Lecture 9 January 23 2016 Biotech 3 Part

Lecture 9 January 23, 2016 Biotech 3

Part I – Module II: Calculate the transformation efficiency of the p. UC 8 ligated vector Transformation Efficiency – calculated by quantitating the number of bacteria (CFU) that were successfully transformed per microgram (μg) of DNA. Reported as CFU/ μg. Dependent on many factors: • Type of bacteria • Bacterial growth phase • Amount of plasmid used • DNA tertiary structure (supercoiled) • Heat shock temperature • Method used to transform cells • Example: 108 vs 105 CFU/ug (electroporation vs chemically competent cells)
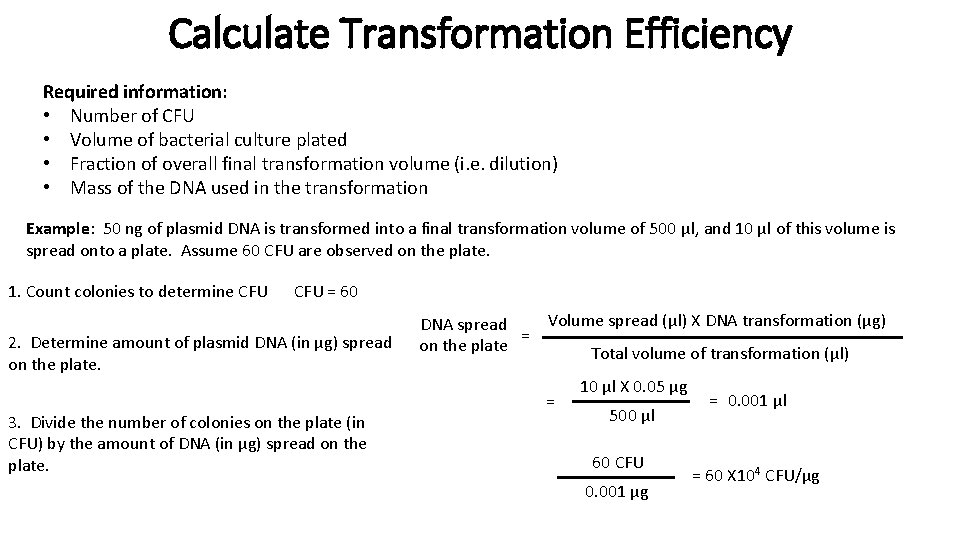
Calculate Transformation Efficiency Required information: • Number of CFU • Volume of bacterial culture plated • Fraction of overall final transformation volume (i. e. dilution) • Mass of the DNA used in the transformation Example: 50 ng of plasmid DNA is transformed into a final transformation volume of 500 μl, and 10 μl of this volume is spread onto a plate. Assume 60 CFU are observed on the plate. 1. Count colonies to determine CFU = 60 2. Determine amount of plasmid DNA (in μg) spread on the plate. 3. Divide the number of colonies on the plate (in CFU) by the amount of DNA (in μg) spread on the plate. Volume spread (μl) X DNA transformation (μg) DNA spread = on the plate Total volume of transformation (μl) = 10 μl X 0. 05 μg 500 μl 60 CFU 0. 001 μg = 0. 001 μl = 60 X 104 CFU/μg
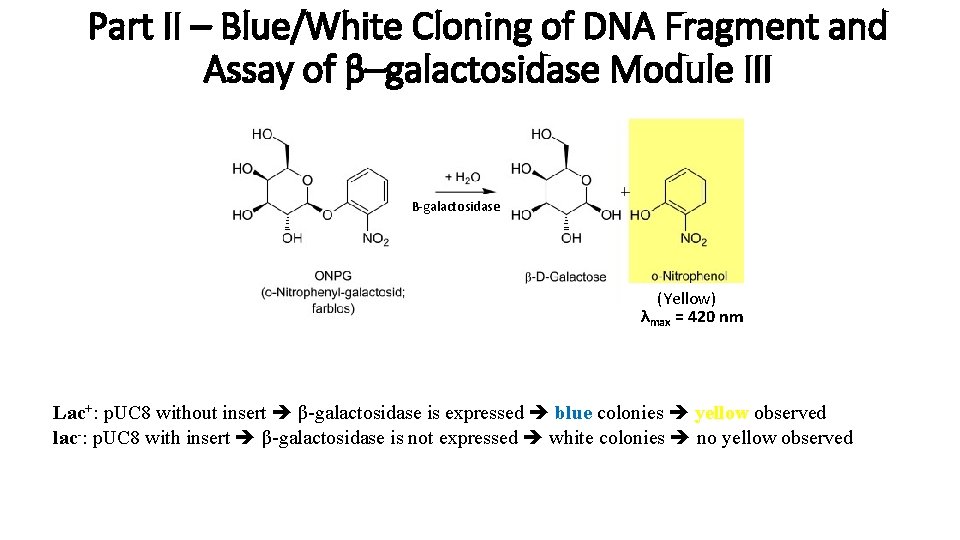
Part II – Blue/White Cloning of DNA Fragment and Assay of β–galactosidase Module III B-galactosidase (Yellow) λmax = 420 nm Lac+: p. UC 8 without insert β-galactosidase is expressed blue colonies yellow observed lac-: p. UC 8 with insert β-galactosidase is not expressed white colonies no yellow observed

Part II – Blue/White Cloning of DNA Fragment and Assay of β–galactosidase Module III calculation Background: 1972, Jeffrey Miller published "Experiments in Molecular Genetics" which contained a protocol for determining the amount of β-Gal with ONPG. Because of this, ONPG/βGal assays are referred to as "Miller" assays, and a standardized amount of β-Gal activity is a "Miller Unit“. 1 Miller Unit = 1000 X A 420 – (1. 75 x A 550)) T x V x A 600 A 420 is the absorbance of the yellow ONP A 550 is the scatter from cell debris, which when multiplied by 1. 75 approximates the scatter observed at A 420 T = reaction time in minutes V = volume of culture assayed A 600 reflects cell density
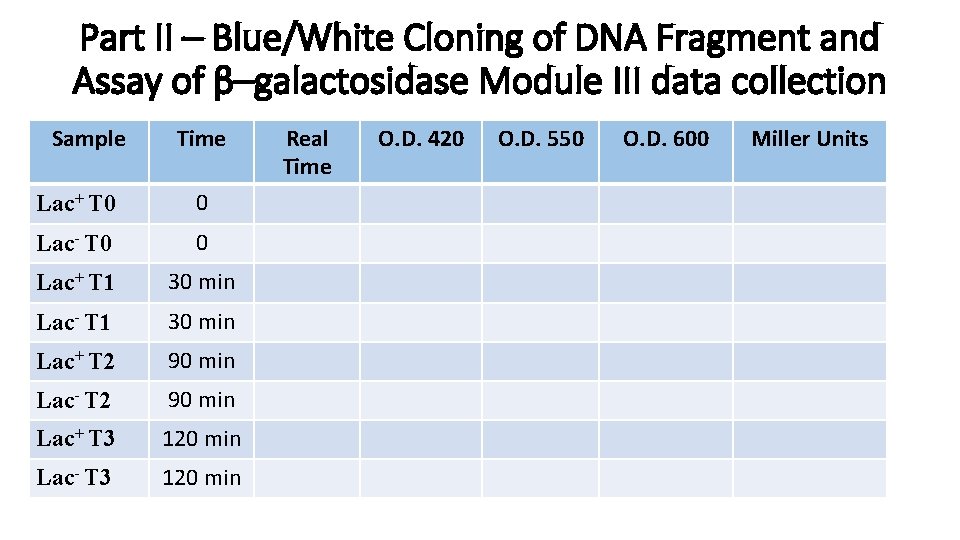
Part II – Blue/White Cloning of DNA Fragment and Assay of β–galactosidase Module III data collection Sample Time Lac+ T 0 0 Lac- T 0 0 Lac+ T 1 30 min Lac- T 1 30 min Lac+ T 2 90 min Lac- T 2 90 min Lac+ T 3 120 min Lac- T 3 120 min Real Time O. D. 420 O. D. 550 O. D. 600 Miller Units
- Slides: 6