LECTURE PRESENTATIONS For CAMPBELL BIOLOGY NINTH EDITION Jane

LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 18 Regulation of Gene Expression Lectures by Erin Barley Kathleen Fitzpatrick © 2011 Pearson Education, Inc.

Gene expression: Bacteria vs. Eukaryotes • prokaryotes and eukaryotes alter gene expression in response to their changing environment – gene expression = refers to the entire process whereby genetic information is decoded into a protein • prokaryotes and eukaryotes carry out gene expression in similar ways – transcription using an RNA polymerase – translation using ribosomes • but there are some differences: – – – 1. RNA polymerases differ – only one in prokaryotes; 3 in eukaryotes 2. transcription factors used by eukaryotes 3. transcription is terminated differently in prokaryotes vs. eukaryotes 4. ribosomes – bacterial ones are smaller 5. lack of compartmentalization in bacteria – transcribe and translate at the same time

So what is a gene? • unit of inheritance • located on chromosomes • region of specific nucleotide sequence located along the length of DNA • DNA sequence that codes for a specific sequence of amino acids • BUT: some DNA sequences are NEVER translated – e. g. r. RNA and t. RNA are transcribed but not translated into anything • so a gene is a region of DNA that is either – 1. translated into a sequence of amino acids (polypeptide) functional protein – OR – 2. transcribed into a RNA molecule

So what is a gene? • molecular components of a gene: – A. coding sequences - eukaryotes have introns within their coding sequence – B. promoter – C. enhancers – found in eukaryotes – D. UTRs – found in eukaryotes – E. poly-adenylation sequence – found within the eukaryotic 3’ UTR

Overview: Conducting the Genetic Orchestra • genetic and biochemical work in bacteria identified two things – 1. protein-binding regulatory sequences associated with genes – 2. proteins that can bind these regulatory sequences – either activating or repressing gene expression • these two components underlie the ability of both prokaryotic and eukaryotic cells to turn genes on and off
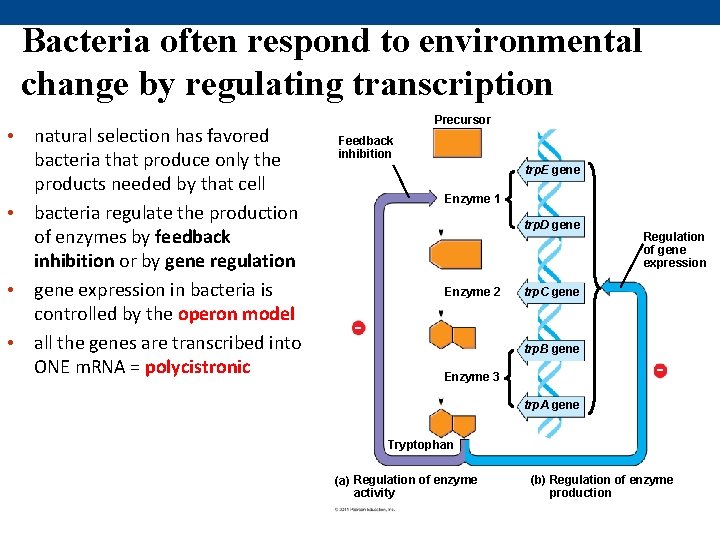
Bacteria often respond to environmental change by regulating transcription • natural selection has favored bacteria that produce only the products needed by that cell • bacteria regulate the production of enzymes by feedback inhibition or by gene regulation • gene expression in bacteria is controlled by the operon model • all the genes are transcribed into ONE m. RNA = polycistronic Precursor Feedback inhibition trp. E gene Enzyme 1 trp. D gene Enzyme 2 Regulation of gene expression trp. C gene trp. B gene Enzyme 3 trp. A gene Tryptophan (a) Regulation of enzyme activity (b) Regulation of enzyme production

Operons: Definitions & Concepts • bacteria group functionally related genes so they can be under coordinated control by a single “on-off regulatory switch” • the regulatory “switch” is a segment of DNA called an operator – a binding site for “regulatory factors” that determine whether RNA polymerase binds the nearby promoter • the operator can be controlled by proteins or nutrients – e. g. can be switched off by a protein called a repressor that binds to the operator and blocks RNA polymerase binding to the promoter – repressor is the product of a separate regulatory gene – repressor can be in an active or inactive form, depending on the presence of other molecules • co-repressor is a molecule that cooperates with a repressor protein to switch an operon off – e. g. the amino acid tryptophan
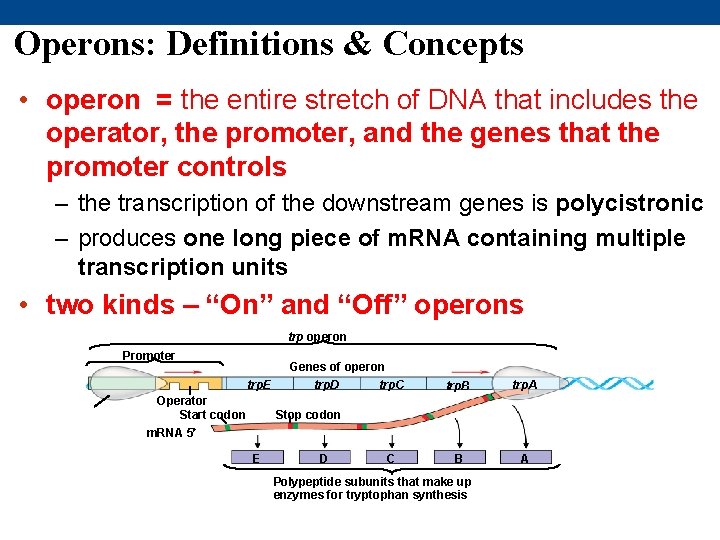
Operons: Definitions & Concepts • operon = the entire stretch of DNA that includes the operator, the promoter, and the genes that the promoter controls – the transcription of the downstream genes is polycistronic – produces one long piece of m. RNA containing multiple transcription units • two kinds – “On” and “Off” operons trp operon Promoter trp. E Operator Start codon m. RNA 5 Genes of operon trp. D trp. C trp. B trp. A B A Stop codon E D C Polypeptide subunits that make up enzymes for tryptophan synthesis

Repressible and Inducible Operons: Two Types of Negative Gene Regulation • OFF operon = repressible operon is one that is usually on but is turned OFF by a repressor – e. g. the trp operon is a repressible operon • ON operon = inducible operon is one that is usually off but is turned ON by an inducer – e. g. lac operon is an inducible operon

The trp Operon: Repressible Operons • E. coli can synthesize the amino acid tryptophan when it is absent from the growth media • by default the trp operon is on and the genes for tryptophan synthesis are transcribed • comprised of the: – 1. operator – capable of binding a repressor protein – 2. genes of the operon – for synthesizing tryptophan when it is missing from the growth media • also a regulatory gene = trp. R – expressed whether tryptophan is absent or present – NOT PART OF THE OPERON!!!!
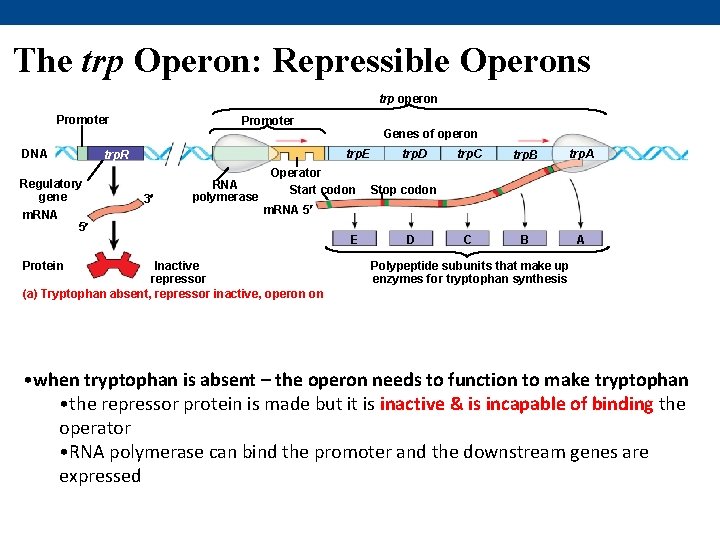
The trp Operon: Repressible Operons trp operon Promoter DNA Protein Genes of operon trp. E trp. R Regulatory gene m. RNA Promoter 3 RNA polymerase Operator Start codon trp. D trp. C trp. B trp. A C B A Stop codon m. RNA 5 5 Inactive repressor (a) Tryptophan absent, repressor inactive, operon on E D Polypeptide subunits that make up enzymes for tryptophan synthesis • when tryptophan is absent – the operon needs to function to make tryptophan • the repressor protein is made but it is inactive & is incapable of binding the operator • RNA polymerase can bind the promoter and the downstream genes are expressed
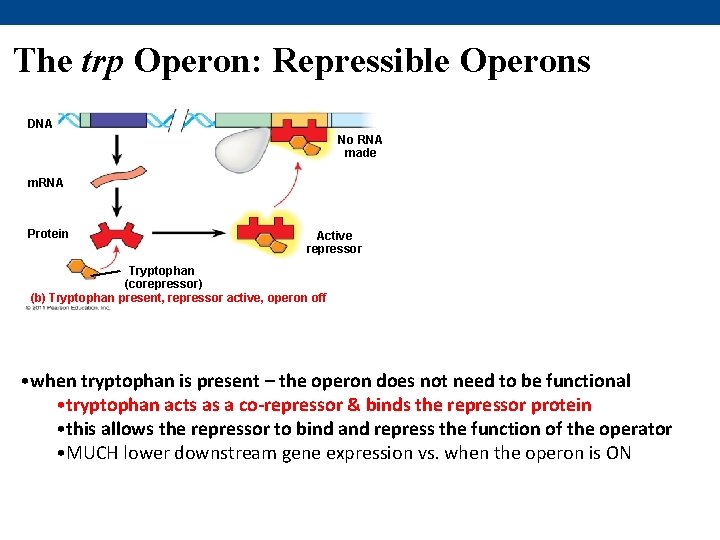
The trp Operon: Repressible Operons DNA No RNA made m. RNA Protein Active repressor Tryptophan (corepressor) (b) Tryptophan present, repressor active, operon off • when tryptophan is present – the operon does not need to be functional • tryptophan acts as a co-repressor & binds the repressor protein • this allows the repressor to bind and repress the function of the operator • MUCH lower downstream gene expression vs. when the operon is ON

The lac Operon: Inducible Operons • proposed by Francois Jacob and Jacques Monod - 1960 s • E. coli can use glucose and other sugars (such as lactose) as their sole source of carbon and energy • the normal situation is for the bacteria to use glucose – levels of a bacterial enzyme called beta-galactosidase (lactose breakdown) are very low • when lactose is given to the bacteria – b-Gal levels increase – said to be induced

The lac Operon: Inducible Operons • the lac operon is an inducible operon – contains genes that code for enzymes used in the hydrolysis and metabolism of lactose • when E. coli are grown with glucose – no need for the enzymes of the lac operon since there is no lactose in the medium – so the operon is turned OFF • but with media containing lactose – need to turn the operon ON to make the enzymes for metabolizing and using lactose

The lac Operon: Inducible Operons • genes of the lac-operon: – – – 1. lac. Z gene = beta-galactosidase – splits the lactose into glucose and galactose 2. lac. Y gene 3. lac. A gene 4. operator = binds the repressor 5. promoter = binds RNA polymerase II & transcription factors PLUS; lac. I gene = codes for a lac repressor NOT PART OF THE OPERON!!! • THIS IS IMPORTANT!!! - without any outside control - the lac regulatory gene lac. I is constitutively active and acts to eventually switch the lac operon OFF – through the constitutive production of a lac repressor protein • a molecule called an inducer is needed to inactivate the repressor to turn the lac operon ON
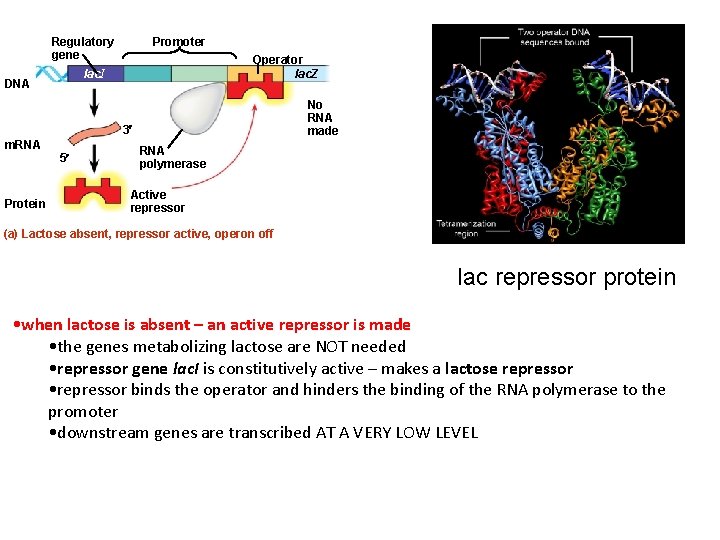
Regulatory gene Promoter Operator lac. Z lac. I DNA No RNA made 3 m. RNA Protein 5 RNA polymerase Active repressor (a) Lactose absent, repressor active, operon off lac repressor protein • when lactose is absent – an active repressor is made • the genes metabolizing lactose are NOT needed • repressor gene lac. I is constitutively active – makes a lactose repressor • repressor binds the operator and hinders the binding of the RNA polymerase to the promoter • downstream genes are transcribed AT A VERY LOW LEVEL
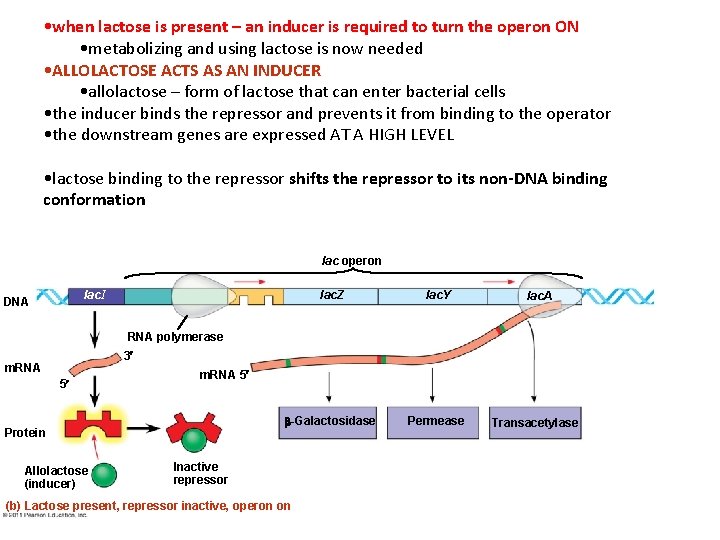
• when lactose is present – an inducer is required to turn the operon ON • metabolizing and using lactose is now needed • ALLOLACTOSE ACTS AS AN INDUCER • allolactose – form of lactose that can enter bacterial cells • the inducer binds the repressor and prevents it from binding to the operator • the downstream genes are expressed AT A HIGH LEVEL • lactose binding to the repressor shifts the repressor to its non-DNA binding conformation lac operon lac. I DNA lac. Z lac. Y -Galactosidase Permease lac. A RNA polymerase 3 m. RNA 5 Protein Allolactose (inducer) Inactive repressor (b) Lactose present, repressor inactive, operon on Transacetylase

• in nature – the inducer of the lab operon is a lactose derivative (allolactose) • in the lab – other inducers can be used to turn the operon on – e. g. IPTG = isopropyl-b-D-thiogalactoside – IPTG is NOT a broken down by -Gal • BUT we can give the bacteria a specific b-Gal substrate that will turn colors once it is broken down – X-Gal – turns blue with broken down by b-gal enzyme – used to identify bacteria containing cloned genes – can insert a customized gene into plasmids containing a version of the lac operon with a b-gal gene • insertion of your desired gene INTO the plasmid disrupts -gal expression • inability to breakdown X-Gal – colonies are white

• inducible enzymes usually function in catabolic pathways – their synthesis is induced by a chemical signal • repressible enzymes usually function in anabolic pathways – their synthesis is repressed by high levels of the end product • regulation of the trp and lac operons involves negative control of genes because operons are switched off by the active form of the repressor

Positive Gene Regulation: CAP proteins • some operons are also subject to positive control • when bacteria are given both lactose AND glucose - the bacteria will use glucose – the enzymes for glycolysis are continually present in bacteria • when lactose is present and glucose is short supply – it makes the enzymes for lactose metabolism • how does the bacteria sense the low levels of glucose? ?
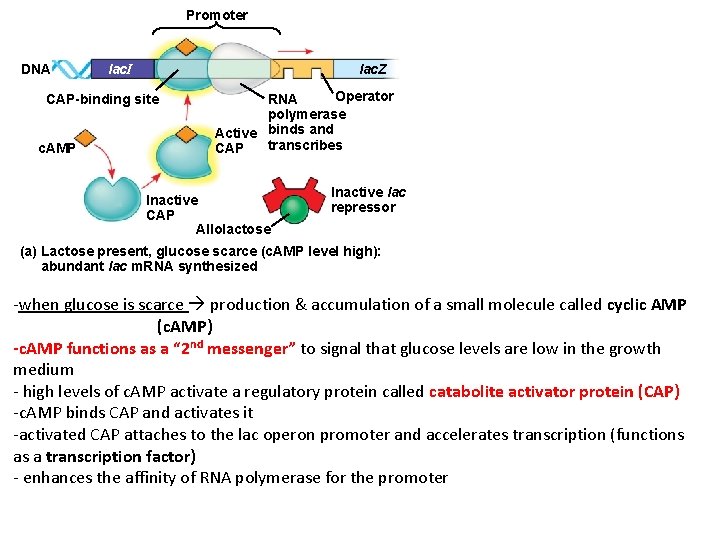
Promoter DNA lac. I lac. Z CAP-binding site c. AMP Operator RNA polymerase Active binds and transcribes CAP Inactive CAP Allolactose Inactive lac repressor (a) Lactose present, glucose scarce (c. AMP level high): abundant lac m. RNA synthesized -when glucose is scarce production & accumulation of a small molecule called cyclic AMP (c. AMP) -c. AMP functions as a “ 2 nd messenger” to signal that glucose levels are low in the growth medium - high levels of c. AMP activate a regulatory protein called catabolite activator protein (CAP) -c. AMP binds CAP and activates it -activated CAP attaches to the lac operon promoter and accelerates transcription (functions as a transcription factor) - enhances the affinity of RNA polymerase for the promoter

• CAP helps regulate other operons that encode enzymes used in catabolic pathways • when glucose levels are low and lactose levels are high – 1. lactose binds the lactose repressor and prevents it from binding the operator and inhibiting gene transcription = genes for lactose metabolism are made – 2. c. AMP activation of CAP and its binding to the lac promoter increases transcription = lac operon genes are made at a higher rate – CAP controls the rate at which the lac operon genes are made

• when glucose levels increase and lactose levels decrease – 1. CAP activation will eventually decrease and so will its enhancement of transcription – 2. the lactose repressor is now able to bind the operator and inhibit transcription

• so the lac operon is actually under dual control as lactose increases and glucose decreases: – positive control – as levels of c. AMP rise – so does CAP activation and the activity of the lac operon – negative control – as repressor activity decreases & the activity of the lac operon increases – THEREFORE: it is the allosteric state of the lac repressor that determines if transcription happens – it is the presence of CAP that controls the rate at which transcription will happen

GO HOME & STUDY!!! Next Lecture EUKARYOTIC GENE REGULATION
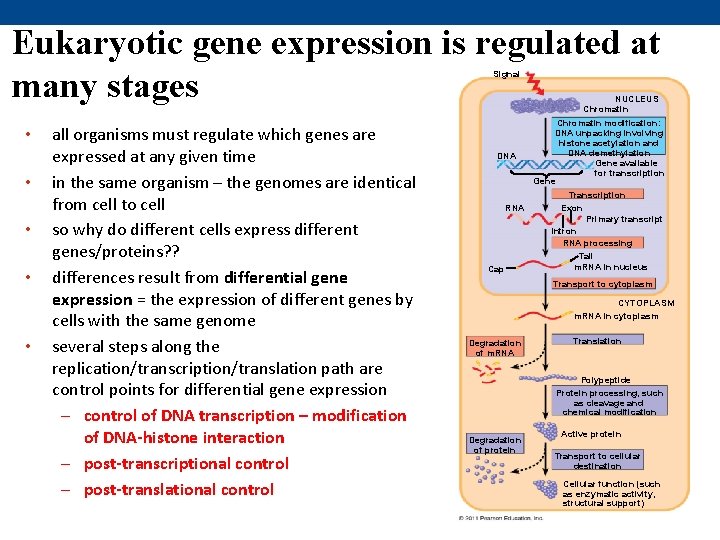
Eukaryotic gene expression is regulated at many stages Signal NUCLEUS Chromatin • • • all organisms must regulate which genes are expressed at any given time in the same organism – the genomes are identical from cell to cell so why do different cells express different genes/proteins? ? differences result from differential gene expression = the expression of different genes by cells with the same genome several steps along the replication/transcription/translation path are control points for differential gene expression – control of DNA transcription – modification of DNA-histone interaction – post-transcriptional control – post-translational control DNA Chromatin modification: DNA unpacking involving histone acetylation and DNA demethylation Gene available for transcription Gene Transcription RNA Cap Exon Primary transcript Intron RNA processing Tail m. RNA in nucleus Transport to cytoplasm CYTOPLASM m. RNA in cytoplasm Degradation of m. RNA Translation Polypeptide Protein processing, such as cleavage and chemical modification Degradation of protein Active protein Transport to cellular destination Cellular function (such as enzymatic activity, structural support)
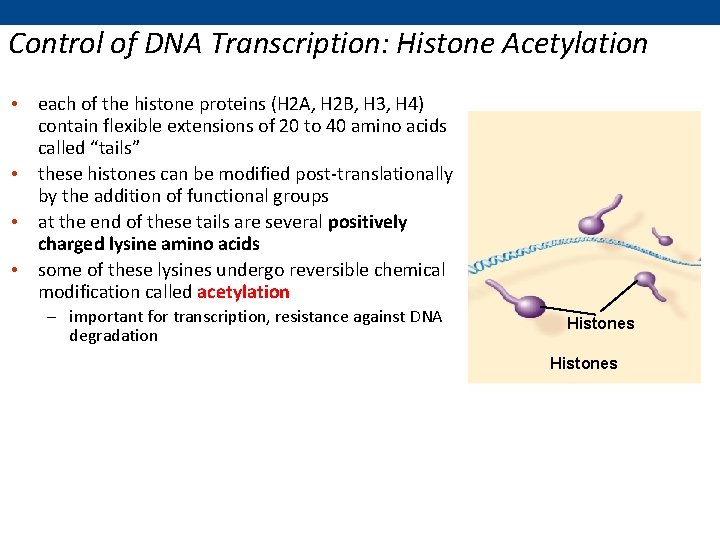
Control of DNA Transcription: Histone Acetylation • each of the histone proteins (H 2 A, H 2 B, H 3, H 4) contain flexible extensions of 20 to 40 amino acids called “tails” • these histones can be modified post-translationally by the addition of functional groups • at the end of these tails are several positively charged lysine amino acids • some of these lysines undergo reversible chemical modification called acetylation – important for transcription, resistance against DNA degradation Histones

Acetylation and Deacetylation of Histones • numerous post-translational modifications can be done to histone proteins – affects how the DNA-histone interacts and ultimately affects the transcription of the DNA • some histone lysines undergo reversible chemical modifications called acetylation and deacteylation • acetylation = transfer of an acetyl group onto the NH 2 terminus of an amino acid – for histones – performed by a family of enzymes called histone acteyltransferases (HATs) • acetylation neutralizes the +ve charge of these lysines – its interaction with the DNA is eliminated – the DNA becomes less tightly associated with the histone – results in better access for the transcriptional machinery to the DNA
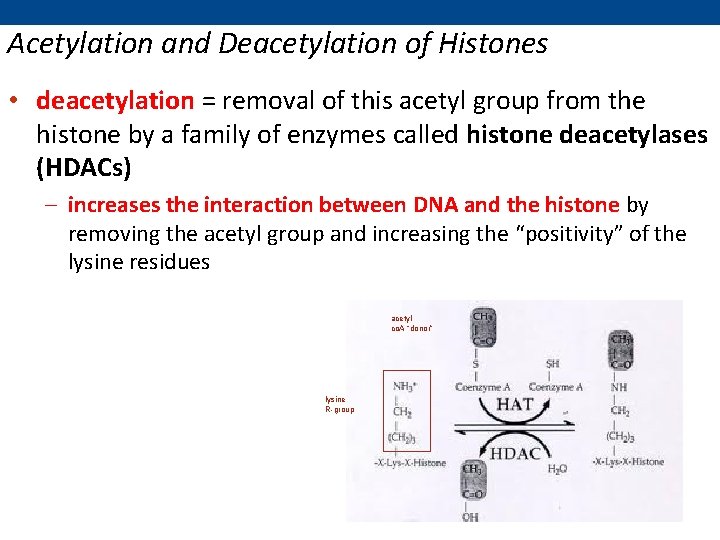
Acetylation and Deacetylation of Histones • deacetylation = removal of this acetyl group from the histone by a family of enzymes called histone deacetylases (HDACs) – increases the interaction between DNA and the histone by removing the acetyl group and increasing the “positivity” of the lysine residues acetyl co. A “donor” lysine R-group
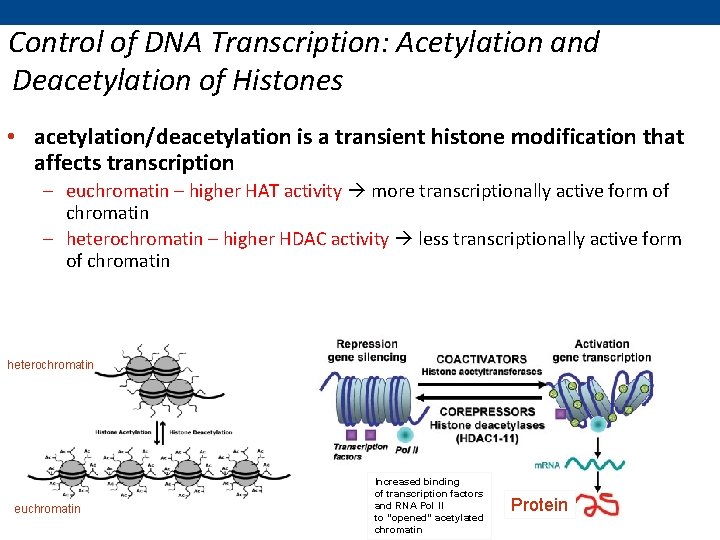
Control of DNA Transcription: Acetylation and Deacetylation of Histones • acetylation/deacetylation is a transient histone modification that affects transcription – euchromatin – higher HAT activity more transcriptionally active form of chromatin – heterochromatin – higher HDAC activity less transcriptionally active form of chromatin heterochromatin euchromatin Increased binding of transcription factors and RNA Pol II to “opened” acetylated chromatin Protein
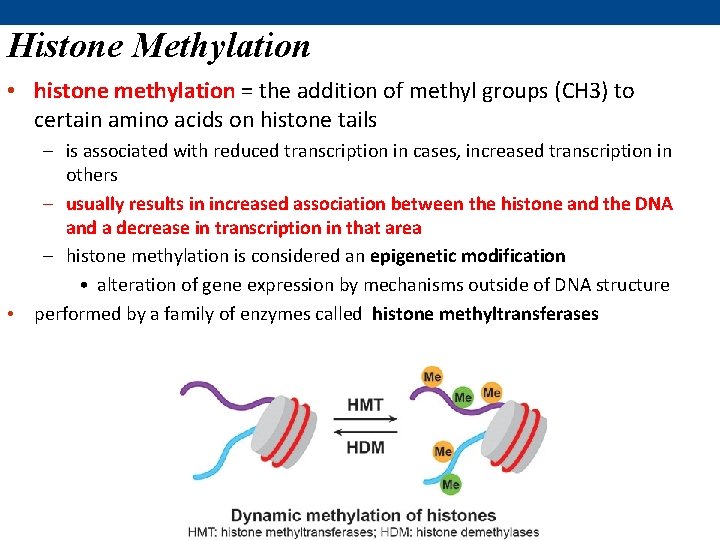
Histone Methylation • histone methylation = the addition of methyl groups (CH 3) to certain amino acids on histone tails – is associated with reduced transcription in cases, increased transcription in others – usually results in increased association between the histone and the DNA and a decrease in transcription in that area – histone methylation is considered an epigenetic modification • alteration of gene expression by mechanisms outside of DNA structure • performed by a family of enzymes called histone methyltransferases

DNA Methylation • in addition to histones – methyl groups can be attached to certain DNA bases = DNA methylation – usually cytosine – done by a different set of enzymes than those that methylate histones = DNA methyltransferases – is associated with reduced transcription in some species – i. e. the more methylated, the more inactive the gene • DNA methylation essential for long-term inactivation of genes during cellular differentiation – DNA methylation can last through several rounds of replication – when a methylated DNA sequence is replicated – the daughter strand is methylated too

Regulation of Transcription Initiation • chromatin-modifying enzymes provide initial control of gene expression by making a region of DNA either more or less able to bind the transcription machinery • additional transcriptional levels are also found – enhancers – promoters
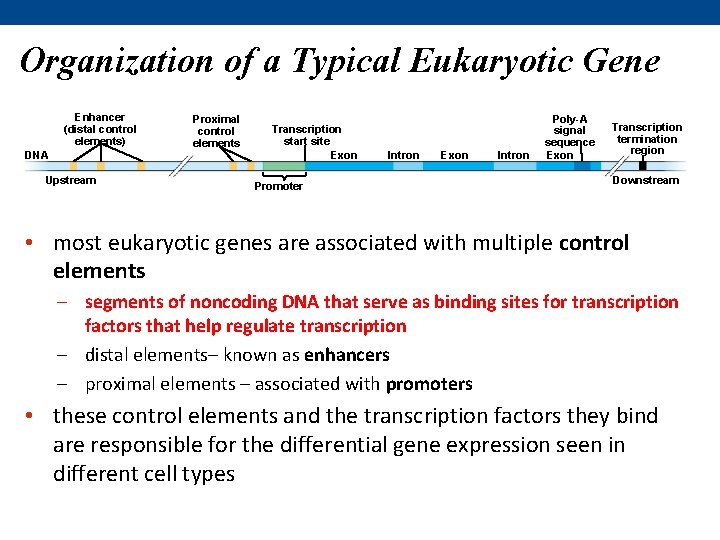
Organization of a Typical Eukaryotic Gene Enhancer (distal control elements) DNA Upstream Proximal control elements Transcription start site Exon Promoter Intron Exon Intron Poly-A signal sequence Exon Transcription termination region Downstream • most eukaryotic genes are associated with multiple control elements – segments of noncoding DNA that serve as binding sites for transcription factors that help regulate transcription – distal elements– known as enhancers – proximal elements – associated with promoters • these control elements and the transcription factors they bind are responsible for the differential gene expression seen in different cell types
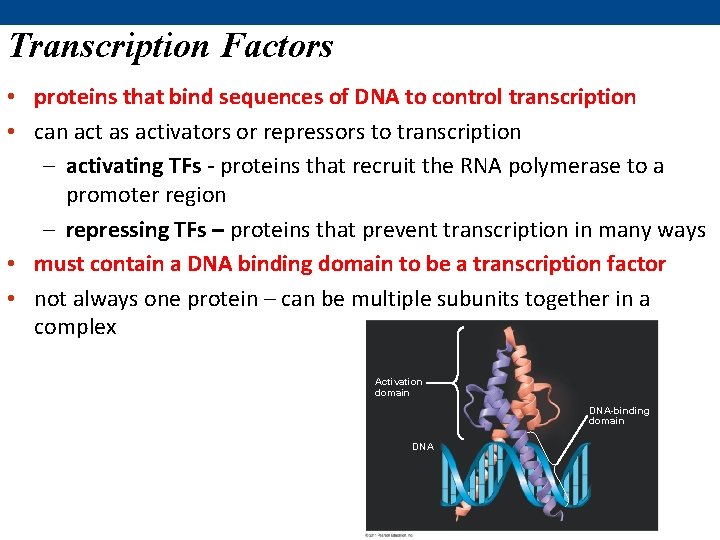
Transcription Factors • proteins that bind sequences of DNA to control transcription • can act as activators or repressors to transcription – activating TFs - proteins that recruit the RNA polymerase to a promoter region – repressing TFs – proteins that prevent transcription in many ways • must contain a DNA binding domain to be a transcription factor • not always one protein – can be multiple subunits together in a complex Activation domain DNA-binding domain DNA
Transcription Factors • two broad categories: – 1. general transcription factors – 2. specific transcription factors Activation domain DNA-binding domain DNA

Transcription Factors • two broad categories: – 1. general transcription factors are essential for the transcription of all proteincoding genes • assist the RNA polymerase in binding the promoter – only give a low level of transcription!! • activity is enhanced by specific transcription factors – 2. specific transcription factors control the high-level, differential expression of specific genes within a specific cell type • • bind the promoter and enhancer regions of a gene can function to activate or repress transcription e. g. Runx-2 – transcription factor that is found in osteoblasts directs the expression of several osteogenic genes involved in making bone
Promoters • sequence of DNA located immediately upstream of the transcription start site • promotes transcription of DNA into RNA • site of RNA polymerase binding in both prokaryotes and eukaryotes • contain sequences for the binding of RNA polymerase and sequences for the binding of transcription factors • in eukaryotes – promoter is made up of several regions of DNA
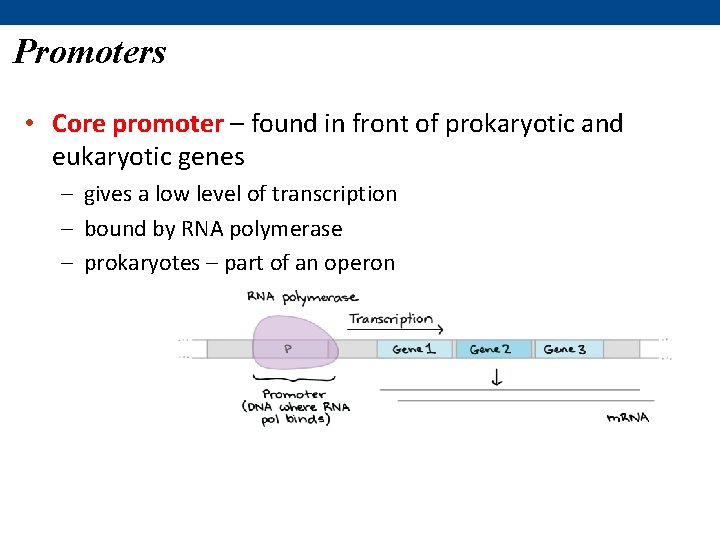
Promoters • Core promoter – found in front of prokaryotic and eukaryotic genes – gives a low level of transcription – bound by RNA polymerase – prokaryotes – part of an operon

Promoters • eukaryotic promoters have additional regions in their promoter called proximal promoter elements/promoter proximal elements (PPEs) – found in addition to the core promoter in many eukaryotic genes – gives regulated transcription – e. g. in certain cells at certain times

Promoters • initial work done in bacteria – found two kinds of DNA sequences controlling transcription • 1. those that are found in the promoters of all bacterial genes – found in what is called the core promoter • 2. those that are found in a more limited number of genes that respond to a specific signal • core bacterial promoter: binds RNA polymerase and an associated sigma factor (part of the RNA polymerase complex) – 10 nucleotides upstream from the start site of transcription is a key region = TATA box (consensus sequence TATAAT) – the core promoter binds a sigma factor called s 70

Promoters • eukaryotic promoters: more complicated than prokaryotic promoters – DEFINITION: several DNA sequences that binds the RNA polymerase II and transcription factors – core promoter required plus additional upstream DNA sequences that regulate transcription – Eukaryotic promoter: – 1. sequences involved in the basic process of transcription = core promoter – 2. sequences active in a particular tissue type or in response to a specific signal – regulated transcription
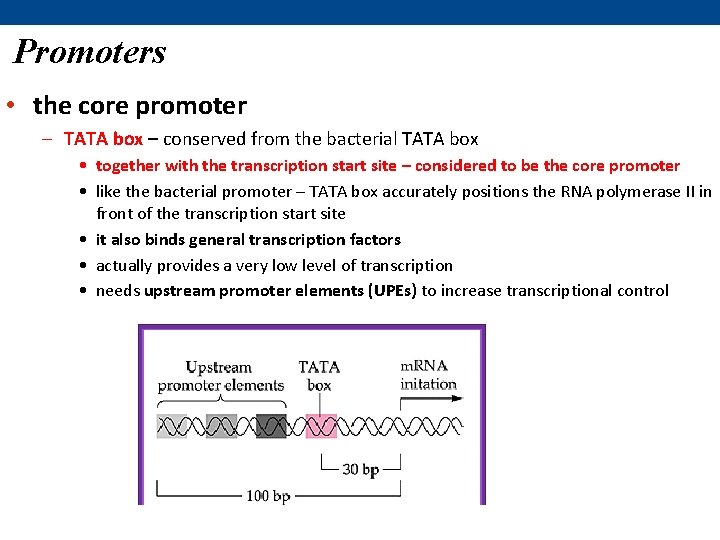
Promoters • the core promoter – TATA box – conserved from the bacterial TATA box • together with the transcription start site – considered to be the core promoter • like the bacterial promoter – TATA box accurately positions the RNA polymerase II in front of the transcription start site • it also binds general transcription factors • actually provides a very low level of transcription • needs upstream promoter elements (UPEs) to increase transcriptional control
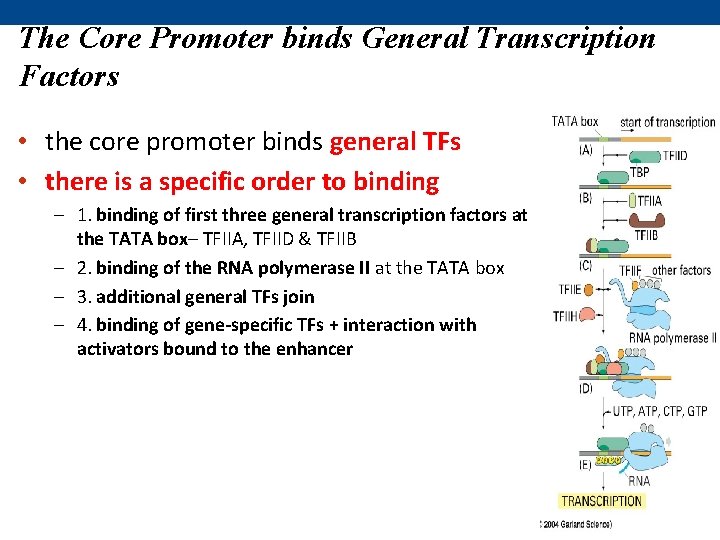
The Core Promoter binds General Transcription Factors • the core promoter binds general TFs • there is a specific order to binding – 1. binding of first three general transcription factors at the TATA box– TFIIA, TFIID & TFIIB – 2. binding of the RNA polymerase II at the TATA box – 3. additional general TFs join – 4. binding of gene-specific TFs + interaction with activators bound to the enhancer

Additional Control Elements for Transcription • core promoter is found in front of genes that are expressed at a high level – e. g. cell cycle genes • eukaryotic genes have additional control elements to allow for controlled expression in specific cell types • 1. promoter proximal elements (PPEs) • 2. enhancer sequence elements or enhancers
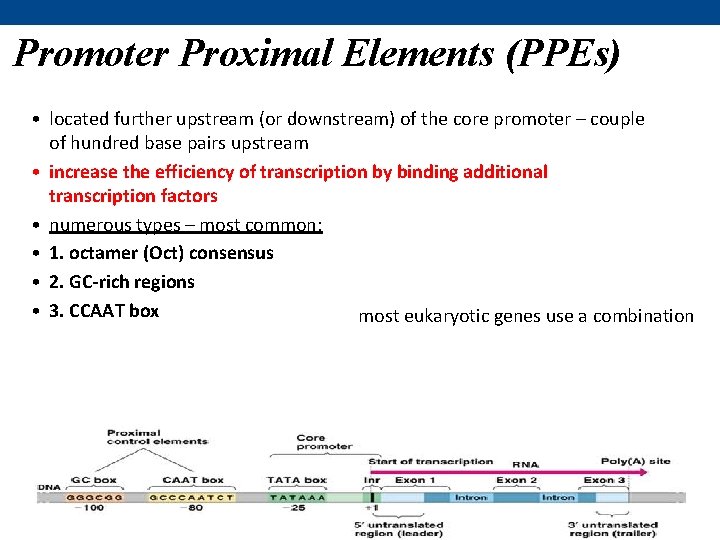
Promoter Proximal Elements (PPEs) • located further upstream (or downstream) of the core promoter – couple of hundred base pairs upstream • increase the efficiency of transcription by binding additional transcription factors • numerous types – most common: • 1. octamer (Oct) consensus • 2. GC-rich regions • 3. CCAAT box most eukaryotic genes use a combination
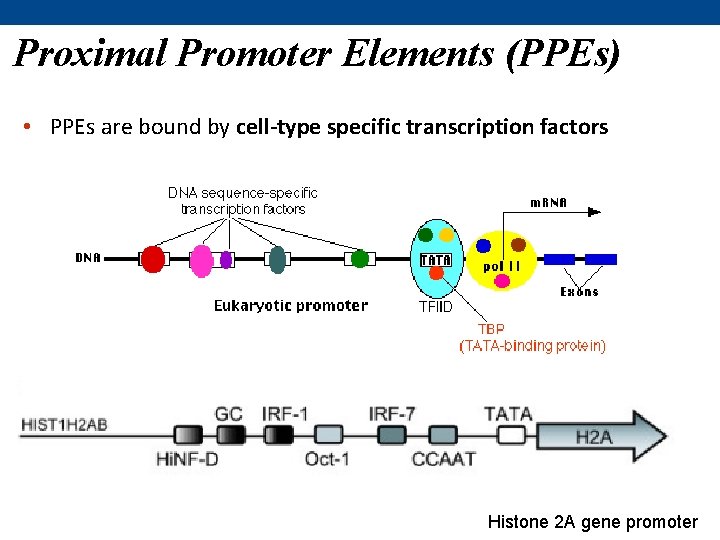
Proximal Promoter Elements (PPEs) • PPEs are bound by cell-type specific transcription factors Histone 2 A gene promoter
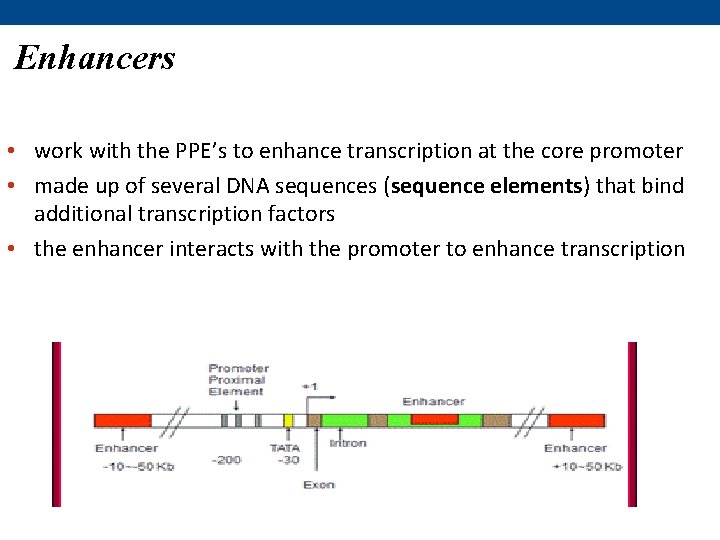
Enhancers • work with the PPE’s to enhance transcription at the core promoter • made up of several DNA sequences (sequence elements) that bind additional transcription factors • the enhancer interacts with the promoter to enhance transcription

Enhancers & their Transcription Factors • transcription factors that bind enhancers are called activators – positively acting transcription factors • activators = proteins that bind to DNA sequences and stimulate/activate transcription of a gene – also called trans-acting factors – can act on adjacent chromosomes • just like transcription factors activators have two domains Activation domain DNA-binding domain DNA 1. DNA binding domain 2. activation domain - site that activates transcription by helping to form the transcription initiation complex

Eukaryotic gene elements: a summary • so the typical eukaryotic gene is controlled by up to 3 distinct control elements – 1. core promoter itself – upstream of the transcription start site • for basic transcriptional control – 2. promoter proximal elements (PPEs) located close to the promoter • required for efficient transcription in any cell – increase transcription rate – 3. distant elements called Enhancers
Next class: Post-transcriptional and translational control

Transcription Initiation • transcription can happen as long as the core promoter is present – but transcription rates will be very low • so efficient transcription of eukaryotic genes requires: the activity of the core promoter, PPEs, enhancers and a multitude of transcription factors together with the RNA polymerase II – RNA polymerase I and III activity is subject to similar kinds of regulation – not as extensive as RNA polymerase II • these components come together to form a transcription initiation complex
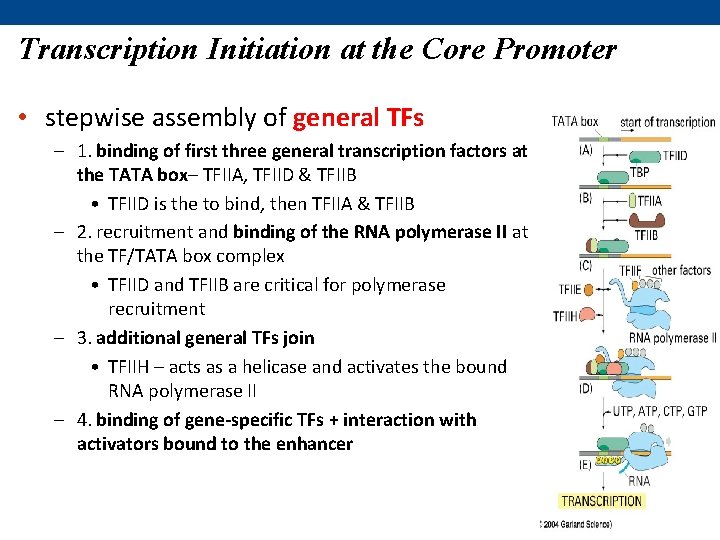
Transcription Initiation at the Core Promoter • stepwise assembly of general TFs – 1. binding of first three general transcription factors at the TATA box– TFIIA, TFIID & TFIIB • TFIID is the to bind, then TFIIA & TFIIB – 2. recruitment and binding of the RNA polymerase II at the TF/TATA box complex • TFIID and TFIIB are critical for polymerase recruitment – 3. additional general TFs join • TFIIH – acts as a helicase and activates the bound RNA polymerase II – 4. binding of gene-specific TFs + interaction with activators bound to the enhancer
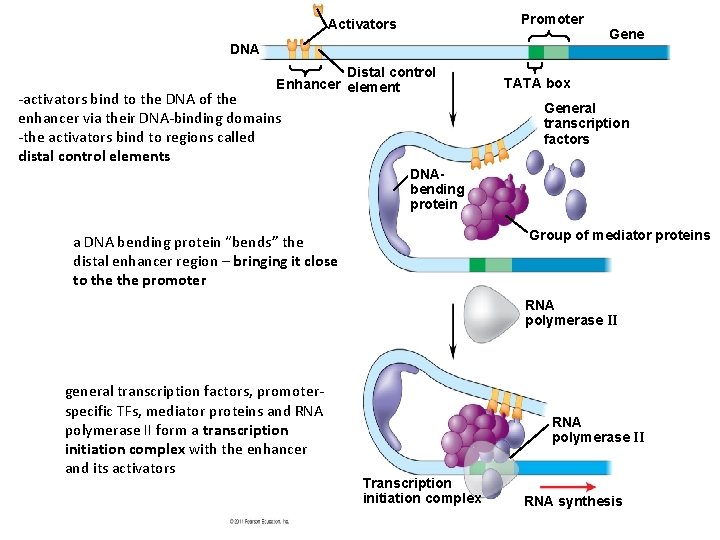
Promoter Activators DNA Distal control Enhancer element -activators bind to the DNA of the enhancer via their DNA-binding domains -the activators bind to regions called distal control elements Gene TATA box General transcription factors DNAbending protein Group of mediator proteins a DNA bending protein “bends” the distal enhancer region – bringing it close to the promoter RNA polymerase II general transcription factors, promoterspecific TFs, mediator proteins and RNA polymerase II form a transcription initiation complex with the enhancer and its activators RNA polymerase II Transcription initiation complex RNA synthesis
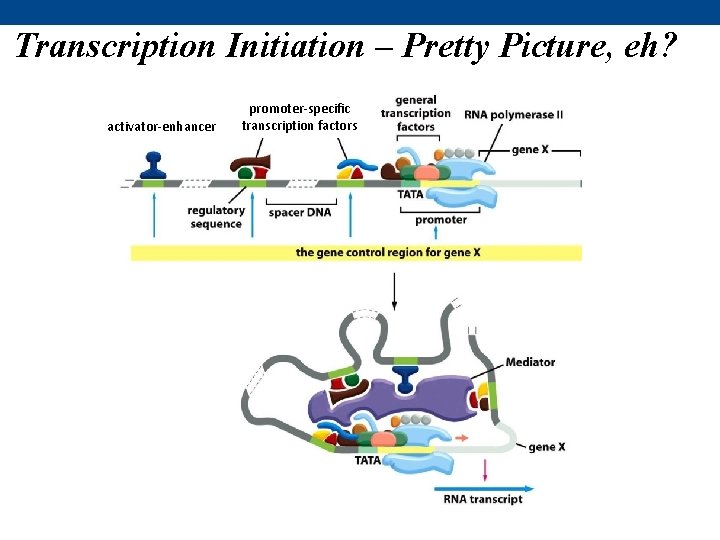
Transcription Initiation – Pretty Picture, eh? activator-enhancer promoter-specific transcription factors

Big Picture Time! – some of these transcription factors can act indirectly by influencing chromatin structure to promote or silence transcription • e. g. some TFs can recruit HAT to the chromatin and increase the degree of acetylation and gene expression

Repressors • some transcription factors can also function as repressors or silencers – – inhibiting expression of a particular gene by a variety of methods some repressors bind activators and prevent their binding to enhancers some bind the distal control elements in the enhancer directly others bind proximal control elements or the promoter
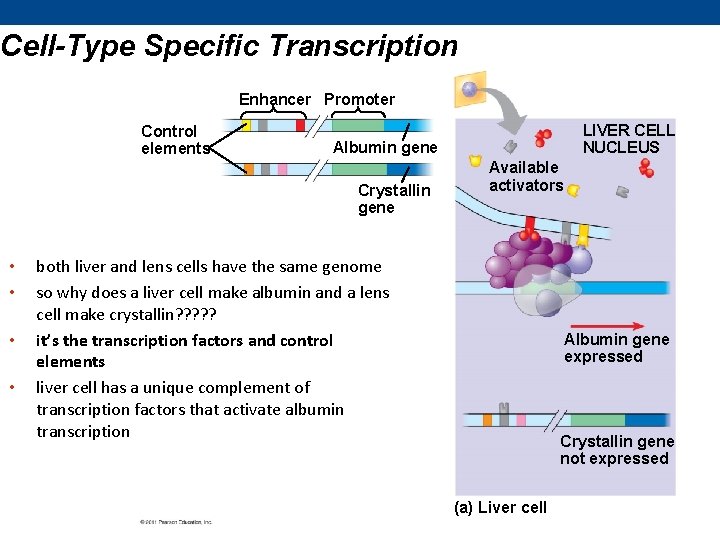
Cell-Type Specific Transcription Enhancer Promoter Control elements Albumin gene Crystallin gene • • LIVER CELL NUCLEUS Available activators both liver and lens cells have the same genome so why does a liver cell make albumin and a lens cell make crystallin? ? ? it’s the transcription factors and control elements liver cell has a unique complement of transcription factors that activate albumin transcription Albumin gene expressed Crystallin gene not expressed (a) Liver cell
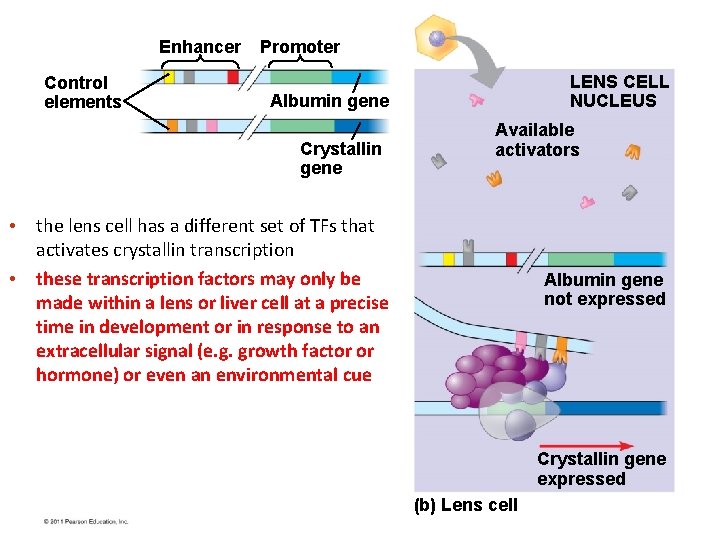
Enhancer Control elements Promoter LENS CELL NUCLEUS Albumin gene Crystallin gene Available activators • the lens cell has a different set of TFs that activates crystallin transcription • these transcription factors may only be made within a lens or liver cell at a precise time in development or in response to an extracellular signal (e. g. growth factor or hormone) or even an environmental cue Albumin gene not expressed Crystallin gene expressed (b) Lens cell

Coordinately Controlled Genes in Eukaryotes • unlike the genes of a prokaryotic operon, each co-expressed eukaryotic gene has a core promoter and several other control elements • these genes can be scattered over different chromosomes, but each has the same combination of control elements as one another • multiple copies of activators recognize these control elements on each gene and promote their simultaneous transcription

GO HAVE LUNCH! Next Lecture POST-TRANSCRIPTIONAL & TRANSLATIONAL REGULATION

Mechanisms of Post-Transcriptional Regulation • transcription alone does not account for gene expression • regulatory mechanisms can operate at various stages after transcription • allow a cell to fine-tune gene expression rapidly in response to environmental changes • post-transcriptional processing: – 1. m. RNA structure – cap and tail; UTRs – 2. m. RNA splicing – 3. m. RNA half life and degradation
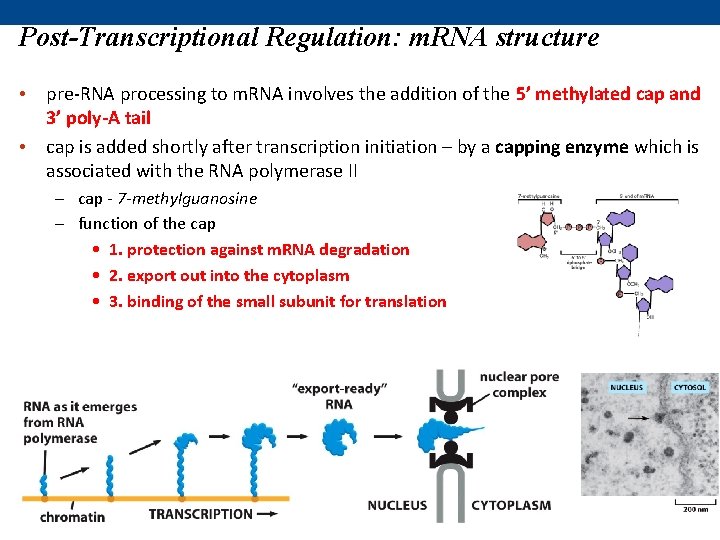
Post-Transcriptional Regulation: m. RNA structure • pre-RNA processing to m. RNA involves the addition of the 5’ methylated cap and 3’ poly-A tail • cap is added shortly after transcription initiation – by a capping enzyme which is associated with the RNA polymerase II – cap - 7 -methylguanosine – function of the cap • 1. protection against m. RNA degradation • 2. export out into the cytoplasm • 3. binding of the small subunit for translation

Post-Transcriptional Regulation: m. RNA structure • cap is followed by the 5’UTR region (untranslated region) – found between the transcription start site and ends one nucleotide before the ATG/start codon of the coding sequence – contains elements for controlling gene expression and m. RNA export – contains sequences that are involved in translation initiation (i. e. docking of the ribosome)
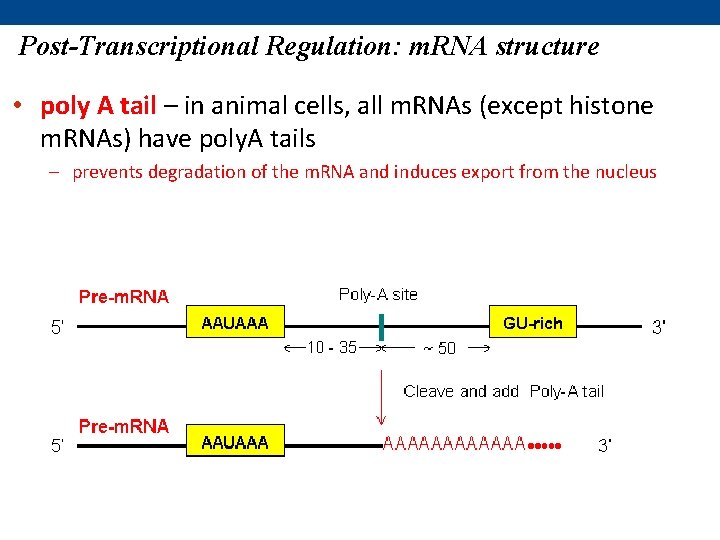
Post-Transcriptional Regulation: m. RNA structure • poly A tail – in animal cells, all m. RNAs (except histone m. RNAs) have poly. A tails – prevents degradation of the m. RNA and induces export from the nucleus
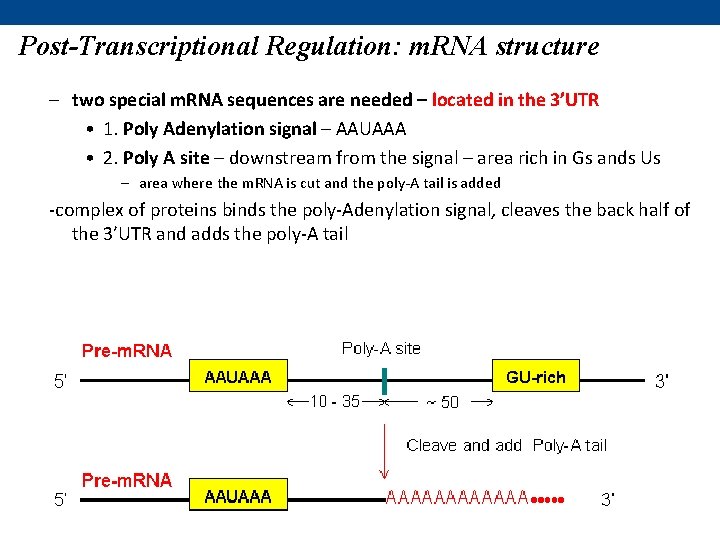
Post-Transcriptional Regulation: m. RNA structure – two special m. RNA sequences are needed – located in the 3’UTR • 1. Poly Adenylation signal – AAUAAA • 2. Poly A site – downstream from the signal – area rich in Gs ands Us – area where the m. RNA is cut and the poly-A tail is added -complex of proteins binds the poly-Adenylation signal, cleaves the back half of the 3’UTR and adds the poly-A tail

Post-Transcriptional Regulation: m. RNA structure • 3’UTR – second of the two UTRs that flank a transcription unit’s coding sequence – controls m. RNA transcription levels – contains numerous regulatory regions for • 1. poly –adenylation – contains the poly. A signal and poly. A site • 2. m. RNA creation – silencer regions to repress transcription of m. RNA • 3. m. RNA stability – contain AU-rich elements (AREs) that increase the stability of the m. RNA • 4. m. RNA export – contains sequences that attract nuclear export proteins • 5. translation efficiency – also affected by AREs

Post-transcriptional control: m. RNA degradation • the life span of m. RNA molecules in the cytoplasm is a key to determining protein synthesis • eukaryotic m. RNA is more long lived than prokaryotic m. RNA • numerous enzymes (RNases) can breakdown m. RNA • the binding of small RNAs called micro. RNAs (mi. RNAs) to the m. RNA can target it for degredation

Noncoding RNAs play multiple roles in controlling gene expression • mi. RNA is a non-coding RNA • only a small fraction of DNA codes for proteins – 30, 000 to 100, 000 genes • the rest of the DNA is either junk or contains sequences important for gene expression BUT do not code for protein = non-coding DNA • most non-coding DNA is transcribed into noncoding RNAs (nc. RNAs) – e. g. mi. RNA – e. g. t. RNA, r. RNA • noncoding RNAs regulate gene expression at two points: m. RNA translation and chromatin configuration
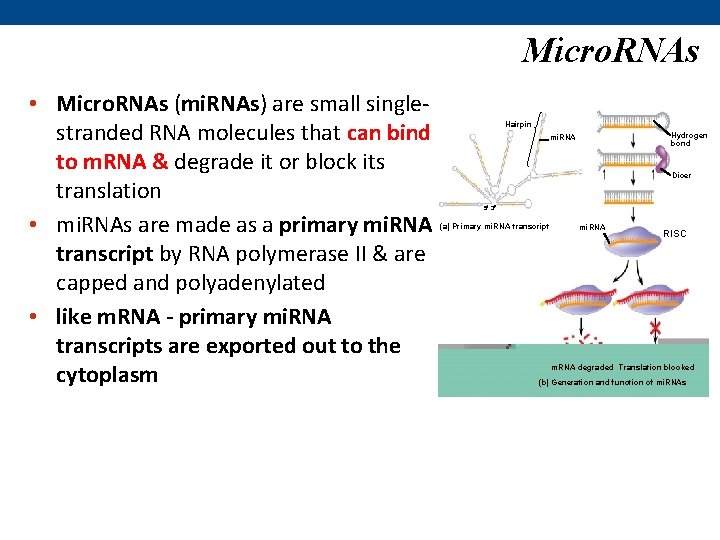
Micro. RNAs • Micro. RNAs (mi. RNAs) are small singlestranded RNA molecules that can bind to m. RNA & degrade it or block its translation • mi. RNAs are made as a primary mi. RNA transcript by RNA polymerase II & are capped and polyadenylated • like m. RNA - primary mi. RNA transcripts are exported out to the cytoplasm Hairpin Hydrogen bond mi. RNA Dicer 5 3 (a) Primary mi. RNA transcript mi. RNA RISC m. RNA degraded Translation blocked (b) Generation and function of mi. RNAs
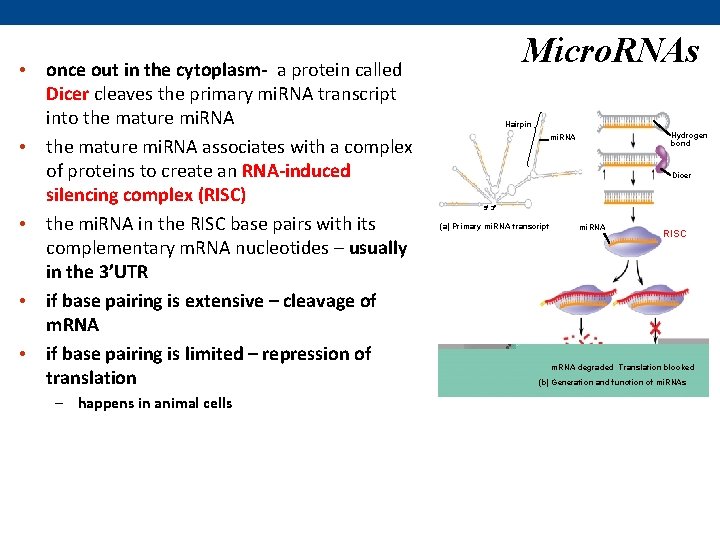
• once out in the cytoplasm- a protein called Dicer cleaves the primary mi. RNA transcript into the mature mi. RNA • the mature mi. RNA associates with a complex of proteins to create an RNA-induced silencing complex (RISC) • the mi. RNA in the RISC base pairs with its complementary m. RNA nucleotides – usually in the 3’UTR • if base pairing is extensive – cleavage of m. RNA • if base pairing is limited – repression of translation – happens in animal cells Micro. RNAs Hairpin Hydrogen bond mi. RNA Dicer 5 3 (a) Primary mi. RNA transcript mi. RNA RISC m. RNA degraded Translation blocked (b) Generation and function of mi. RNAs
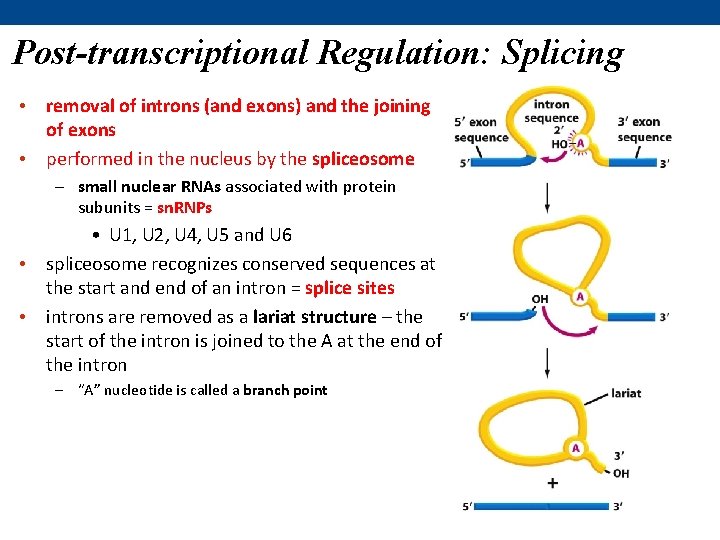
Post-transcriptional Regulation: Splicing • removal of introns (and exons) and the joining of exons • performed in the nucleus by the spliceosome – small nuclear RNAs associated with protein subunits = sn. RNPs • U 1, U 2, U 4, U 5 and U 6 • spliceosome recognizes conserved sequences at the start and end of an intron = splice sites • introns are removed as a lariat structure – the start of the intron is joined to the A at the end of the intron – “A” nucleotide is called a branch point
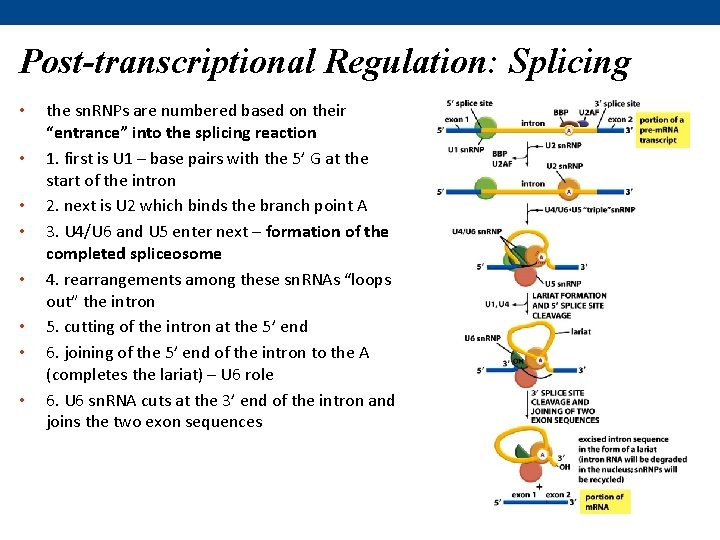
Post-transcriptional Regulation: Splicing • • the sn. RNPs are numbered based on their “entrance” into the splicing reaction 1. first is U 1 – base pairs with the 5’ G at the start of the intron 2. next is U 2 which binds the branch point A 3. U 4/U 6 and U 5 enter next – formation of the completed spliceosome 4. rearrangements among these sn. RNAs “loops out” the intron 5. cutting of the intron at the 5’ end 6. joining of the 5’ end of the intron to the A (completes the lariat) – U 6 role 6. U 6 sn. RNA cuts at the 3’ end of the intron and joins the two exon sequences
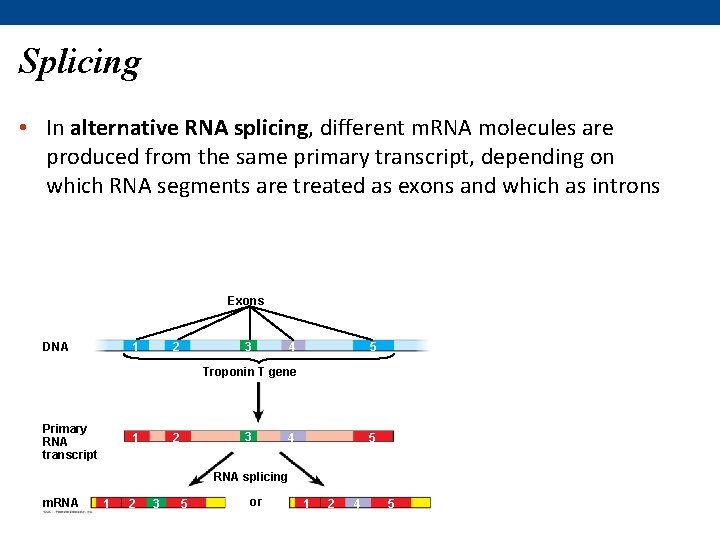
Splicing • In alternative RNA splicing, different m. RNA molecules are produced from the same primary transcript, depending on which RNA segments are treated as exons and which as introns Exons DNA 1 4 3 2 5 Troponin T gene Primary RNA transcript 3 2 1 5 4 RNA splicing m. RNA 1 2 3 5 or 1 2 4 5

Initiation of Translation • initiation of translation of select m. RNAs can be blocked by regulatory proteins that bind to sequences or structures of the m. RNA • alternatively, translation of all m. RNAs in a cell may be regulated simultaneously – for example, translation initiation factors are simultaneously activated in an egg following fertilization

Protein Processing • after translation - various types of protein processing, including folding, cleavage and the addition of chemical groups take place • known as post-translational processing • numerous kinds of chemical additions – enzymes adding fatty acids, hydrophobic groups, additional peptides like ubiquitin – enzymes cleaving the proteins – addition of chemical groups like phosphate groups, acetyl groups, carbohydrates

Protein Folding: Chaperones • protein function is completely dependent upon 3 D structure • the information for folding is contained within the amino acid sequence of the polypeptide chain – the hydrophobic residues are “buried” within the center of the folding protein – spontaneous process – large numbers of interactions between the R groups of the AAs form • ionic • van der waals – between hydrophobic groups • disulfide bridges • folding can begin the minute the PP chain emerges from the ribosome • but most protein don’t • these proteins are met at the ribosome by a class of proteins called molecular chaperones
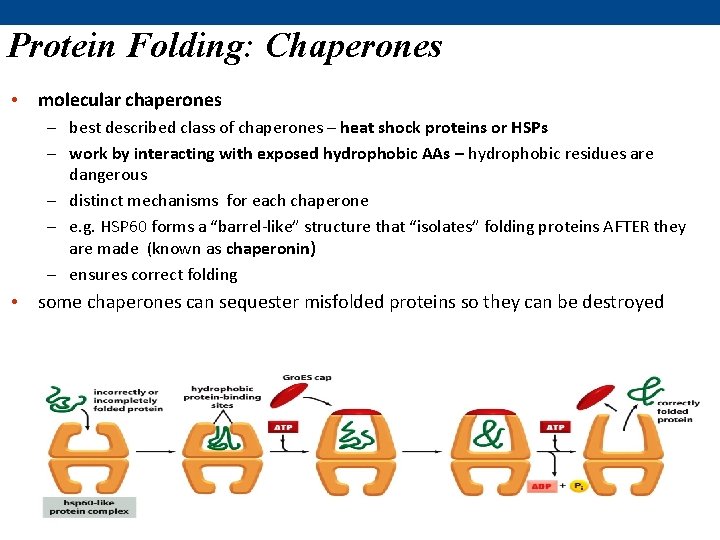
Protein Folding: Chaperones • molecular chaperones – best described class of chaperones – heat shock proteins or HSPs – work by interacting with exposed hydrophobic AAs – hydrophobic residues are dangerous – distinct mechanisms for each chaperone – e. g. HSP 60 forms a “barrel-like” structure that “isolates” folding proteins AFTER they are made (known as chaperonin) – ensures correct folding • some chaperones can sequester misfolded proteins so they can be destroyed
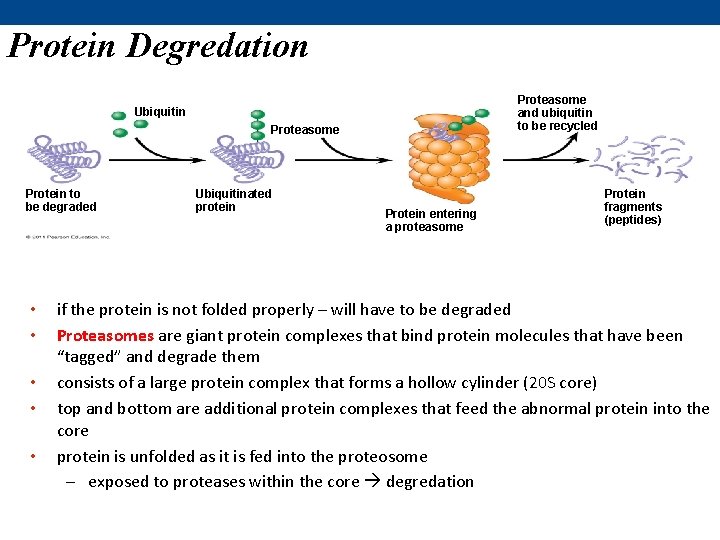
Protein Degredation Proteasome and ubiquitin to be recycled Ubiquitin Proteasome Protein to be degraded • • • Ubiquitinated protein Protein entering a proteasome Protein fragments (peptides) if the protein is not folded properly – will have to be degraded Proteasomes are giant protein complexes that bind protein molecules that have been “tagged” and degrade them consists of a large protein complex that forms a hollow cylinder (20 S core) top and bottom are additional protein complexes that feed the abnormal protein into the core protein is unfolded as it is fed into the proteosome – exposed to proteases within the core degredation
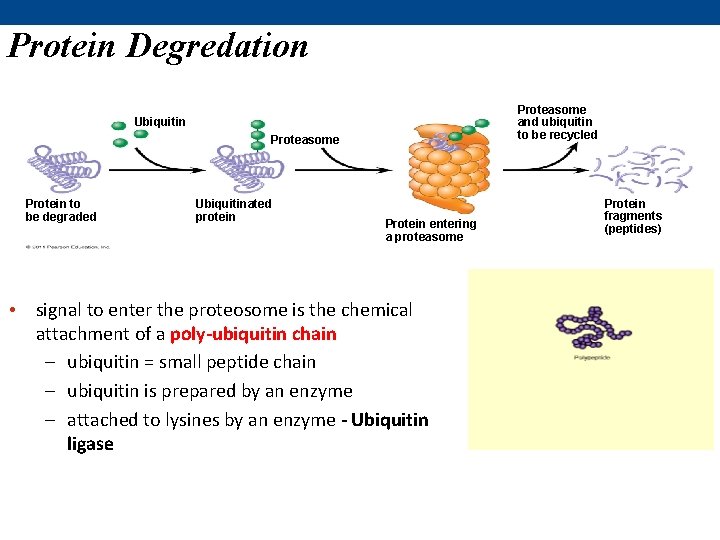
Protein Degredation Proteasome and ubiquitin to be recycled Ubiquitin Proteasome Protein to be degraded Ubiquitinated protein Protein entering a proteasome • signal to enter the proteosome is the chemical attachment of a poly-ubiquitin chain – ubiquitin = small peptide chain – ubiquitin is prepared by an enzyme – attached to lysines by an enzyme - Ubiquitin ligase Protein fragments (peptides)
- Slides: 80