Lecture Power Point to accompany Molecular Biology Fourth

Lecture Power. Point to accompany Molecular Biology Fourth Edition Robert F. Weaver Chapter 10 Eukaryotic RNA Polymerases and Their Promoters Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display.

10. 1 Multiple Forms of Eukaryotic RNA Polymerase • There at least two RNA polymerases operating in eukaryotic nuclei – One transcribes major ribosomal RNA genes – One or more to transcribe rest of nuclear genes • Ribosomal genes are different from other nuclear genes – Different base composition from other nuclear genes – Unusually repetitive – Found in different compartment, the nucleolus 2

Separation of the Three Nuclear Polymerases • Eukaryotic nuclei contain three RNA polymerases – These can be separated by ion-exchange chromatography • RNA polymerase I found in nucleolus – Location suggests in transcribes r. RNA genes • RNA polymerases II and III are found in the nucleoplasm 3

Roles of the Three RNA Polymerases • Polymerase I makes large r. RNA precursor • Polymerase II makes – Heterogeneous nuclear RNA (hn. RNA) – Small nuclear RNA • Polymerase III makes precursors to t. RNAs, 5 S r. RNA and other small RNA 4

RNA Polymerase Subunit Structures 5

Polymerase II Structure • For enzymes like eukaryotic RNA polymerases, can be difficult to tell: – Which polypeptides copurify with polymerase activity – Which are actually subunits of the enzyme • Technique to help determine whether a polypeptide copurifies or is a subunit is called epitope tagging 6
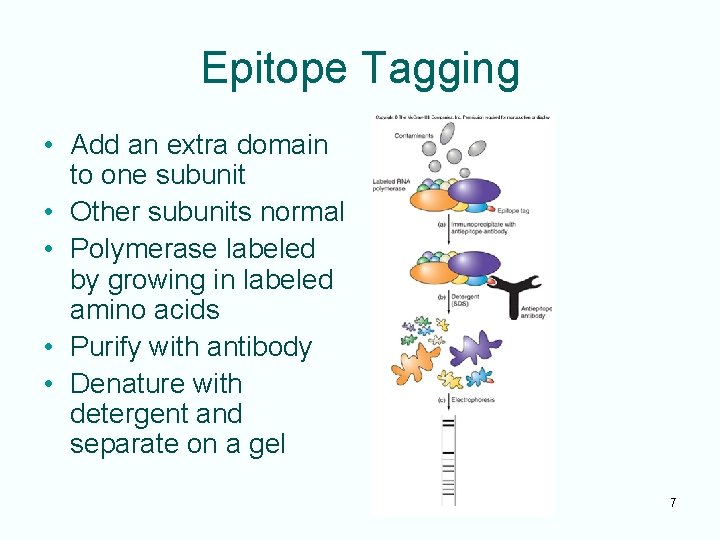
Epitope Tagging • Add an extra domain to one subunit • Other subunits normal • Polymerase labeled by growing in labeled amino acids • Purify with antibody • Denature with detergent and separate on a gel 7

Polymerase II Original 10 subunits are placed in 3 groups: • Core – related in structure and function to bacterial core subunits • Common – found in all 3 nuclear RNA polymerases • Nonessential subunits – conditionally dispensable for enzymatic activity 8

Core Subunits • Three polypeptides, Rpb 1, Rpb 2, Rpb 3 are absolutely required for enzyme activity • These are homologous to b’-, b-, and a-subunits • Both Rpb 1 and b’-subunit binds DNA • Rpb 2 and b-subunit are at or near the nucleotide -joining active site • Rpb 3 does not resemble a-subunit – There is one 20 -amino acid subunit of great similarity – 2 subunits are about same size, same stoichiometry – 2 monomers per holoenzyme 9

Common Subunits • There are five common subunits – Rpb 5 – Rpb 6 – Rpb 8 – Rpb 10 – Rpb 12 • Little known about function • They are all found in all 3 polymerases • Suggests play roles fundamental in transcription 10

Subunits Nonessential for Elongation • Rpb 4 and Rpb 7 – – – Dissociate fairly easily from polymerase Found in substoichiometric quantities Might shuttle from one polymerase II to another Rpb 4 may help anchor Rpb 7 to the enzyme Mutants without Rpb 4 and Rpb 7 transcribes well, but cannot initiate at a real promoter • Rpb 7 is an essential subunit, so must not be completely absent in the mutant 11

Heterogeneity of the Rpb 1 Subunit • RPB 1 gene product is subunit II • Subunit IIa is the primary product in yeast – Can be converted to IIb by proteolytic removal of the carboxyl-terminal domain (CTD) which is 7 -peptide repeated over and over – Converts to IIo by phosphorylating 2 ser in the repeating heptad of the CTD – Enzyme with IIa binds to the promoter – Enzyme with IIo is involved in transcript elongation 12

The Three-Dimensional Structure of RNA Polymerase II • Structure of yeast polymerase II (specifically pol II 4/7) at atomic resolution reveals a deep cleft that accepts a linear DNA template from one end to another • Catalytic center lies at the bottom of the cleft and contains a Mg 2+ ion • A second Mg 2+ ion present in low concentrations • Geometry allows enough space for: – TFIID to bind at the TATA box of the promoter – TFIIB to link the polymerase to TFIID – Places polymerase correctly to initiate transcription 13

3 -D Structure - RNA Polymerase II in an Elongation Complex • Structure of polymerase II bound to DNA template and RNA product in an elongation complex has been determined • When nucleic acids are present, the clamp region of the polymerase has shifted closed over the DNA and RNA – Closed clamp ensures that transcription is processive – able to transcribe a whole gene without falling off and terminating prematurely 14
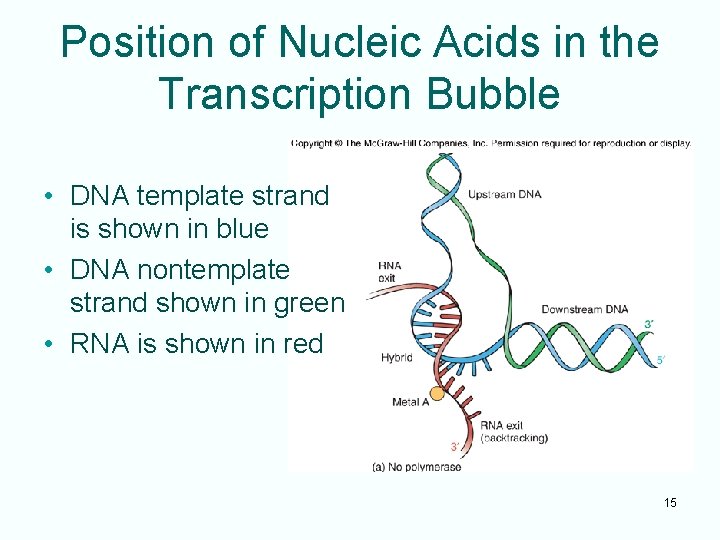
Position of Nucleic Acids in the Transcription Bubble • DNA template strand is shown in blue • DNA nontemplate strand shown in green • RNA is shown in red 15
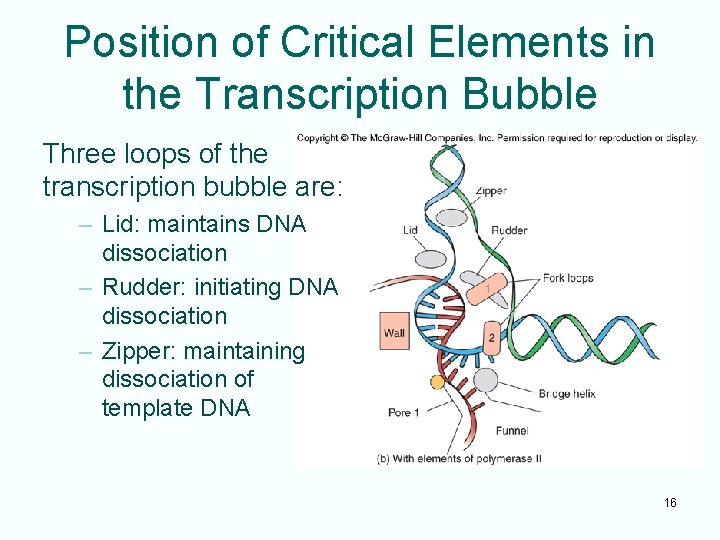
Position of Critical Elements in the Transcription Bubble Three loops of the transcription bubble are: – Lid: maintains DNA dissociation – Rudder: initiating DNA dissociation – Zipper: maintaining dissociation of template DNA 16
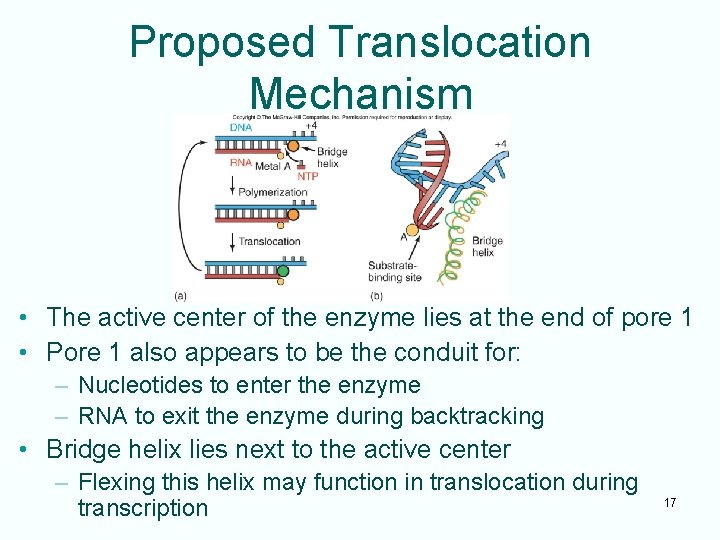
Proposed Translocation Mechanism • The active center of the enzyme lies at the end of pore 1 • Pore 1 also appears to be the conduit for: – Nucleotides to enter the enzyme – RNA to exit the enzyme during backtracking • Bridge helix lies next to the active center – Flexing this helix may function in translocation during transcription 17

3 -D Structure - RNA Polymerase II in the Posttranslocation State • X-ray crystallography has shown the lid of Rpb 1 interacts with the DNA-RNA hybrid to force the hybrid open after base pair -8 • The lid then interacts with bases of the nascent RNA to keep the hybrid melted beyond base pair -8 • The rudder of Rpb 1 collaborates with lid to keep the hybrid melted by interacting with bases -9 and -10 • Fork loop 1 of Rpb 2 interacts with bases -5, -6, and -7 of the RNA to keep the RNA-DNA hybrid together 18

Structural Basis of Nucleotide Selection • Moving through the entry pore toward the active site of RNA polymerase II, incoming nucleotide first encounters the E (entry) site – E site is inverted relative to its position in the A site (active) where phosphodiester bonds form – E and A sites partially overlap – Rotation of nucleotide between the E and A sites may play a role in base and sugar specificity • Two metal ions (Mg 2+ or Mn 2+) are present at the active site – One is permanently bound to the enzyme – The other enters the active site complexed to the incoming nucleotide 19

The Role of Rpb 4 and Rpb 7 • Structure of the 12 -subunit RNA polymerase II reveals that, with Rpb 4/7 in place, clamp is forced shut • Initiation occurs, with its clamp shut, it appears that the promoter DNA must melt to permit template DNA strand to enter the active site • The Rpb 4/7 extends the dock region of the polymerase, which makes binding of transcription factors easier 20

10. 2 Promoters • Three eukaryotic RNA polymerases have: – Different structures – Transcribe different classes of genes • Expect that the 3 polymerases would recognize different promoters 21
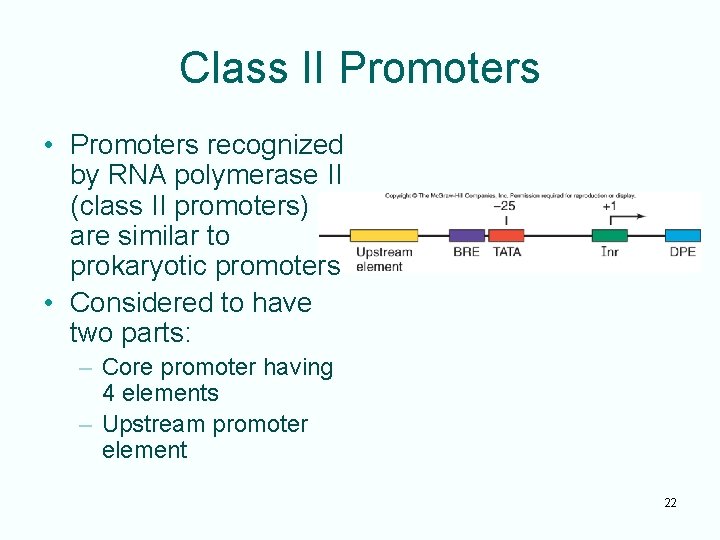
Class II Promoters • Promoters recognized by RNA polymerase II (class II promoters) are similar to prokaryotic promoters • Considered to have two parts: – Core promoter having 4 elements – Upstream promoter element 22

Core Promoter Elements – TATA Box • TATA box – Found on the nontemplate strand – Very similar to the prokaryotic -10 box – There are frequently TATA-less promoters • Housekeeping genes that are constitutively active in nearly all cells as they control common biochemical pathways • Developmentally regulated genes 23

Linker Scanning • Systematically substitute a 10 -bp linker for 10 -bp sequences throughout the promoter • Found that mutations within the TATA box destroyed promoter activity 24

Core Promoter Elements • In addition to TATA box, core promoters are: – TFIIB recognition element (BRE) – Initiator (Inr) – Downstream promoter element (DPE) • At least one of the four core elements is missing in most promoters • TATA-less promoters tend to have DPEs • Promoters for highly specialized genes tend to have TATA boxes • Promoters for housekeeping genes tend to lack them 25

Upstream Elements • Upstream promoter elements are usually found upstream of class II core promoters • Differ from core promoters in binding to relatively gene-specific transcription factors – GC boxes bind transcription factor Sp 1 – CCAAT boxes bind CTF (CCAAT-binding transcription factor) • Upstream promoter elements can be orientationindependent, yet are relatively positiondependent 26

Class I Promoters • Class I promoters are not well conserved in sequence across species • General architecture of the promoter is well conserved – two elements: – Core element surrounding transcription start site – Upstream promoter element (UPE) 100 bp farther upstream – Spacing between these elements is important 27

Class III Promoters • RNA polymerase III transcribes a set of short genes • These have promoters that lie wholly within the genes • There are 3 types of these promoters 28

Promoters of Some Polymerase III Genes • Type I (5 S r. RNA) has 3 regions: – Box A – Short intermediate element – Box C • Type II (t. RNA) has 2 regions: – Box A – Box B • Type III (nonclassical) resemble those of type II 29

10. 3 Enhancers and Silencers • These are position- and orientationindependent DNA elements that stimulate or depress, respectively, transcription of associated genes • Are often tissue-specific in that they rely on tissue-specific DNA-binding proteins for their activities • Some DNA elements can act either as enhancer or silencer depending on what is bound to it 30
- Slides: 30