LESSON SEQUENCE PROCESSING Goals Introduce DNA Assembly and

LESSON: SEQUENCE PROCESSING Goals: - Introduce DNA Assembly and Alignment Practice rebuilding full sequences from reads
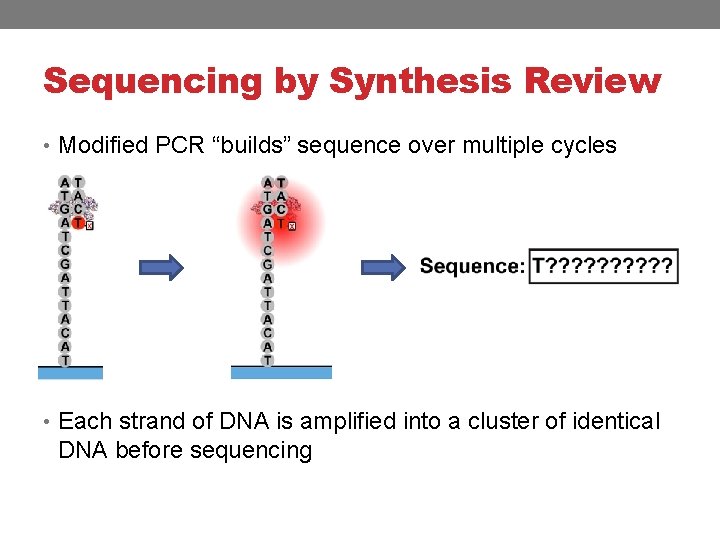
Sequencing by Synthesis Review • Modified PCR “builds” sequence over multiple cycles • Each strand of DNA is amplified into a cluster of identical DNA before sequencing

Sequencing by Synthesis Review • Multiple clusters are sequenced at once • Clusters can be: • From different samples OR from the sample • Short regions OR long regions that have been broken into shorter pieces • Unique tags (indices) identify the source of each cluster • The sequence from each cluster is referred to as a “read”

Before analysis can begin: • Sequence information needs to be stored • FASTA files store sequence information in a text format • Long regions that were broken up for sequencing need to be rebuilt • Assembly rebuilds long regions using overlapping sequences • Alignment rebuilds long regions by matching reads to a reference • “References” are the results from the previous times a genome or region was sequenced. • This can also be called the “consensus” sequence since it is the agreed upon complete version of the sequence.
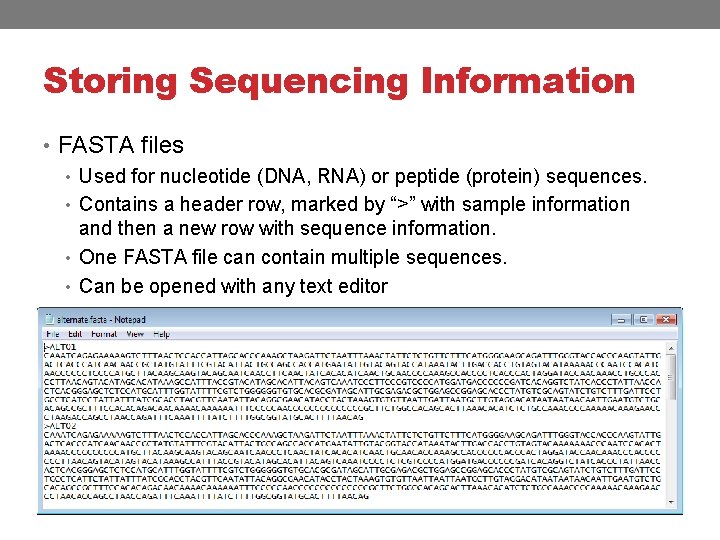
Storing Sequencing Information • FASTA files • Used for nucleotide (DNA, RNA) or peptide (protein) sequences. • Contains a header row, marked by “>” with sample information and then a new row with sequence information. • One FASTA file can contain multiple sequences. • Can be opened with any text editor
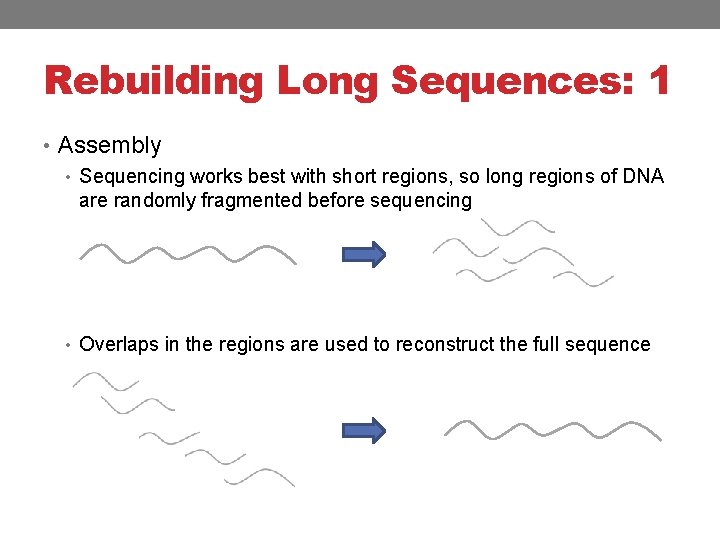
Rebuilding Long Sequences: 1 • Assembly • Sequencing works best with short regions, so long regions of DNA are randomly fragmented before sequencing • Overlaps in the regions are used to reconstruct the full sequence

Assembly Details • DNA is amplified before fragmentation. Lots of copies being randomly fragmented means a lot of overlap. • The more short fragments which overlap with one another allow more certainty that the long region has been correctly assembled. Read 1: ATCCGCATTGAC Read 2: TGACCTAGCGCA OR Read 3: GCAATACGTGAC Read 2: TGACCTAGCGCA ? Read 4: CATTGACCTAG

Practice Assembly • Sequence Processing OR Read Assembly Activity • All groups get only the reads • Think about the following: • How many “reads” were necessary to cover the entire “genome”? • How sure are you of the final sequence? • Are there any regions of ambiguity? • What information would you want to help resolve that ambiguity?
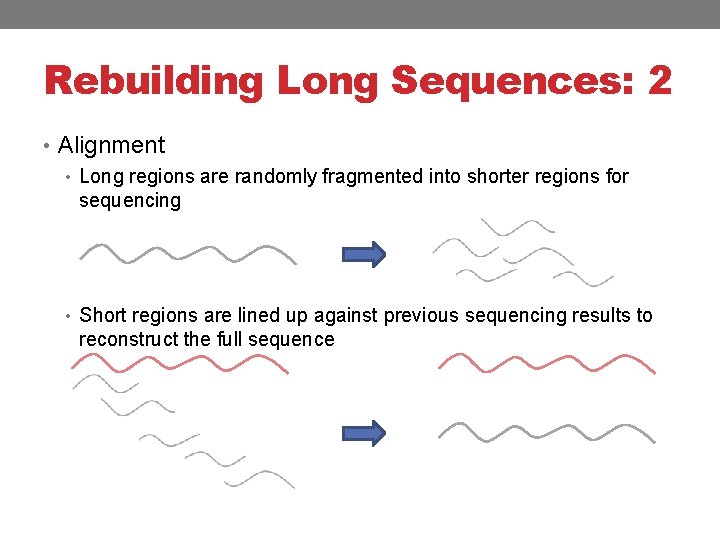
Rebuilding Long Sequences: 2 • Alignment • Long regions are randomly fragmented into shorter regions for sequencing • Short regions are lined up against previous sequencing results to reconstruct the full sequence

Alignment Details • Points of variation between the read and reference are noted and stored in a “Variant Call File” (VCF) • The more short fragments which include a variation, the more certain we can be that variation isn’t just a sequencing error. • Reads can vary from a reference in different ways • Changes in a nucleotide • Insertions • Deletions Reference: Read: ATCCCGGA-TCGTTA |||| || The | indicates ATC-CGGAATCGATA a perfect match

Storing Variation Information • Variant Call File (VCF) • Indicates differences compared to a reference. • Contains header rows, marked by “##”, and a table of variants • Can be opened in text or spreadsheet editors

Practice Aligning • Sequence Processing OR Read Assembly Activity • All groups get reads and a reference copy of the original text • For more practice with alignment: Aligning Short Texts Activity • Think about the following: • How are you deciding on the “best” alignment? • What benefit is there to having multiple “reads” for each text? • Multiple Alignment: • When more than two sequences are being aligned

Evaluating Alignments • Goal: maximize overlap between sequences • Scoring • Way of quantifying overlap so different alignments can be compared • Different scoring systems exist, but a simple one would be • Matches: +1 • Mismatches: -1 • Gaps: -2 • To use this system: Score = (number of matches) – (number of mismatches) – 2*(number of gaps)

Comparing Alignments Score = (number of matches) – (number of mismatches) – 2*(number of gaps) • Alignment 1 Reference: Read: • Alignment 2 Reference: Read: • Alignment 3 Reference: Read: GTCGAATGAAACGATTAA |||| | TCGATTTAACGATTA GTCGAATGAAACGATTAA || |||| TCGATTTAACGATTA GTCGAATGAAACGATTAA |||| | | ||||||| TCGATTTA-ACGATTA
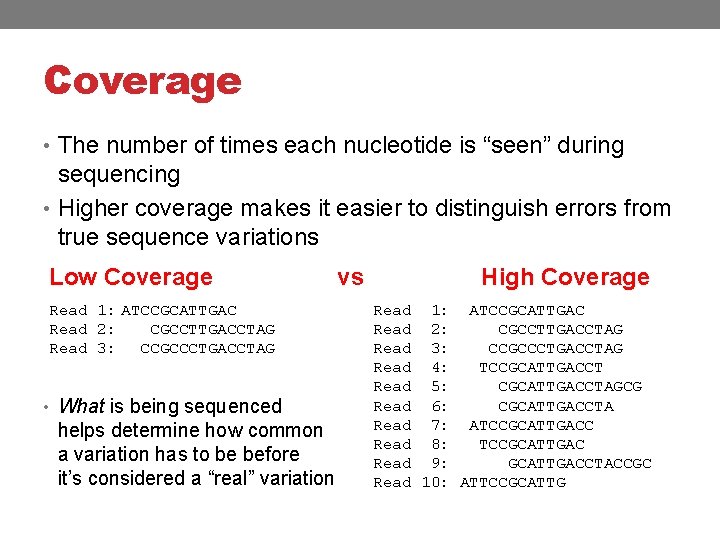
Coverage • The number of times each nucleotide is “seen” during sequencing • Higher coverage makes it easier to distinguish errors from true sequence variations Low Coverage Read 1: ATCCGCATTGAC Read 2: CGCCTTGACCTAG Read 3: CCGCCCTGACCTAG • What is being sequenced helps determine how common a variation has to be before it’s considered a “real” variation vs High Coverage Read 1: ATCCGCATTGAC Read 2: CGCCTTGACCTAG Read 3: CCGCCCTGACCTAG Read 4: TCCGCATTGACCT Read 5: CGCATTGACCTAGCG Read 6: CGCATTGACCTA Read 7: ATCCGCATTGACC Read 8: TCCGCATTGAC Read 9: GCATTGACCTACCGC Read 10: ATTCCGCATTG

Types of Sequencing Analysis • De Novo Sequencing • Used the first time a gene or genome is ever sequenced • Uses assembly to stitch short regions into a longer whole • Resequencing • Used subsequent times a genome is sequenced • Uses alignment to identify short sequences using a reference 16

Compare methods • Sequence Processing OR Read Assembly Activity • Use a different text, provide half the groups a “Reference” sheet • Think about the following: • How long are the “reads”? • How long is the “genome”? • How easy was this task with vs without a “reference” text? • How fast was this task with vs without a “reference” text? • How long are sequencing reads? • How long are genomes? • How easy/fast would using real sequencing data be?

Role of computers in analysis • Computers can: • Automate tasks • Work faster than humans • Process long sequences just as easily as short sequences • Bioinformatics: use of computers for analyzing complex biological data. • Lots of bioinformatics tools exist for you to use in analyzing your sequence 18
- Slides: 18