Lecture Power Point to accompany Molecular Biology Fourth

Lecture Power. Point to accompany Molecular Biology Fourth Edition Robert F. Weaver Chapter 16 Other RNA Processing Events Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display.

16. 1 Ribosomal RNA Processing • r. RNA genes of both eukaryotes and bacteria are transcribed as larger precursors must be processed to yield r. RNAs of mature size • Several different r. RNA molecules are embedded in a long, precursor and each must be cut out 2

Eukaryotic r. RNA Processing • Ribosomal RNAs are made in eukaryotic nucleoli as precursors that must be processed to release mature r. RNAs • Order of RNAs in the precursor is – 18 S – 5. 8 S – 28 S in all eukaryotes – Exact sizes of the mature r. RNAs vary from one species to another 3
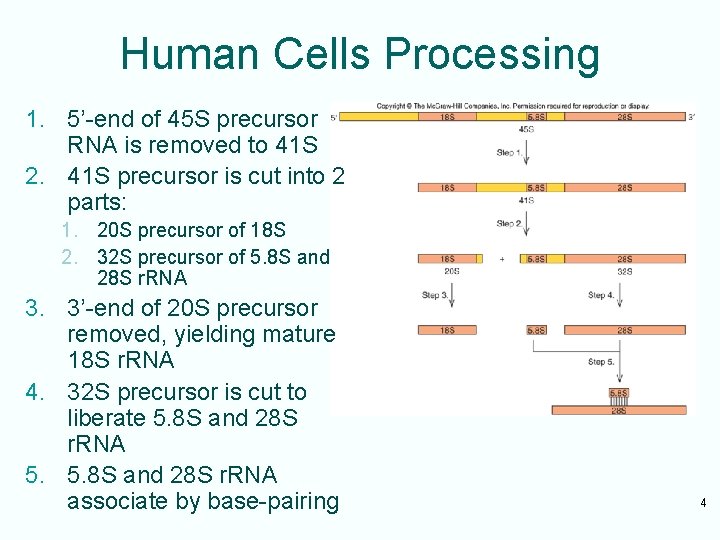
Human Cells Processing 1. 5’-end of 45 S precursor RNA is removed to 41 S 2. 41 S precursor is cut into 2 parts: 1. 20 S precursor of 18 S 2. 32 S precursor of 5. 8 S and 28 S r. RNA 3. 3’-end of 20 S precursor removed, yielding mature 18 S r. RNA 4. 32 S precursor is cut to liberate 5. 8 S and 28 S r. RNA 5. 5. 8 S and 28 S r. RNA associate by base-pairing 4

Bacterial r. RNA Processing • Bacterial r. RNA precursors contain t. RNA and all 3 r. RNA • r. RNA are released from their precursors by RNase III and RNase E – RNase III is the enzyme that performs at least the initial cleavages that separate the individual large r. RNAs – RNase E is another ribonuclease that is responsible for removing the 5 S r. RNA from the precursor 5
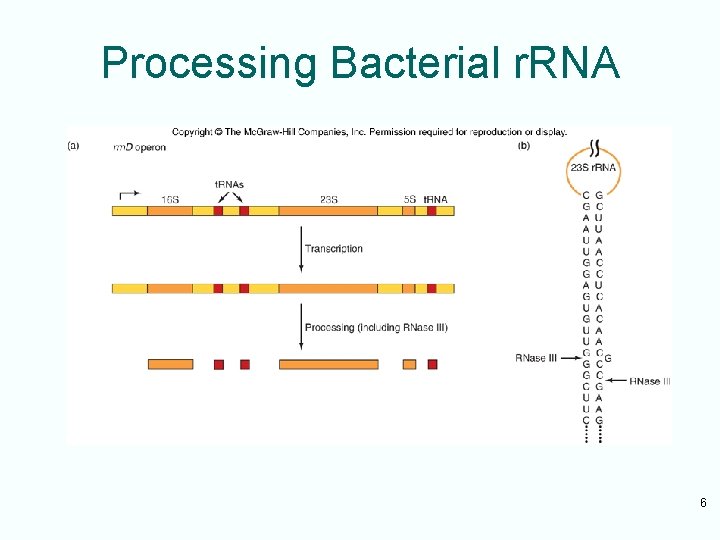
Processing Bacterial r. RNA 6

16. 2 Transfer RNA Processing • Transfer RNAs are made in all cells as overly long precursors – These must be processed by removing RNA at both ends • Nuclei of eukaryotes contain precursors of a single t. RNA • In bacteria, precursor may contain one or more t. RNA 7

Cutting Apart Polycistronic Precursors • In processing bacterial RNA that contain more than one t. RNA – First step is to cut precursor up into fragments with just one t. RNA each • Cutting between t. RNAs in precursors having 2 or more t. RNA • Cutting between t. RNAs and r. RNAs in precursors – Enzyme that performs both chores is the RNase III 8

Forming Mature 5’-Ends • Extra nucleotides are removed from the 5’ends of pre-t. RNA in one step by an endonucleolytic cleavage catalyzed by RNase P • RNase P from bacteria and eukaryotic nuclei have a catalytic RNA subunit called M 1 RNA • Spinach chloroplast RNase P appears to lack an RNA subunit 9

RNase P Action • RNase P makes a cut at the site that becomes mature 5’end of a t. RNA • This enzyme is all that is needed to form mature 5’-ends 10

Forming Mature 3’-Ends • RNase II and polynucleotide phosphorylase cooperate – To remove most of extra nucleotides at the end of a t. RNA precursor – Stopping at the +2 stage, with 2 extra nucleotides remaining • RNases PH and T are most active in removing the last 2 nucleotides from RNA – RNase T is the major participant in removing very last nucleotide 11
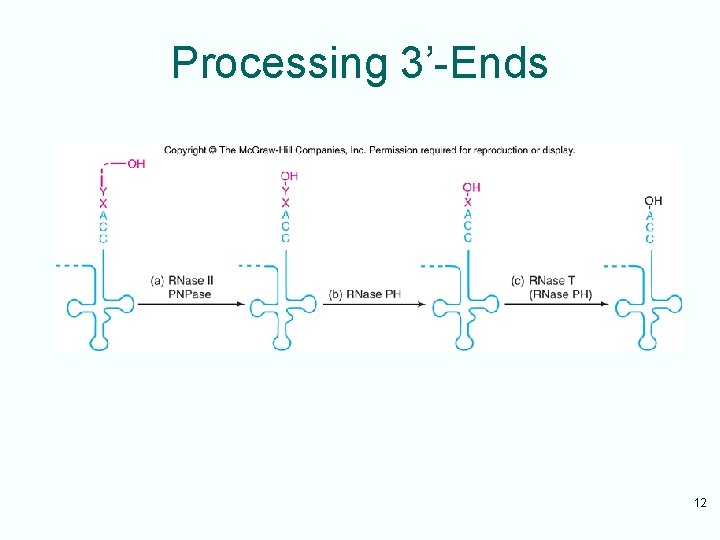
Processing 3’-Ends 12

16. 3 Trans-Splicing • Splicing that occurs in all eukaryotic species is called cis-splicing because it involves 2 or more exons that exist together in the same gene • Alternatively, trans-splicing has exons that are not part of the same gene at all, may not even be on the same chromosome 13

The Mechanism of Trans. Splicing • Trans-splicing occurs in several organisms – Parasitic and free-living worms – First discovered in trypanosomes • Trypanosome m. RNA are formed by transsplicing between a short leader exon and any one of many independent coding exons 14
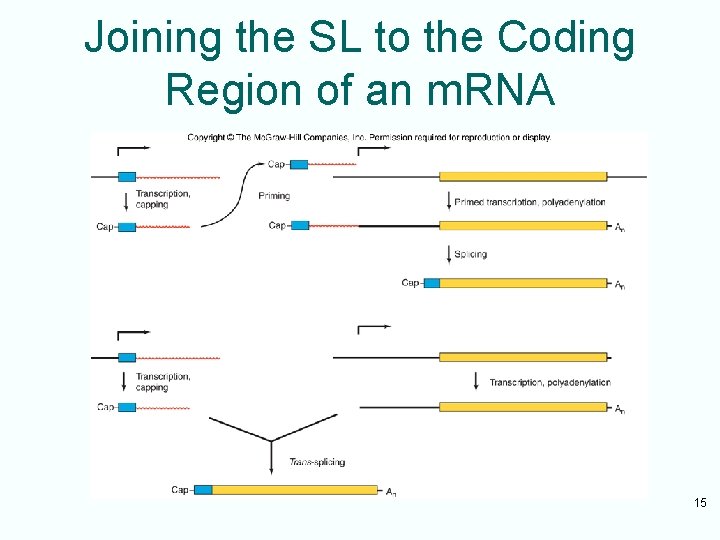
Joining the SL to the Coding Region of an m. RNA 15
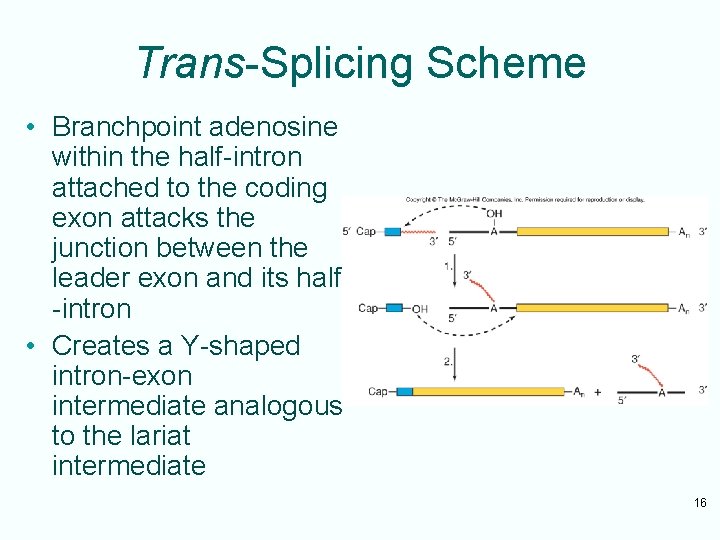
Trans-Splicing Scheme • Branchpoint adenosine within the half-intron attached to the coding exon attacks the junction between the leader exon and its half -intron • Creates a Y-shaped intron-exon intermediate analogous to the lariat intermediate 16

16. 4 RNA Editing • Pseudogenes are a duplicate copy of a gene that has been mutated so it does not function and is no longer used • Cryptogenes are incomplete genes • Trypanosomatid mitochondria encode incomplete m. RNA that must be edited before being translated • Editing occurs in the 3’ 5’ direction by successive action of one or more guide RNAs 17

Mechanism of Editing • Unedited transcripts can be found along with edited versions of the same m. RNAs • Editing occurs in the poly(A) tails of m. RNAs that are added posttranscriptionally • Partially edited transcripts have been isolated, always edited at their 3’-ends but not at their 5’-ends 18

Role of g. RNA in Editing • Guide RNAs (g. RNA) could direct the insertion and deletion of UMPs over a stretch of nucleotides in the m. RNA • When editing is done, g. RNA could hybridize near the 5’-end of newly edited region 19
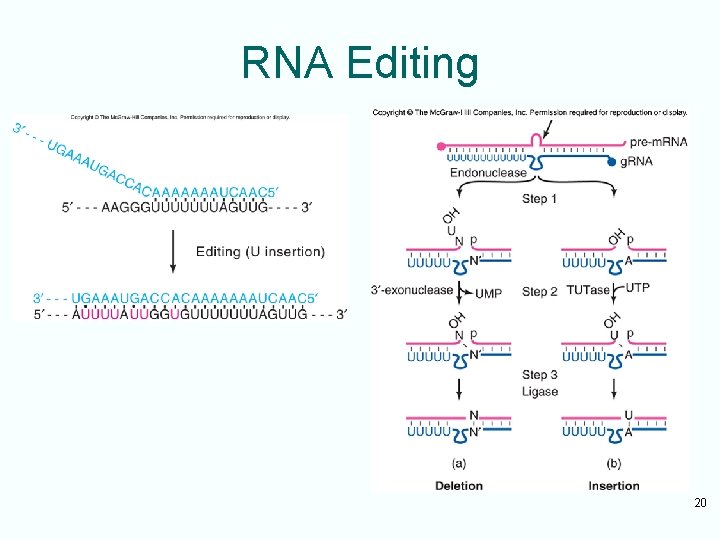
RNA Editing 20

Guide RNA Editing • 5’-end of the first g. RNA hybridizes to an unedited region at the 3’-border of editing I the pre-m. RNA • The 5’-ends of the rest of the g. RNAs hybridize to edited regions progressively closer to the 5’-end of the region to be edited in the pre-m. RNA • All of these g. RNAs provide A’s and G’s as templates for the incorporation of U’s missing from the m. RNA 21

Mechanism of Removing U’s • Sometimes the g. RNA is missing an A or G to pair with a U in the m. RNA – In this case the U is removed • Mechanism of removing U’s involves – Cutting pre-m. RNA just beyond U to be removed – Removal of U by exonuclease – Ligating the two pieces of pre-m. RNA together • Mechanism of adding U’s uses same first and last step • Middle step involves addition of one or more U’s from UTP by TUTase 22

Editing by Nucleotide Deamination • Some adenosines in m. RNAs of higher eukaryotes, including fruit flies and mammals, must be deaminated to inosine posttranscriptionally for m. RNA to code for proper proteins • Enzymes know as adenosine deaminases active on RNAs (ADARs) carry out this kind of RNA editing • Some cytidines must be deaminated to uridine for an m. RNA to code properly 23

16. 5 Posttranscriptional Control of Gene Expression • A common form of posttranscriptional control of gene expression is control of m. RNA stability • When mammary gland tissue is stimulated by prolactin, synthesis of casein protein increases dramatically – Most of this increase in casein is not due to increased rate of transcription of the casein gene – It is caused by an increase in half-life of casein m. RNA 24

Transferrin Receptor m. RNA Stability • Transferrin receptor-Tf. R concentration is low when iron concentration is high • Loss of Tf. R is largely due to decreased stability of the Tf. R m. RNA • Response to iron depends on the 3’-UTR of the m. RNA which contains 5 stem loops called iron response elements (IREs) 25

Rapid Turnover Determinant • Instability of this m. RNA is caused by a rapid turnover determinant that lies in the 3’-UTR • Iron response elements A and E along with the large central loop of the Tf. R 3’-UTR can be deleted without altering the response to iron • Removing all of the IREs, or either 1 or 2 non. IRE stem loops renders the Tf. R m. RNA constitutively stable • Each of the non-IRE stem loops and one of IREs B to D are part of a rapid turnover determinant • Removing a C from IREs B-D render the Tf. R m. RNA constitutively unstable, unable to bind 26

Determining Tf. R m. RNA Stability • When iron concentration is high, Tf. R m. RNA decays rapidly • When iron concentration is low, Tf. R m. RNA decays much more slowly – Difference is about 20 -fold – Important in determining the rate of accumulation of Tf. R m. RNA • TRS-3 m. RNA – Has no rapid turnover determinant – Is constitutively stable • TRS-4 m. RNA has no ability to bind IRE-binding protein and is constitutively unstable 27

Tf. R m. RNA Degradation Pathway • Initiating event in Tf. R m. RNA degradation seems to be endonucleolytic cleavage of m. RNA more than 1000 nt from its 3’-end within the IRE region • Cleavage does not require prior deadenylation of m. RNA – Iron controls Tf. R m. RNA stability 28

Role of Iron Level • When iron concentration is low, aconitase exists in an apoprotein form lacking iron – Binds to the IREs in the Tf. R m. RNA and protects RNA against attack by RNase • High iron concentration – Aconitase apoprotein binds to iron – Cannot bind to Tf. R m. RNA IREs – Leaves RNA vulnerable to degradation 29
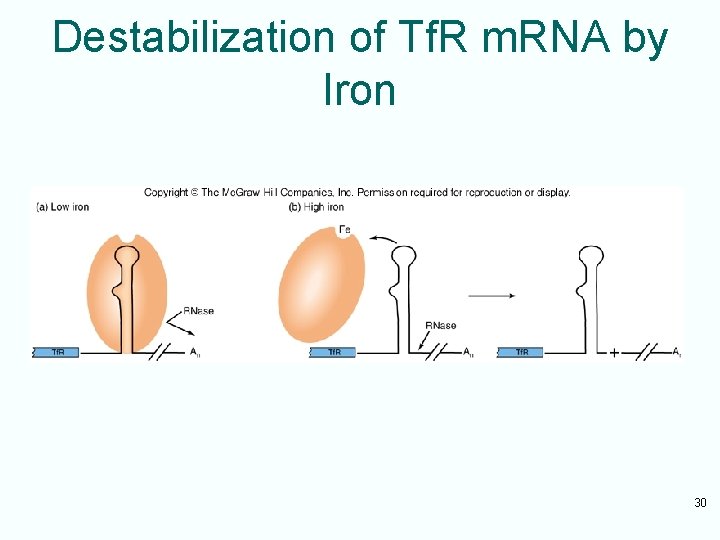
Destabilization of Tf. R m. RNA by Iron 30

RNA Interference • RNA interference occurs when a cell encounters ds. RNA from: – Virus – Transposon – Transgene • Trigger ds. RNA is degraded into 21 -23 nt fragments (si. RNAs) by an RNase III-like enzyme called Dicer • Double-stranded si. RNA, with Dicer and Dicerassociated protein R 2 D 2 form a complex called complex B 31

Complex B • Complex B delivers the si. RNA to the RISC loading complex (RLC) – Separates 2 strands of si. RNA – Transfers guide strand to RNA-induced silencing complex (RISC) that introduces a protein, Ago 2 • The guide strand of si. RNA base-pairs with target m. RNA in the active site of PIWI domain of Ago 2 – Ago 2 is an RNase H-like enzyme known as a slicer – Slicer cleaves the target m. RNA in middle of the region of its base-pairing with the si. RNA – ATP-dependent step has cleaved RNA ejected from RISC which then accepts a new molecule of m. RNA for degradation 32

Amplification of si. RNA • si. RNA is amplified during RNAi when antisense si. RNAs hybridize to target m. RNA and prime synthesis of full-length antisense RNA by RNA-dependent RNA polymerase • New ds. RNA is digested by Dicer into new pieces of si. RNA 33

Role of RNAi Machinery in Heterochromatin Formation • RNAi machinery is involved in heterochromatization at yeast centromeres and silent mating-type regions • In fission, yeast at the outermost regions of centromeres active transcription of the reverse strand occurs – Forward transcripts can base-pair with the reverse transcript to kick off RNAi – In turn this recruits a histone methyltransferase that methylates Lys-9 of H 3 – This recruits Swi 6 causing heterochromatization 34

Heterochromatin Formation in Plants and Mammals • In plants and mammals, formation of heterochromatin is abetted by DNA methylation • This methylation can also attract heterochromatization machinery • Individual genes can be silenced in mammals by RNAi that targets the gene’s control region rather than the coding region • Silencing process involves DNA methylation rather than m. RNA destruction 35

Micro. RNAs and Gene Silencing • Micro. RNAs are 18 -25 nt RNAs produced naturally in plant and animal cells by cleavage from 75 -nt stem-loop precursor RNA • In last step of mi. RNA synthesis, Dicer RNase cleaves ds stem part of the precursor to yield mi. RNA in ds form • Single-stranded form of mi. RNAs can team up with Argonaute protein in a RISC to control gene expression by base-pairing to their m. RNAs 36
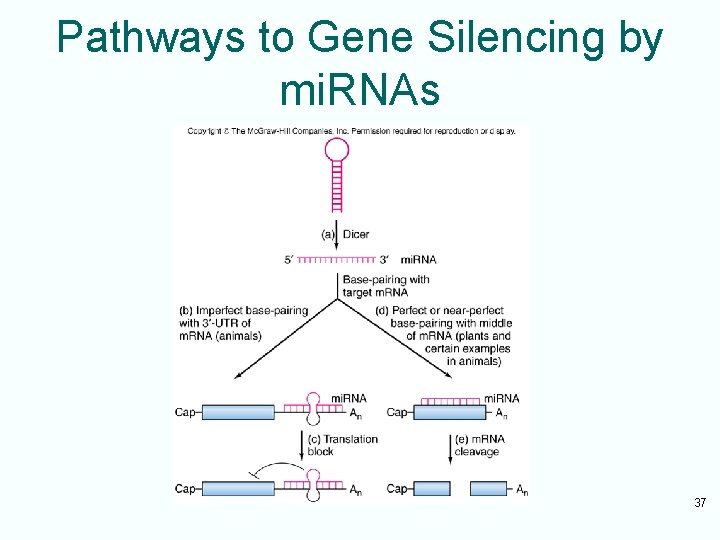
Pathways to Gene Silencing by mi. RNAs 37

Animal and Plant mi. RNA • In animals, mi. RNAs tend to base-pair imperfectly to the 3’-UTRs of their target m. RNAs • This leads to inhibition of protein product accumulation of such m. RNA • In plants, mi. RNAs tend to base-pair perfectly or near-perfectly with target m. RNAs • There are some exceptions which can lead to translation blockage 38
- Slides: 38