LECTURE PRESENTATIONS For CAMPBELL BIOLOGY NINTH EDITION Jane

LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 18 Regulation of Gene Expression Lectures by Erin Barley Kathleen Fitzpatrick © 2011 Pearson Education, Inc.

Overview: Gene Expression • Prokaryotes & eukaryotes use different methods to turn genes “on” or “off” • Ultimately gene expression is about efficient use of resources/energy – If you don’t need it, don’t make it © 2011 Pearson Education, Inc.
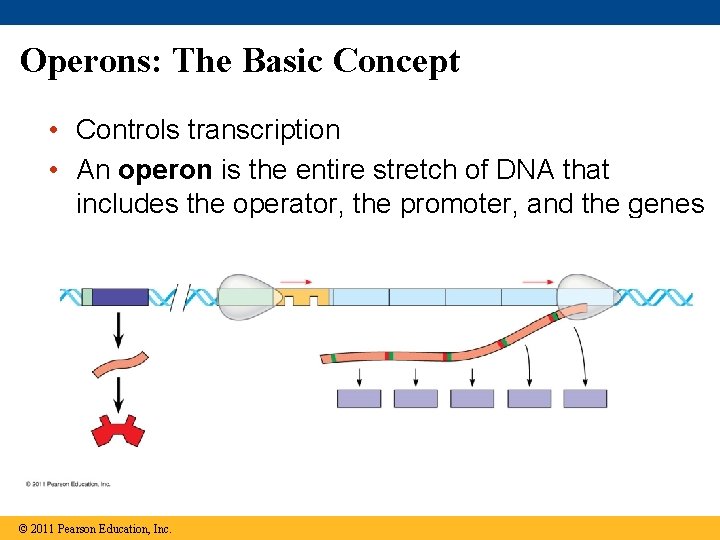
Operons: The Basic Concept • Controls transcription • An operon is the entire stretch of DNA that includes the operator, the promoter, and the genes that they control © 2011 Pearson Education, Inc.

Parts of the operon • Promoter (RNA Polymerase binding site) – Operator (Repressor binding site) • Genes
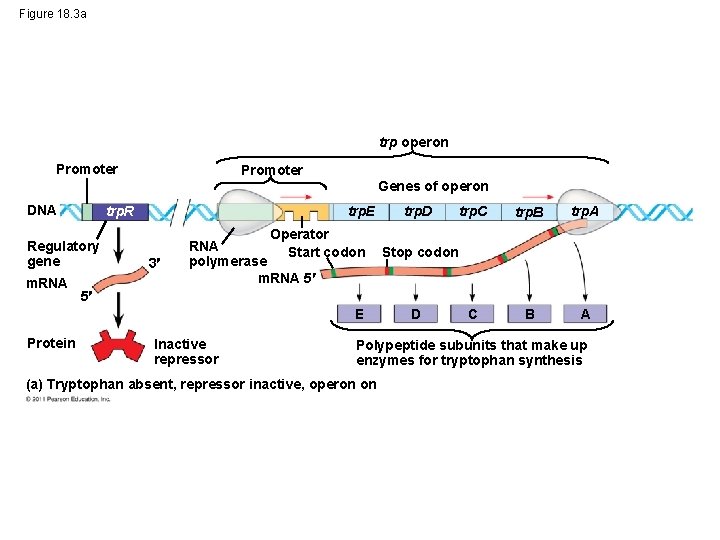
Figure 18. 3 a trp operon Promoter Genes of operon DNA trp. R Regulatory gene m. RNA trp. E 3 Operator RNA Start codon polymerase m. RNA 5 trp. C trp. B trp. A C B A Stop codon 5 E Protein trp. D Inactive repressor D Polypeptide subunits that make up enzymes for tryptophan synthesis (a) Tryptophan absent, repressor inactive, operon on

Other players: • Repressor: A protein that can turn the operon “off” or “on” by binding to the operator, blocking RNA polymerase • regulatory gene: makes the repressor (usually “upstream” from the operon. • Co-repressor: A molecule that binds to the repressor protein to change its shape © 2011 Pearson Education, Inc.
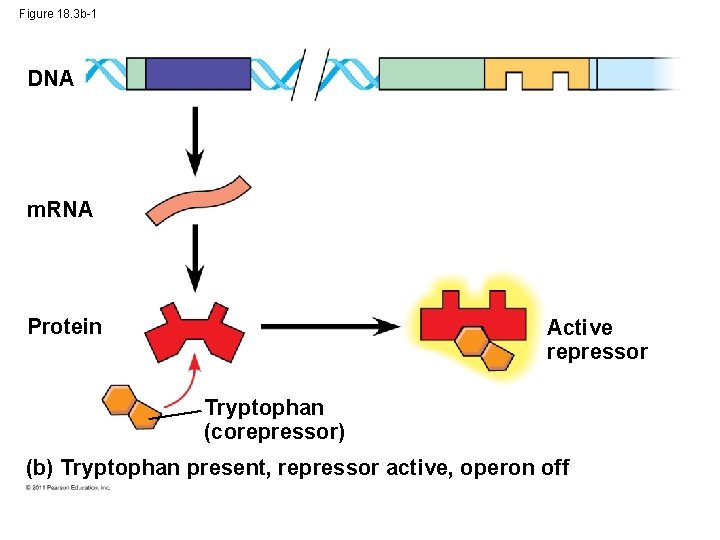
Figure 18. 3 b-1 DNA m. RNA Protein Active repressor Tryptophan (corepressor) (b) Tryptophan present, repressor active, operon off
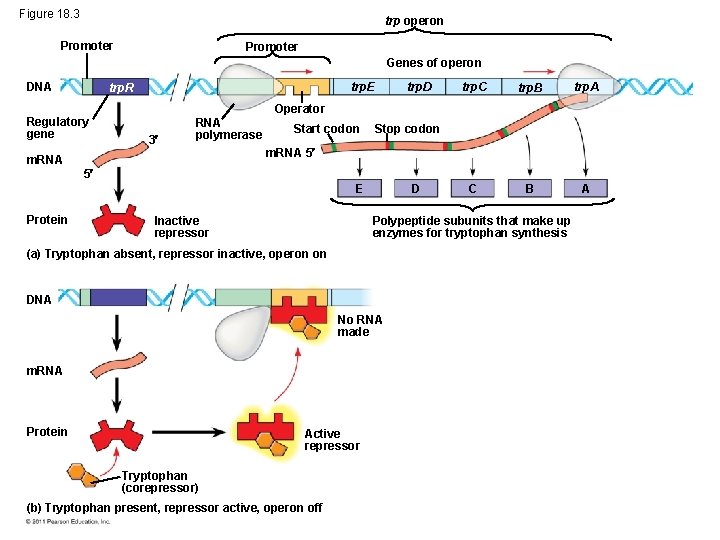
Figure 18. 3 trp operon Promoter Genes of operon DNA trp. E trp. R Regulatory gene trp. D trp. C trp. B trp. A C B A Operator 3 RNA polymerase m. RNA Start codon Stop codon m. RNA 5 5 E Protein Inactive repressor D Polypeptide subunits that make up enzymes for tryptophan synthesis (a) Tryptophan absent, repressor inactive, operon on DNA No RNA made m. RNA Protein Active repressor Tryptophan (corepressor) (b) Tryptophan present, repressor active, operon off

Repressible and Inducible Operons: Negative or Positive Feedback? • Repressible operon: default is “on; ” binding of a repressor shuts off transcription (ex: trp operon) • inducible operon default is “off”; a molecule called an inducer inactivates the repressor and turns on transcription (ex. lac operon) © 2011 Pearson Education, Inc.
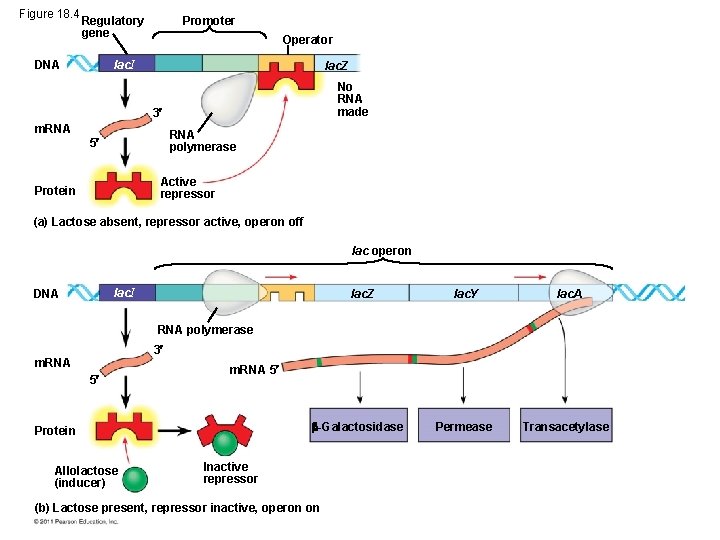
Figure 18. 4 Regulatory gene DNA Promoter Operator lac. I lac. Z No RNA made 3 m. RNA polymerase 5 Active repressor Protein (a) Lactose absent, repressor active, operon off lac operon lac. I DNA lac. Z lac. Y lac. A Permease Transacetylase RNA polymerase 3 m. RNA 5 -Galactosidase Protein Allolactose (inducer) Inactive repressor (b) Lactose present, repressor inactive, operon on

When do we see inducible vs. repressible feedback? • Inducible enzymes usually in catabolic pathways; • Repressible enzymes usually function in anabolic pathways © 2011 Pearson Education, Inc.

Eukaryotic gene expression is regulated at many stages • • Chromatin modification Transcription RNA Processing Transport to cytoplasm Degradation of m. RNA Protein processing Degradation of protein Transport to cellular destination © 2011 Pearson Education, Inc.
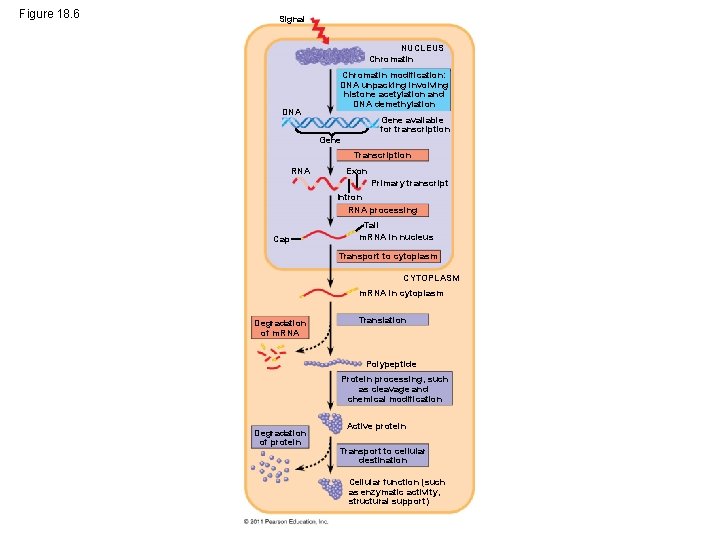
Figure 18. 6 Signal NUCLEUS Chromatin DNA Chromatin modification: DNA unpacking involving histone acetylation and DNA demethylation Gene available for transcription Gene Transcription RNA Exon Primary transcript Intron RNA processing Cap Tail m. RNA in nucleus Transport to cytoplasm CYTOPLASM m. RNA in cytoplasm Degradation of m. RNA Translation Polypeptide Protein processing, such as cleavage and chemical modification Degradation of protein Active protein Transport to cellular destination Cellular function (such as enzymatic activity, structural support)

Differential Gene Expression • Cell differentiation results from gene expression, • Abnormalities in gene expression can lead to diseases including cancer © 2011 Pearson Education, Inc.

Regulation of Chromatin Structure • Histone proteins, part of Chromatin are modified to allow for gene expression 1) Histone acetylation: acetyl groups are attached to positively charged lysines in histone tails 2) Phosphorylation: phosphate groups added next to methyl groups • This “loosens” the structure of the chromatin and allows for transcription. • (See video: DNA Packing) © 2011 Pearson Education, Inc.
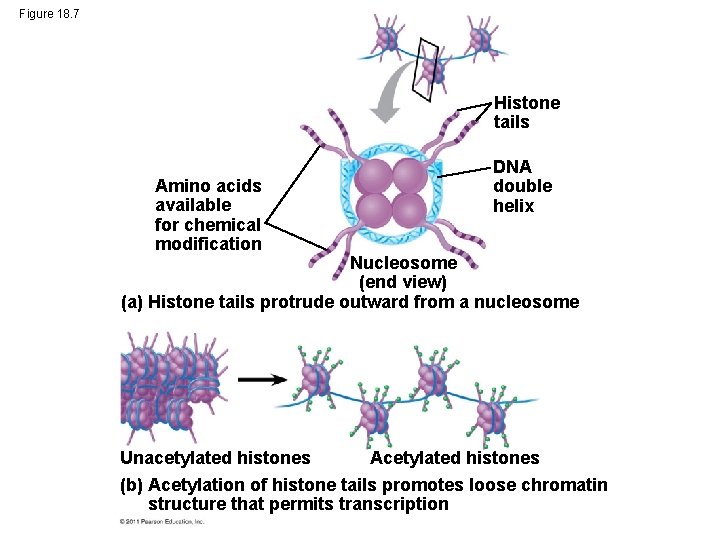
Figure 18. 7 Histone tails Amino acids available for chemical modification DNA double helix Nucleosome (end view) (a) Histone tails protrude outward from a nucleosome Acetylated histones Unacetylated histones (b) Acetylation of histone tails promotes loose chromatin structure that permits transcription

DNA Methylation • reduces transcription • addition of methyl groups, • can cause long-term inactivation of genes in cellular differentiation • Phosphorylation counteracts methylation © 2011 Pearson Education, Inc.

Epigenetic Inheritance • Although the chromatin modifications just discussed do not alter DNA sequence, they may be passed to future generations of cells • The inheritance of traits transmitted by mechanisms not directly involving the nucleotide sequence is called epigenetic inheritance © 2011 Pearson Education, Inc.

Regulation of Transcription Initiation • Chromatin-modifying enzymes provide initial control of gene expression by making a region of DNA either more or less able to bind the transcription machinery © 2011 Pearson Education, Inc.
- Slides: 19