Lecture Power Point to accompany Molecular Biology Fourth

Lecture Power. Point to accompany Molecular Biology Fourth Edition Robert F. Weaver Chapter 14 Messenger RNA Processing I: Splicing Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display.

14. 1 Genes in Pieces • Consider the sequence of the human b-globin gene as a sentence: This is bhgty the human b-globin qwtzptlrbn gene. • Two italicized regions make no sense – Contain sequences unrelated to the globin coding sequences surrounding them – Called intervening sequences, IVSs – Usually called introns • Parts of the gene making sense – Coding regions – Exons • Some lower eukaryotic genes have no introns 2

Evidence for Split Genes • Most higher eukaryotic genes coding for m. RNA and t. RNA are interrupted by unrelated regions called introns • Other parts of the gene, surrounding the introns, are called exons • Exons contain the sequences that finally appear in the mature RNA product – Genes for m. RNAs have been found with anywhere from 0 to 362 exons – t. RNA genes have either 0 or 1 exon 3

RNA Splicing • Introns are present in genes but not in mature RNA • How does the information not find its way into mature RNA products of the genes? – Introns are never transcribed • Polymerase somehow jumps from one exon to another – Introns are transcribed • Primary transcript result, an overlarge gene product is cut down by removing introns • This is correct process • Process of cutting introns out of immature RNAs and stitching together the exons to form final product is RNA splicing 4

Splicing Outline • Introns are transcribed along with exons in the primary transcript • Introns are removed as the exons are spliced together 5

Stages of RNA Splicing • Messenger RNA synthesis in eukaryotes occurs in stages • First stage: – Synthesis of primary transcript product – This is an m. RNA precursor containing introns copied from the gene if present – Precursor is part of a pool of large nuclear RNAs – hn. RNAs • Second stage: – m. RNA maturation – Removal of introns in a process called splicing 6

Splicing Signals • Splicing signals in nuclear m. RNA precursors are remarkably uniform – First 2 bases of introns are GU – Last 2 are AG • 5’- and 3’-splice sites have consensus sequences extending beyond GU and AG motifs • Whole consensus sequences are important to proper splicing • Abnormal splicing can occur when the consensus sequences are mutated 7

14. 2 Mechanism of Splicing of Nuclear m. RNA Precursors • Intermediate in nuclear m. RNA precursor splicing is branched – looks like a lariat • 2 -step model – 2’-OH group of adenosine nucleotide in middle of intron attacks phosphodiester bond between 1 st exon and G beginning of intron • Forms loop of the lariat • Separates first exon from intron – 3’-OH left at end of 1 st exon attacks phosphodiester bond linking intron to 2 nd exon • Forms the exon-exon phosphodiester bond • Releases intron in lariat form at same time 8
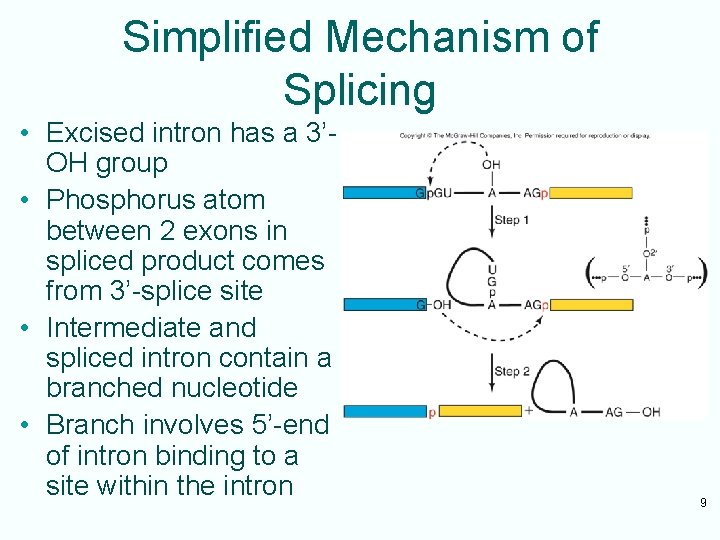
Simplified Mechanism of Splicing • Excised intron has a 3’OH group • Phosphorus atom between 2 exons in spliced product comes from 3’-splice site • Intermediate and spliced intron contain a branched nucleotide • Branch involves 5’-end of intron binding to a site within the intron 9
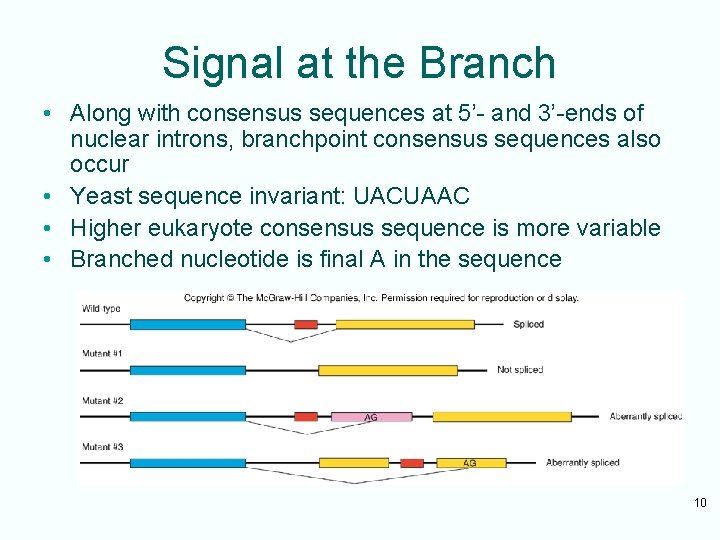
Signal at the Branch • Along with consensus sequences at 5’- and 3’-ends of nuclear introns, branchpoint consensus sequences also occur • Yeast sequence invariant: UACUAAC • Higher eukaryote consensus sequence is more variable • Branched nucleotide is final A in the sequence 10

Spliceosomes • Splicing takes place on a particle called a spliceosome • Yeast spliceosomes and mammalian spliceosomes have sedimentation coefficients of 40 S and 60 S • Spliceosomes contain the pre-m. RNA – Along with sn. RNPs and protein splicing factors – These recognize key splicing signals and orchestrate the splicing process 11

sn. RNPs • Small nuclear RNAs coupled to proteins are abbreviated as sn. RNPs, small nuclear ribonuclear proteins • The sn. RNAs (small nuclear RNAs) can be resolved on a gel: – U 1, U 2, U 4, U 5, U 6 – All 5 sn. RNAs join the spliceosome to play crucial roles in splicing 12
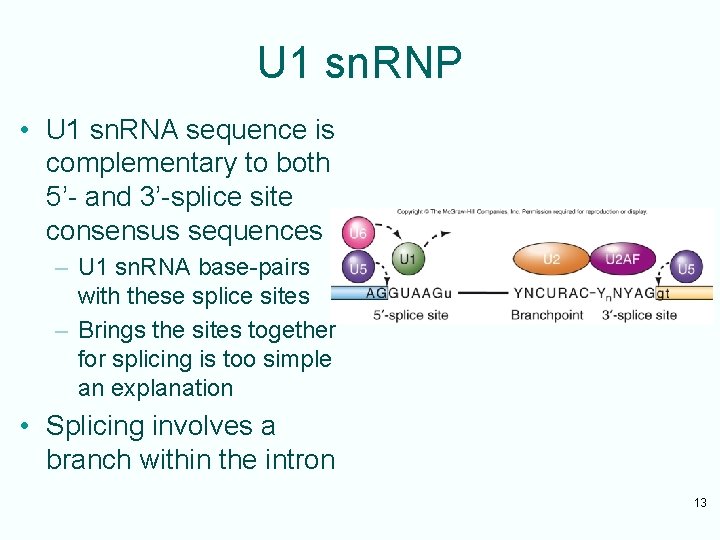
U 1 sn. RNP • U 1 sn. RNA sequence is complementary to both 5’- and 3’-splice site consensus sequences – U 1 sn. RNA base-pairs with these splice sites – Brings the sites together for splicing is too simple an explanation • Splicing involves a branch within the intron 13

Wild-Type and Mutant U 1 sn. RNA • Genetic experiments completed • Base pairing between U 1 sn. RNA and 5’-splice site of m. RNA precursor – Is necessary – Not sufficient for binding 14

U 6 sn. RNP • U 6 sn. RNP associates with the 5’-end of the intron by base pairing through the U 6 RNA • Occurs first prior to formation of lariat intermediate • Character may change after first step in splicing • Association between U 6 and splicing substrate is essential for the splicing process • U 6 also associates with U 2 during splicing 15

U 2 sn. RNP • U 2 sn. RNA base-pairs with the conserved sequence at the splicing branchpoint • This base pairing is essential for splicing • U 2 also forms base pairs with U 6 – This region is called helix I – Helps orient sn. RNPs for splicing • 5’-end of U 2 interacts with 3’-end of U 6 – This interaction forms a region called helix II – This region is important in splicing in mammalian cells, not in yeast cells 16
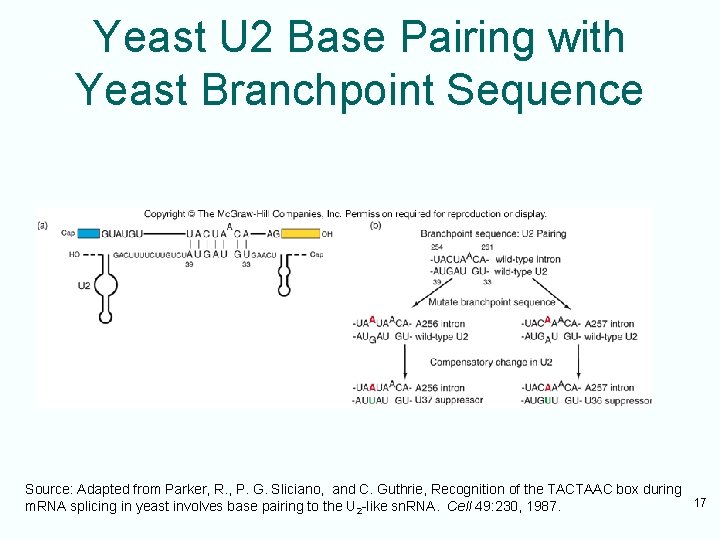
Yeast U 2 Base Pairing with Yeast Branchpoint Sequence Source: Adapted from Parker, R. , P. G. Sliciano, and C. Guthrie, Recognition of the TACTAAC box during 17 m. RNA splicing in yeast involves base pairing to the U 2 -like sn. RNA. Cell 49: 230, 1987.

U 5 sn. RNP • U 5 sn. RNA associates with the last nucleotide in one exon and the first nucleotide of the next exon • This should result in the two exons lining up for splicing 18

U 4 sn. RNP • U 4 base-pairs with U 6 • Its role seems to be to bind U 6 • When U 6 is needed in a splicing reaction U 4 is removed • U 6 gene is split by an m. RNA-type intron in at least two yeast species 19

sn. RNP Involvement in m. RNA Splicing • Spliceosomal complex contains: – – Substrate U 2 U 5 U 6 • The complex ready for the 2 nd step in splicing can be drawn as a group II intron at same stage of splicing • Spliceosomal sn. RNPs substitute for elements at center of catalytic activity of group II introns at same stage of splicing 20

Spliceosome Catalytic Activity • Catalytic center of spliceosome appears to include Mg 2+ and a base-paired complex of 3 RNAs: – U 2 sn. RNA – U 6 sn. RNA – Branchpoint region of the intron • Protein-free fragments of these RNAs can catalyze a reaction related to the first step in splicing 21

Spliceosome Assembly and Function • Spliceosome is composed of many components – proteins and RNA • These components assemble stepwise • The spliceosome cycle: – Assembly – Function – Disassembly • By controlling assembly of the spliceosome, a cell can regulate quality and quantity of splicing and so regulate gene expression 22

Spliceosome Cycle • Assembly begins with binding of U 1 to splicing substrate forming a commitment complex, a unit committed to to splicing out the intron • U 2 joins the complex next, followed by the others • U 2 binding requires ATP • U 6 dissociates from U 4 and displaces U 1 at the 5’-splice site – This step is ATP-dependent – Activates the spliceosome – Allows U 1 and U 4 to be released 23

sn. RNP Structure • All sn. RNP’s have the same set of 7 Sm proteins – Common targets of antibodies in patients with systemic autoimmune diseases – Sm protein binds to a common Sm site on the sn. RNAs: AAUUUGUGG • U 1 sn. RNP has 3 specific proteins – 70 K has an Mr of 52 k. D – A has an Mr of 31 k. D – C has an Mr of 17. 5 k. D • Sm proteins form a doughnut-shaped structure with a hole through the middle, like a flattened funnel 24

Sm Site and RNA • Five sn. RNPs participate in splicing • All contain a common set of 7 Sm proteins and several other proteins that are specific to sn. RNP • Structure of U 1 sn. RNP reveals that the Sm proteins form a doughnut-shaped structure to which the other proteins are attached 25

A Minor Spliceosome • A minor class of introns with variant but highly conserved 5’-splice sites and branchpoints can be spliced with the help of a variant class of sn. RNAs • Cells can contain minor sn. RNAs: – U 11 performs like U 1 – U 12 acts like U 2 – U 4 atac and U 6 atac perform like U 4 and U 6 respectively 26

Commitment, Splice Site Selection and Alternative Splicing • sn. RNPs do not have enough specificity and affinity to bind exclusively and tightly at exon-intron boundaries • Additional splicing factors are needed to help sn. RNPs bind • Some splicing factors are needed to bridge across introns and exons and so define these RNA elements 27

3’-Splice Site Selection • Splicing factor Slu 7 is required for correct 3’ -splicing site selection • Without Slu 7, splicing to correct 3’-splice site AG is suppressed and splicing to aberrant AG’s within 30 nt of the branchpoint is activated • U 2 AF is also required for 3’-splice site recognition • 65 -k. D U 2 AF subunit binds to polypyrimidine tract upstream of 3’-splice site and 35 -k. D subunit binds to the 3’-splice site AG 28

Commitment • Commitment to splice at a given site is determined by an RNA-binding protein • This protein binds to splicing substrate and recruits other spliceosomal components • The first component to follow is U 1 • SR proteins SC 35 and SF 2/ASF commit splicing on human b-globin pre-m. RNA and HIV tat pre-m. RNA • Part of the commitment involves attraction of U 1 in some cases 29

Bridging Proteins and Commitment • Yeast commitment complex has a branchpoint bridging protein (BBP) binds to: – U 1 sn. RNP protein at the 5’-end of the intron – Mud 2 p near the 3’-end of the intron – RNA near the 3’-end of the intron • Bridges the intron and could play a role defining intron prior to splicing • Mammalian BBP is SF 1, may serve same bridging function 30
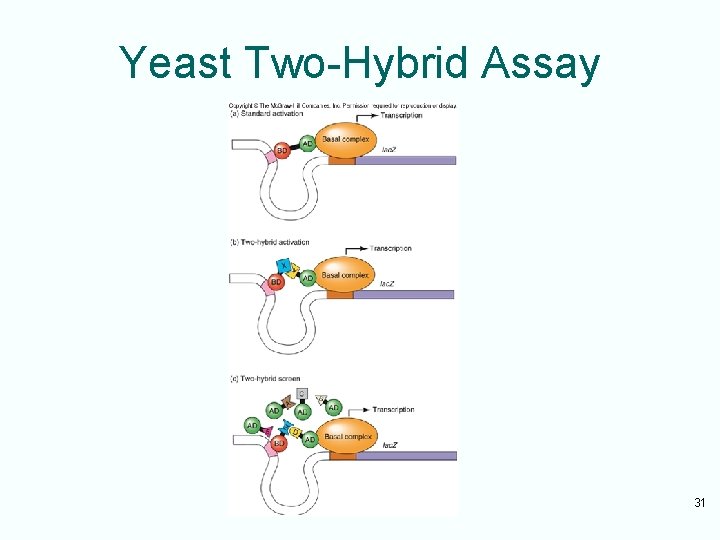
Yeast Two-Hybrid Assay 31
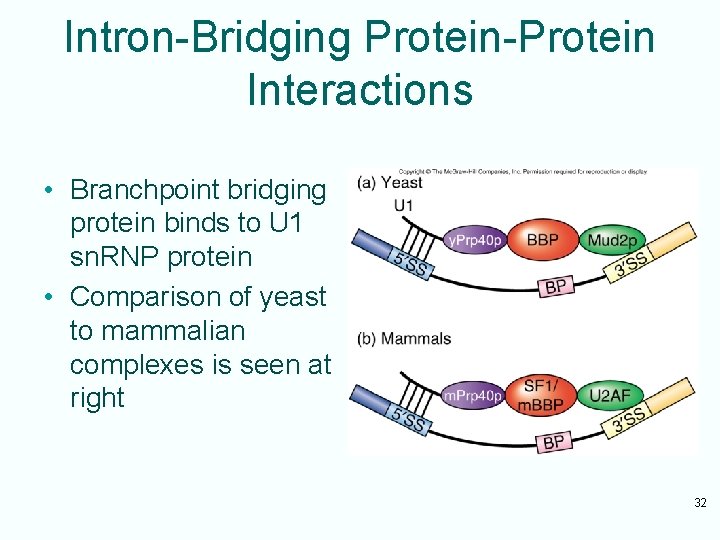
Intron-Bridging Protein-Protein Interactions • Branchpoint bridging protein binds to U 1 sn. RNP protein • Comparison of yeast to mammalian complexes is seen at right 32

Role of the RNA Polymerase II CTD • C-terminal domain of the Rpb 1 subunit of RNA polymerase II stimulates splicing of substrates that use exon definition • This does not apply to those that use intron definition to prepare for splicing • CTD binds to splicing factors and could assemble the factors at the end of exons to set them off for splicing 33

Alternative Splicing • Transcripts of many eukaryotic genes are subject to alternative splicing – This splicing can have profound effects on the protein products of a gene – Can make a difference between: • Secreted or membrane-bound protein • Activity and inactivity • Products of 3 genes in sex determination pathway of the fruit fly are subject to alternative splicing 34

Sex-Specific Splicing • Female-specific splicing of transcript gives: – An active product that causes female-specific splicing of dsx pre-m. RNA – This produces a female fruit fly • Male-specific splicing of transcript gives: – An inactive product that allows male-specific splicing of dsx pre-m. RNA – This produces a male fruit fly 35

Tra and Tra-2 • Tra and its partner Tra-2 act in conjunction with one or more other SR proteins to commit splicing at the female-specific splice site on the dsx pre-m. RNA • Commitment is probably the basis of most, if not all, alternative splicing schemes 36
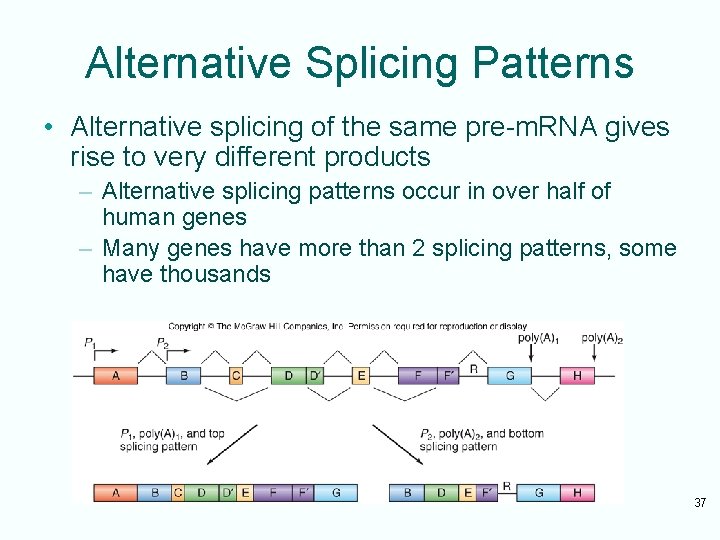
Alternative Splicing Patterns • Alternative splicing of the same pre-m. RNA gives rise to very different products – Alternative splicing patterns occur in over half of human genes – Many genes have more than 2 splicing patterns, some have thousands 37

Types of Alternative Splicing • Begin transcripts at alternative promoters • Some exons can simply be ignored resulting in deletion of the exon • Alternative 5’-splice sites can lead to inclusion or deletion of part of an exon • Alternative 3’-splice sites can lead to inclusion or deletion of part of an exon • A retained intron can be retained in the m. RNA if it is not recognized as an intron • Polyadenylation causes cleavage of pre-m. RNA and loss of downstream exons 38
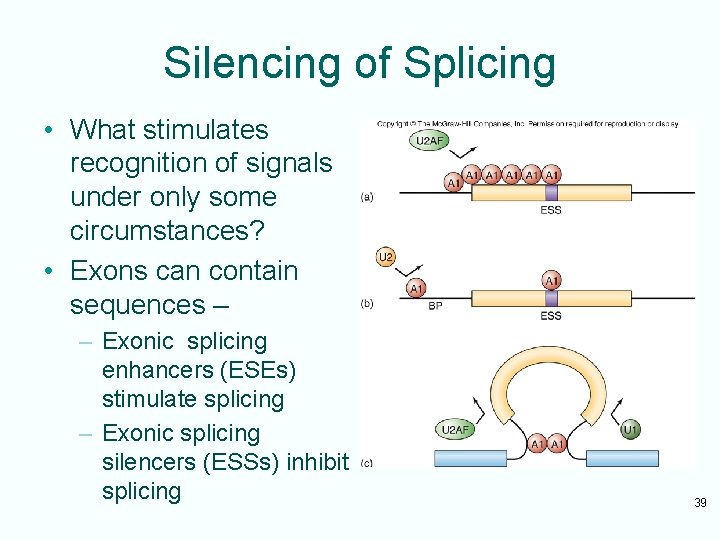
Silencing of Splicing • What stimulates recognition of signals under only some circumstances? • Exons can contain sequences – – Exonic splicing enhancers (ESEs) stimulate splicing – Exonic splicing silencers (ESSs) inhibit splicing 39
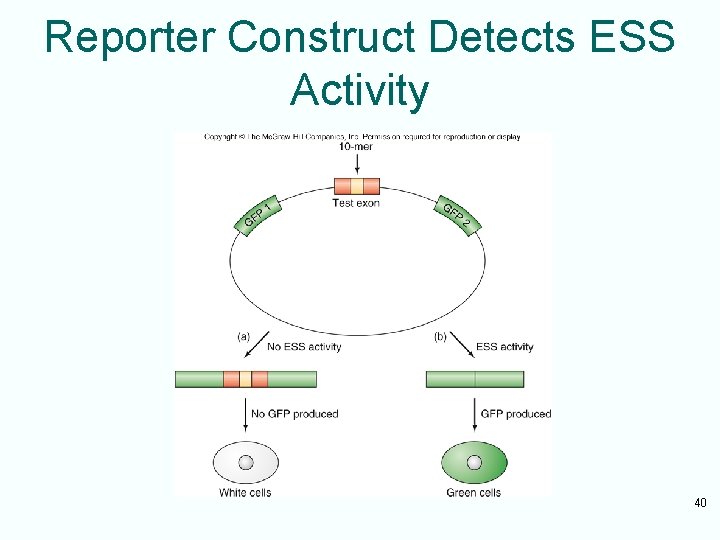
Reporter Construct Detects ESS Activity 40

14. 3 Self-Splicing RNAs • Some RNAs could splice themselves without aid from a spliceosome or any other protein • Tetrahymena 26 S r. RNA gene has an intron, splices itself in vitro – Group I introns are a group of self-splicing RNAs – Another group, Group II introns also have some self-splicing members 41

Group I Introns • Group I introns can be removed in vitro with no help from protein • Reaction begins with attack by a guanine nucleotide on the 5’-splice site – Adds G to the 5’-end of the intron – Releases the first exon • Second step, first exon attacks the 3’-splice site – Ligates 2 exons together – Releases the linear intron • Intron cyclizes twice, losing nucleotides each time, then linearizes a last time 42
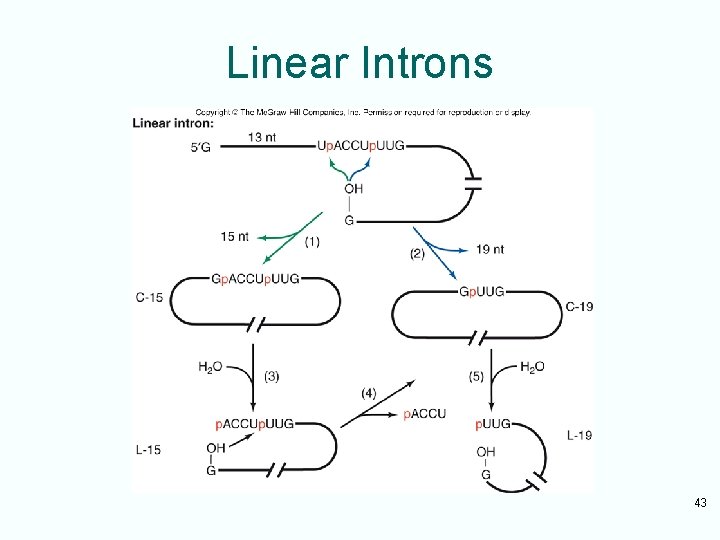
Linear Introns 43

Group II Introns • RNAs containing group II introns selfsplice by a pathway using an A-branched lariat intermediate, like spliceosome lariats • Secondary structures of the splicing complexes involving spliceosomal systems and group II introns are very similar 44
- Slides: 44