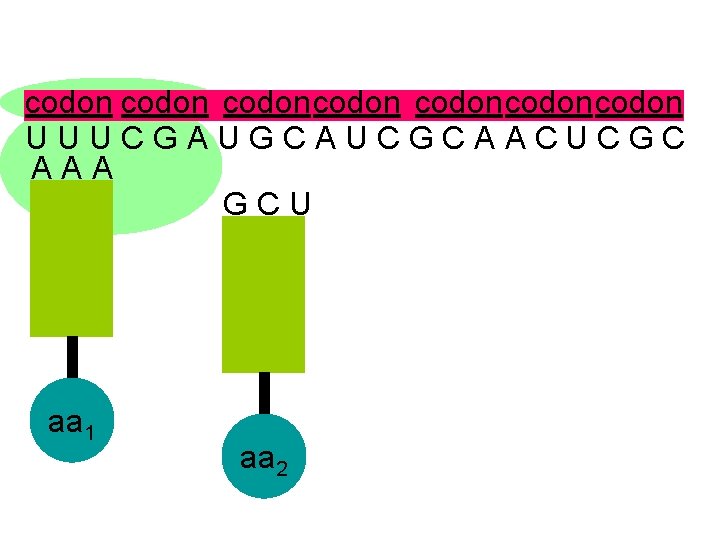
codon codoncodon U U U C G A U G C A U C G C A A C U C G C A A A G C U aa 1 aa 2
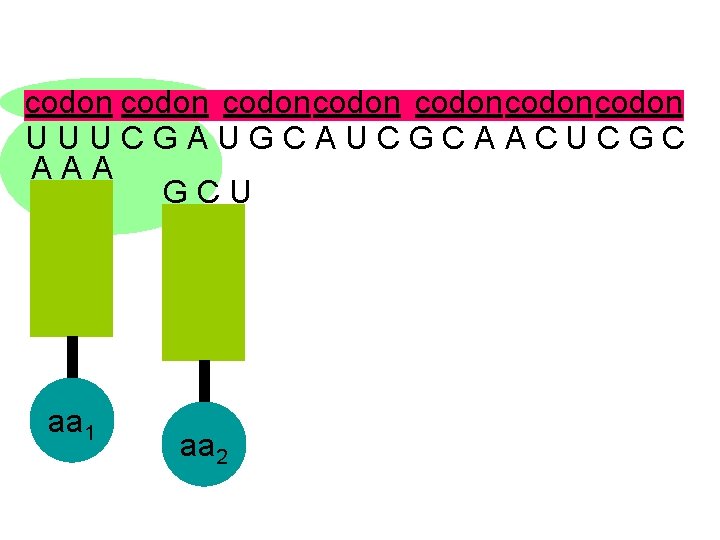
codon codoncodon U U U C G A U G C A U C G C A A C U C G C A A A G C U aa 1 aa 2
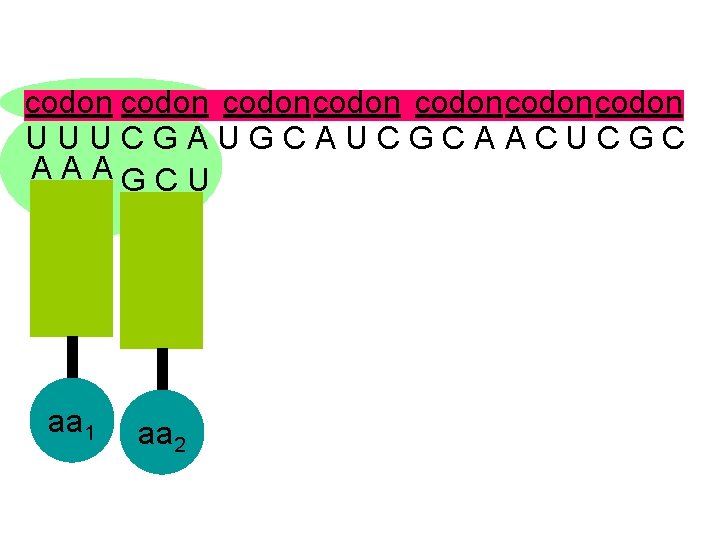
codon codoncodon U U U C G A U G C A U C G C A A C U C G C A A A G C U aa 1 aa 2
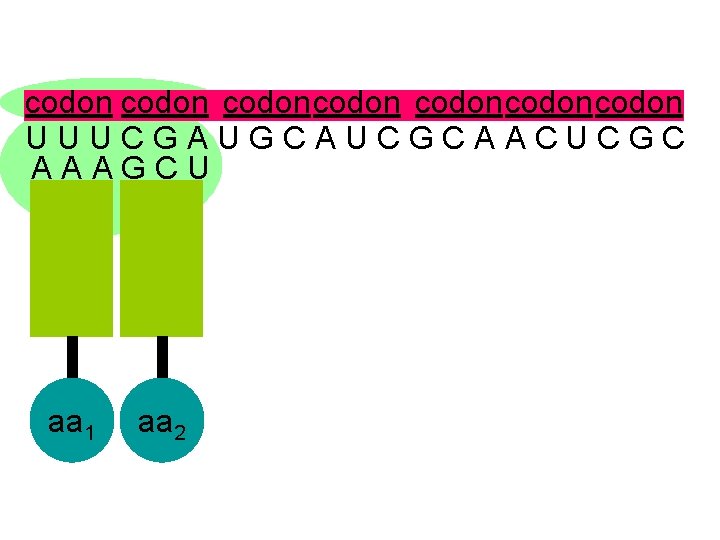
codon codoncodon U U U C G A U G C A U C G C A A C U C G C A A A G C U aa 1 aa 2
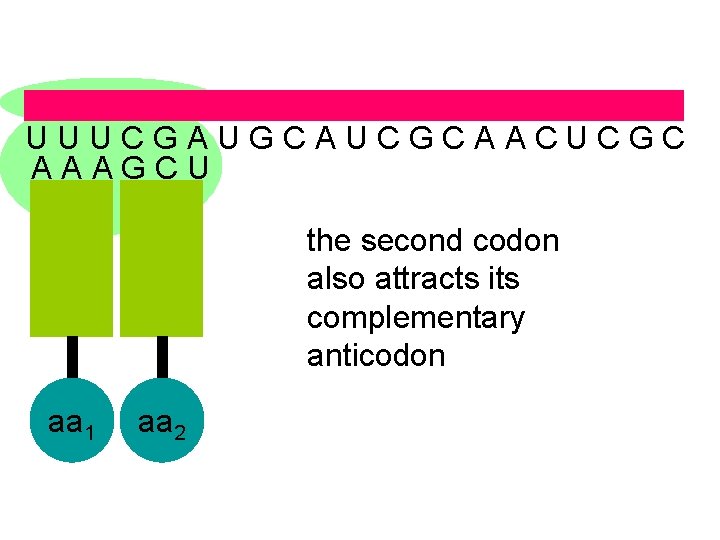
U U U C G A U G C A U C G C A A C U C G C A A A G C U the second codon also attracts its complementary anticodon aa 1 aa 2
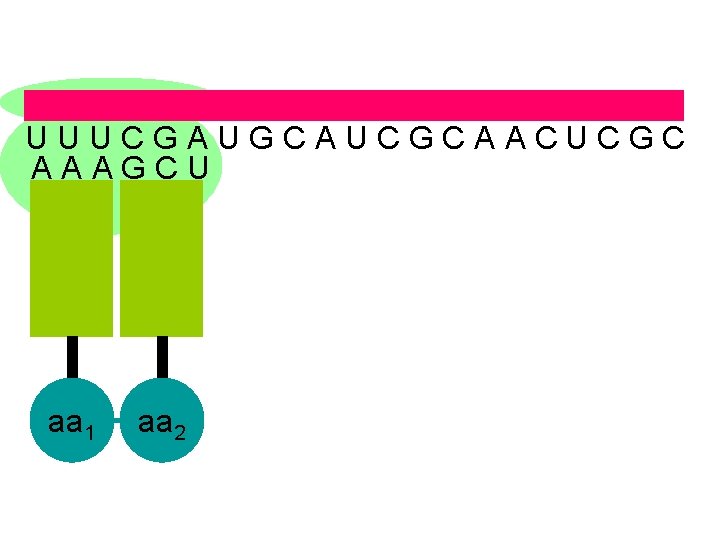
U U U C G A U G C A U C G C A A C U C G C A A A G C U aa 1 aa 2
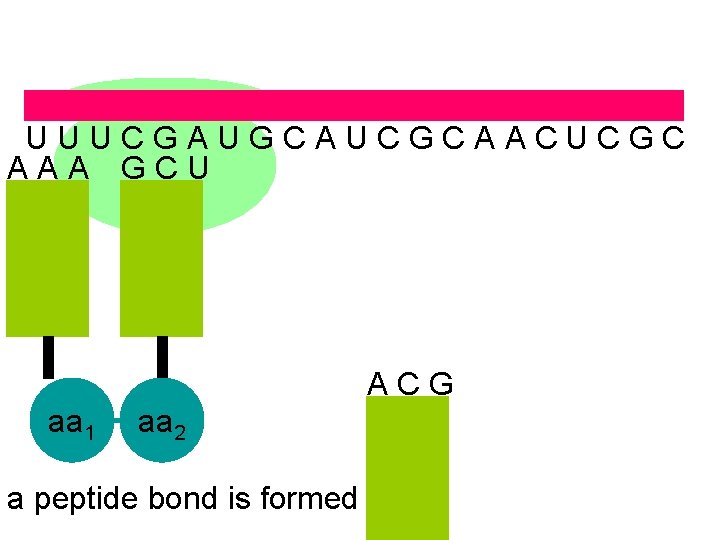
U U U C G A U G C A U C G C A A C U C G C A A A G C U A C G aa 1 aa 2 a peptide bond is formed
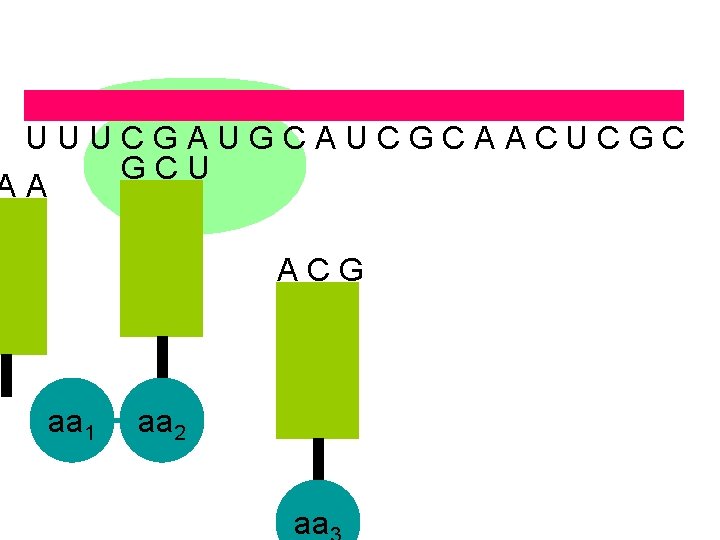
U U U C G A U G C A U C G C A A C U C G C U A A A C G aa 1 aa 2 aa
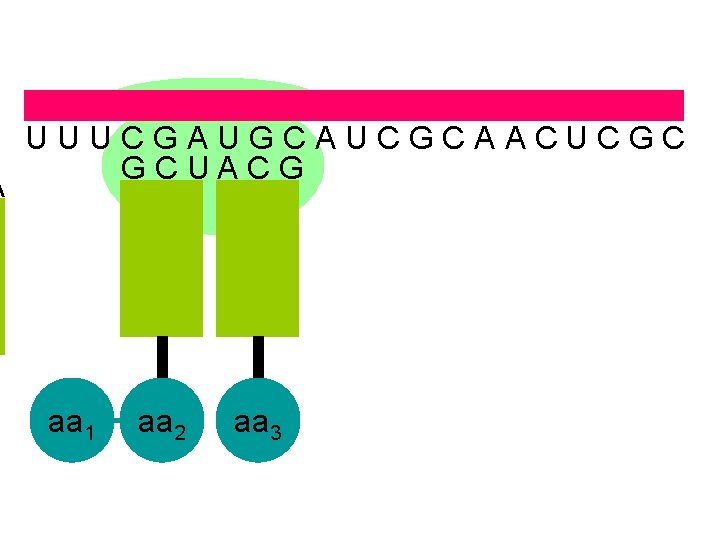
A U U U C G A U G C A U C G C A A C U C G C U A C G aa 1 aa 2 aa 3
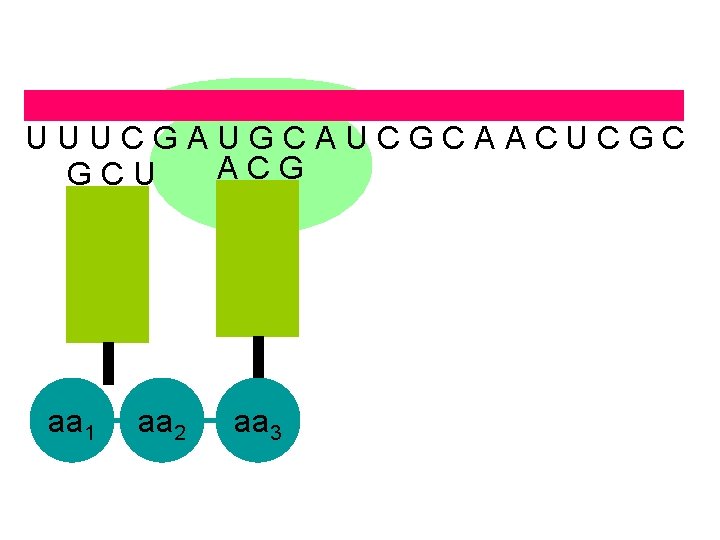
U U U C G A U G C A U C G C A A C U C G C A C G G C U aa 1 aa 2 aa 3
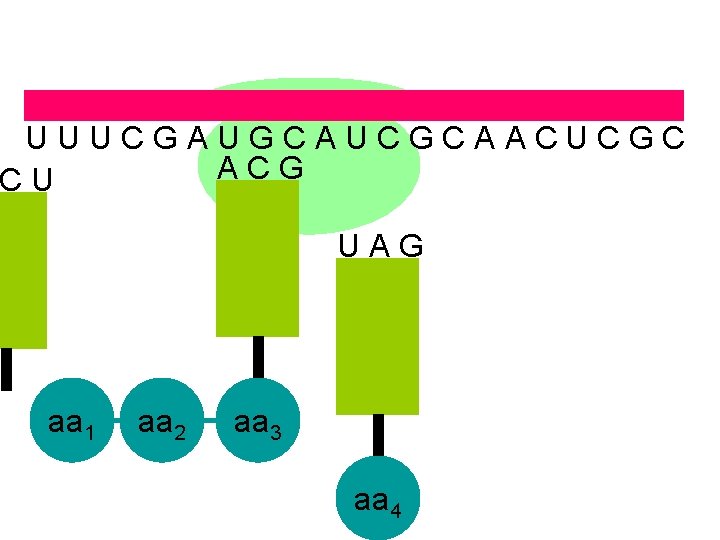
U U U C G A U G C A U C G C A A C U C G C A C G C U U A G aa 1 aa 2 aa 3 aa 4
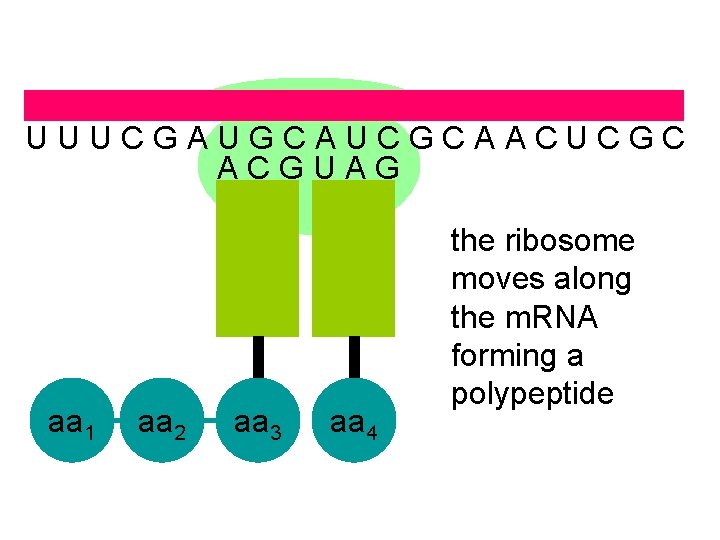
U U U C G A U G C A U C G C A A C U C G C A C G U A G aa 1 aa 2 aa 3 aa 4 the ribosome moves along the m. RNA forming a polypeptide
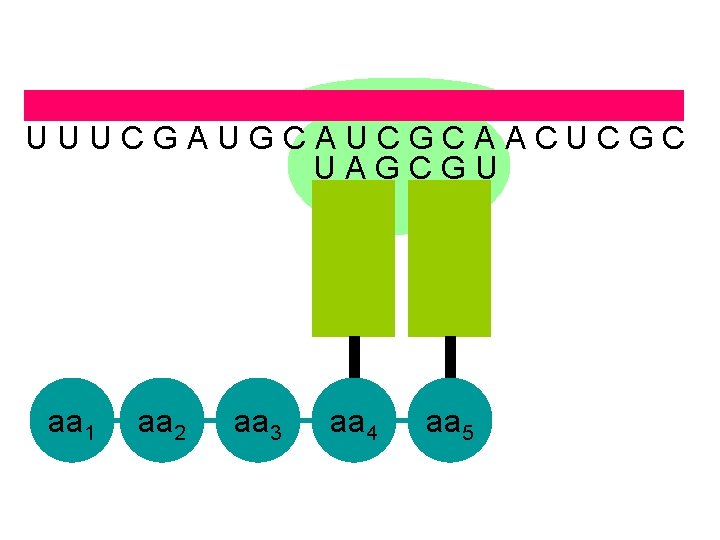
U U U C G A U G C A U C G C A A C U C G C U A G C G U aa 1 aa 2 aa 3 aa 4 aa 5
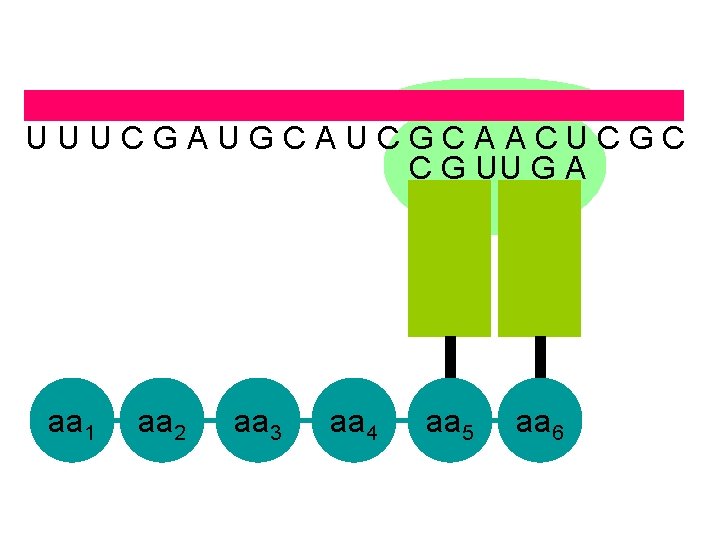
U U U C G A U G C A U C G C A A C U C G C C G U U G A aa 1 aa 2 aa 3 aa 4 aa 5 aa 6
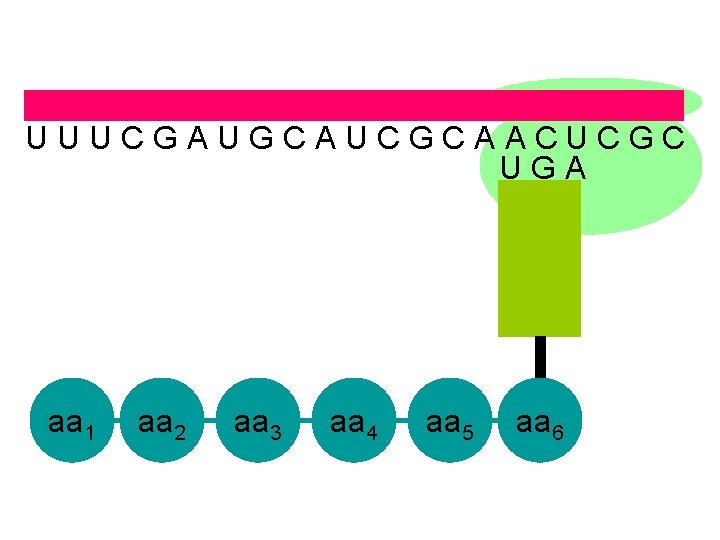
U U U C G A U G C A U C G C A A C U C G C U G A aa 1 aa 2 aa 3 aa 4 aa 5 aa 6
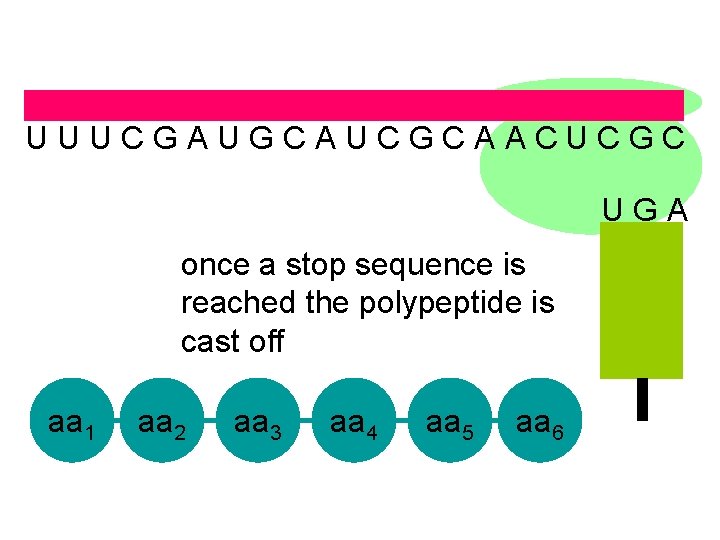
U U U C G A U G C A U C G C A A C U C G C U G A once a stop sequence is reached the polypeptide is cast off aa 1 aa 2 aa 3 aa 4 aa 5 aa 6
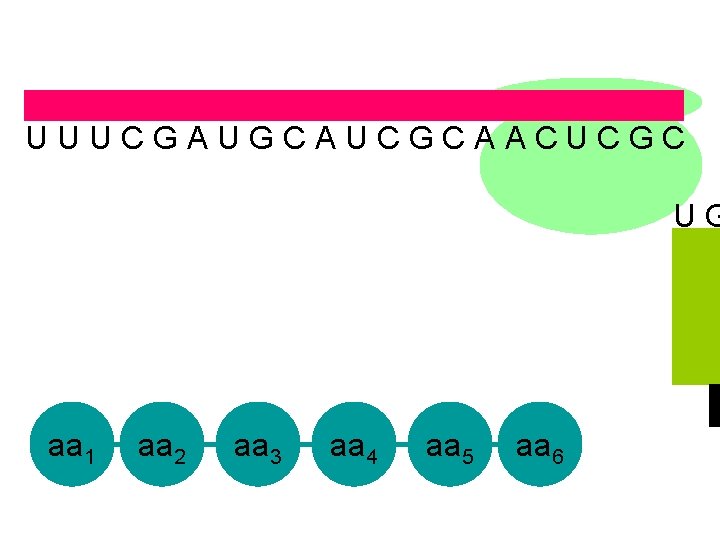
U U U C G A U G C A U C G C A A C U C G C U G aa 1 aa 2 aa 3 aa 4 aa 5 aa 6
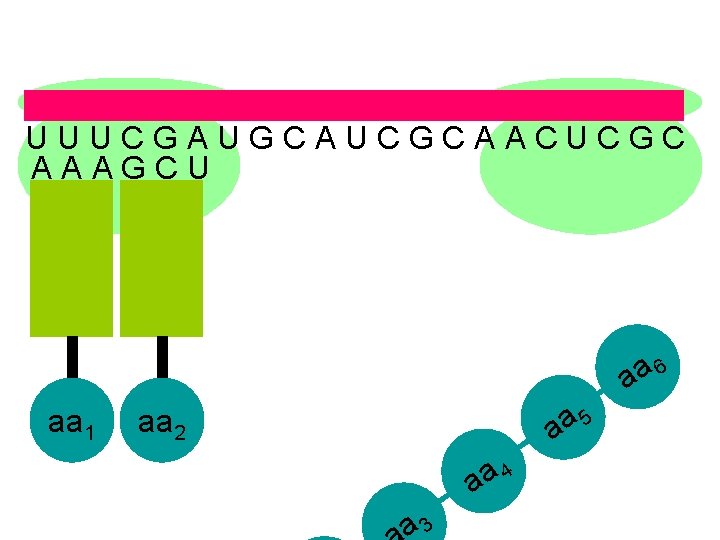
U U U C G A U G C A U C G C A A C U C G C A A A G C U aa 6 aa 1 5 a a aa 2 aa 4 a 3
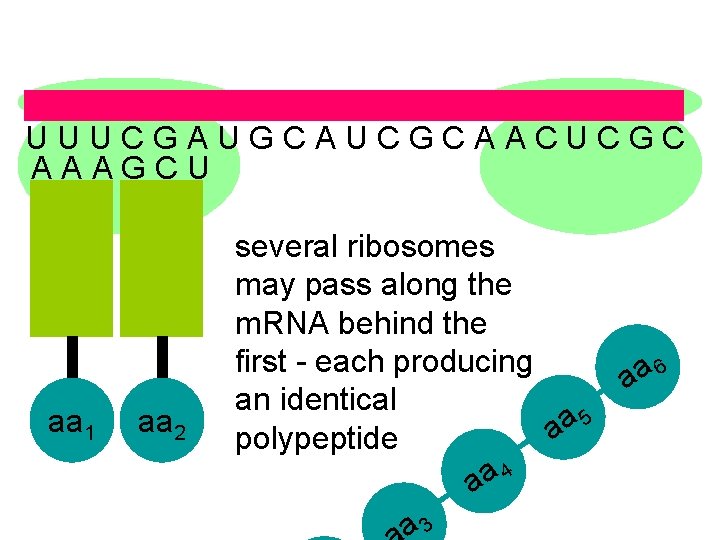
U U U C G A U G C A U C G C A A C U C G C A A A G C U aa 1 aa 2 several ribosomes may pass along the m. RNA behind the first - each producing aa 6 an identical 5 a a polypeptide aa 4 a 3
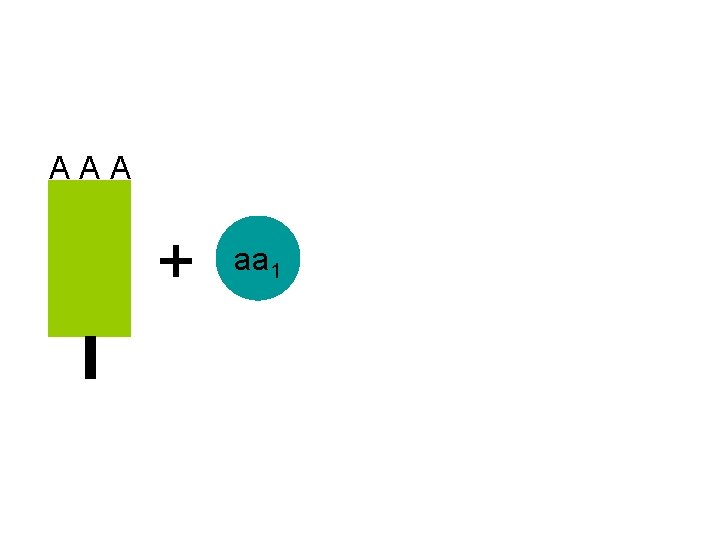
A A A + aa 1
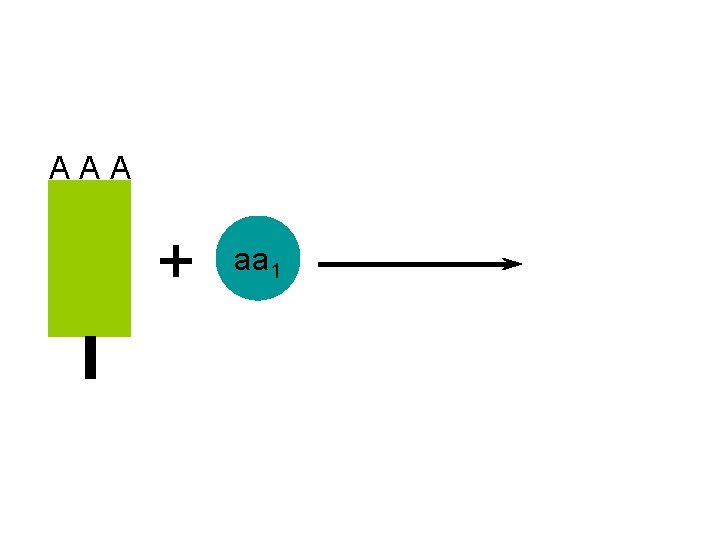
A A A + aa 1
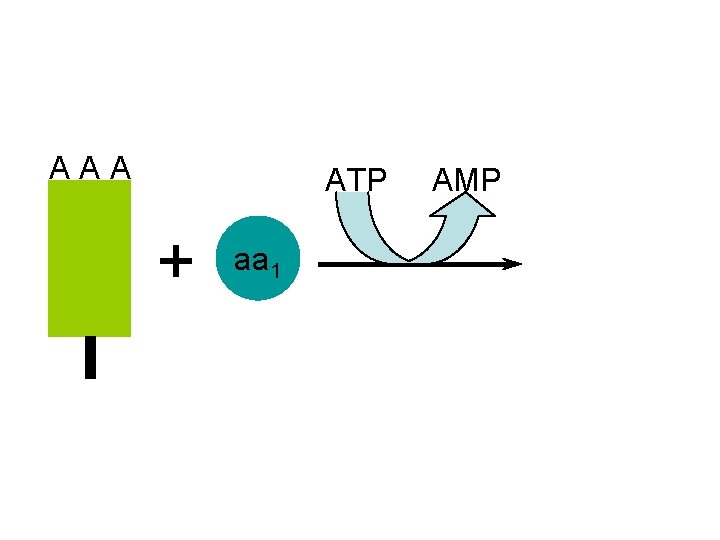
A A A ATP AMP + aa 1
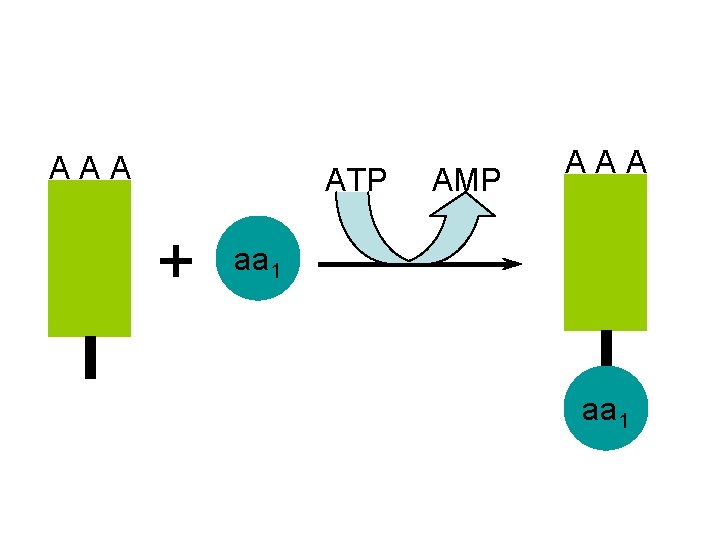
A A A ATP AMP + A A A aa 1
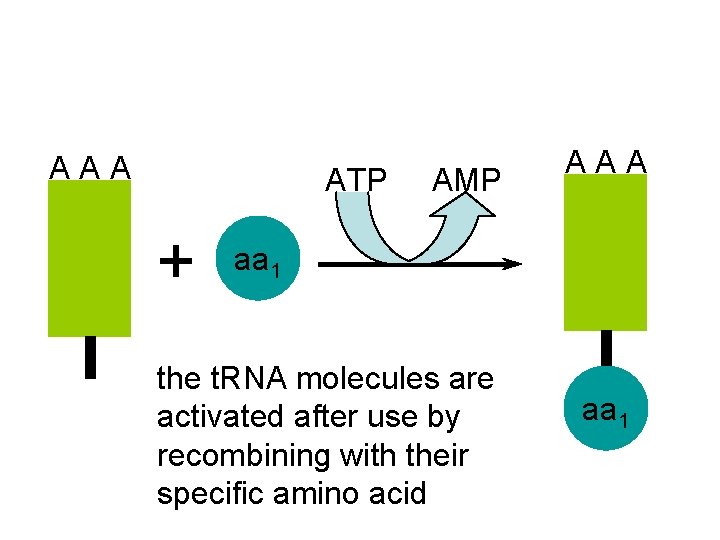
A A A ATP AMP + A A A aa 1 the t. RNA molecules are activated after use by recombining with their specific amino acid aa 1
Plenary • _____is the second stage of protein synthesis • _________ assemble to make a protein • This happens in a sequence dictated by the sequence of _____(triplet of bases of m. RNA) • The genetic code for this is copied from ______in the nucleus on to m. RNA and now translated into a sequence of amino acids, attached to moleucles of _____ • ______are the site of protein synthesis. They can be found free in the cytoplasm or attached to endoplasmic reticulum (rough ER)

Plenary • Translation is the second stage of protein synthesis • Amino acids assemble to make a protein • This happens in a sequence dictated by the sequence of codons (triplet of bases of m. RNA) • The genetic code for this is copied from DNA in the nucleus on to m. RNA and now translated into a sequence of amino acids, attached to moleucles of t. RNA • Ribosomes are the site of protein synthesis. They can be found free in the cytoplasm or attached to endoplasmic reticulum (rough ER)

Cyclic AMP– protein activation • Some proteins need to be activated before they will work • Protein activation is also controlled by molecules e. g. hormones and sugars • Some of these molecules work by binding to cell membranes and triggering the production of c. AMP inside the cell • c. AMP activates proteins – alters 3 D structure – Enzymes can be made active by c. AMP binding to protein and moving the active site so that it is open

Cyclic AMP – protein activation Example – protein kinase A (PKA) 1. PKA has 4 subunits 2. When c. AMP is not bound, the four subunits are bound together and are therefore inactive 3. When c. AMP binds at receptor sites, it causes a change in the enzyme’s 3 D structure, releasing the active subunits. 4. Now PKA is active Your task: Read the stretch and challenge section on page 107. Draw out a series of diagrams to go with the above description of c. AMP activity in protein activation. Answer questions A and B to consolidate this note.

Quiz Questions 1. What are the two stages of protein synthesis called? 2. Where does the first stage of protein synthesis take place? 3. When does RNA polymerase stop making m. RNA? 4. Where does the second stage of protein synthesis take place?

Exam questions 1. A drug that inhibits cell growth is found to be able to bind to DNA, preventing RNA polymerase from binding. Explain how this drug will affect protein synthesis (2) 2. A polypeptide chain (protein) from a eukaryotic cell is 10 amino acids long. a) Predict how ling the m. RNA for this protein would be in nucleotides (without the start and stop codons). Explain your answer. (2) b) Describe how m. RNA is translated into the polypeptide chain (6)

Exam answers 1. The drug binds to DNA, preventing RNA polymerase from binding, so transcription cannot take place and no m. RNA can be made (1 mark) This means that there is no m. RNA for translation and so protein synthesis is inhibited (1 mark) 2. (a) 10 x 3 = 30 nucleotides long (1 mark) each amino acid is coded for by three nucleotides (a codon), so the m. RNA length is nucleotides is the number of amino acids multiplied by three (1 mark)

Exam answers 2(b) • The m. RNA molecule attaches itself to a ribosome and transfer RNA (t. RNA) molecule carry amino acids to the ribosome • A t. RNA molecule, with an anticodon that is complementary to the first codon on the m. RNA (the start codon), attaches itself to the m. RNA by complementary base paring. • A second t. RNA molecule attaches itself to the codon on the m. RNA in the same way • The two amino acids attached to the t. RNA molecules are joined by a peptide bond and the first t. RNA molecule moves away, leaving its amino acid behind • A third t. RNA molecule binds to the next codon on the m. RNA and its amino acid binds to the first two and the second t. RNA molecule moves away • This process continues, producing a chain of linked amino acids (a polypeptide chain), until there is a stop codon on the m. RNA molecule

- Slides: 71