Lecture 7 DNA repair Chapter 10 Problems 2

Lecture 7 DNA repair Chapter 10 Problems 2, 4, 6, 8, 10, 12, and 14 Quiz 3 due today at 4: 00 PM
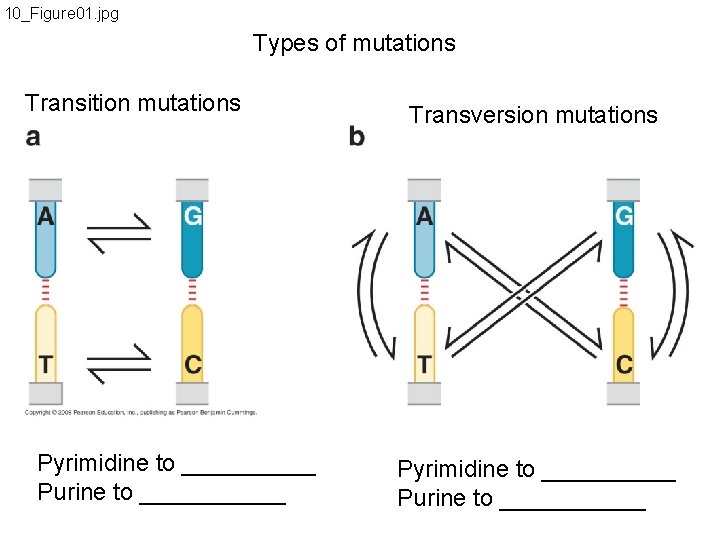
10_Figure 01. jpg Types of mutations Transition mutations Pyrimidine to _____ Purine to ______ Transversion mutations Pyrimidine to _____ Purine to ______
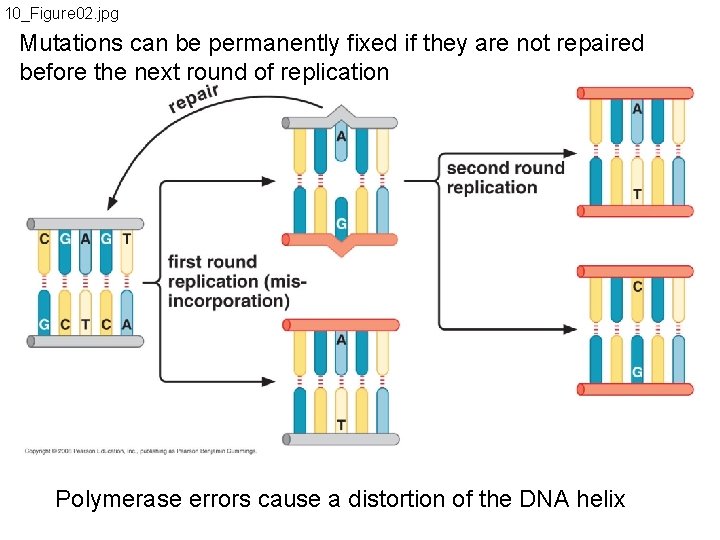
10_Figure 02. jpg Mutations can be permanently fixed if they are not repaired before the next round of replication Polymerase errors cause a distortion of the DNA helix
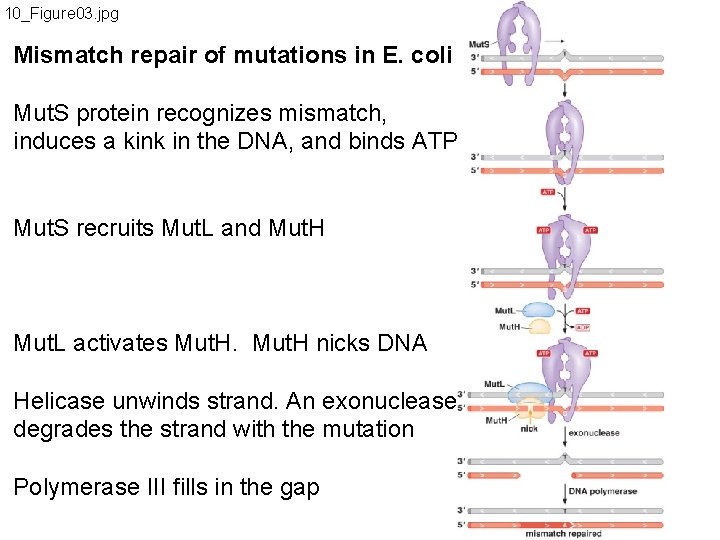
10_Figure 03. jpg Mismatch repair of mutations in E. coli Mut. S protein recognizes mismatch, induces a kink in the DNA, and binds ATP Mut. S recruits Mut. L and Mut. H Mut. L activates Mut. H nicks DNA Helicase unwinds strand. An exonuclease degrades the strand with the mutation Polymerase III fills in the gap
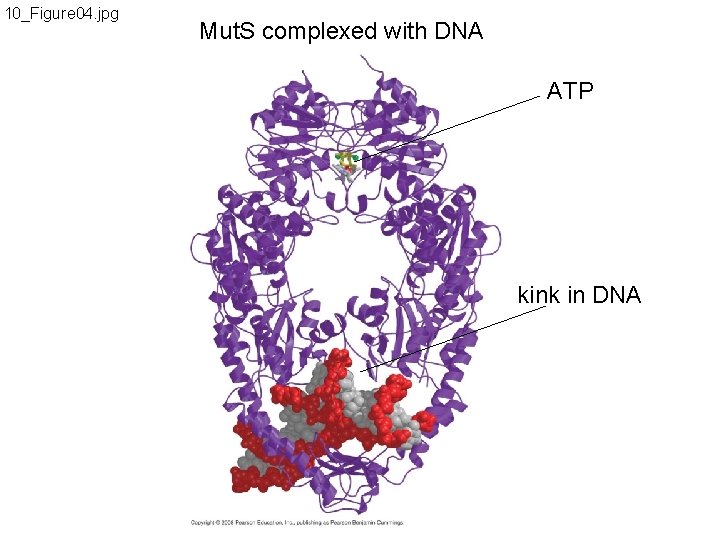
10_Figure 04. jpg Mut. S complexed with DNA ATP kink in DNA
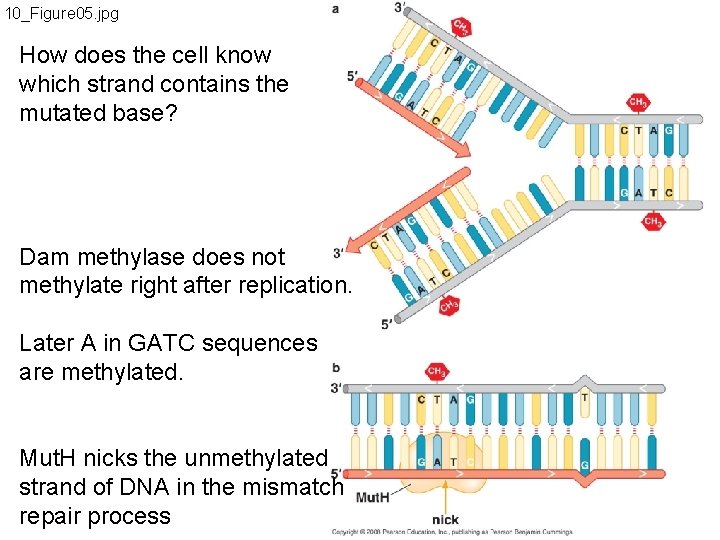
10_Figure 05. jpg How does the cell know which strand contains the mutated base? Dam methylase does not methylate right after replication. Later A in GATC sequences are methylated. Mut. H nicks the unmethylated strand of DNA in the mismatch repair process
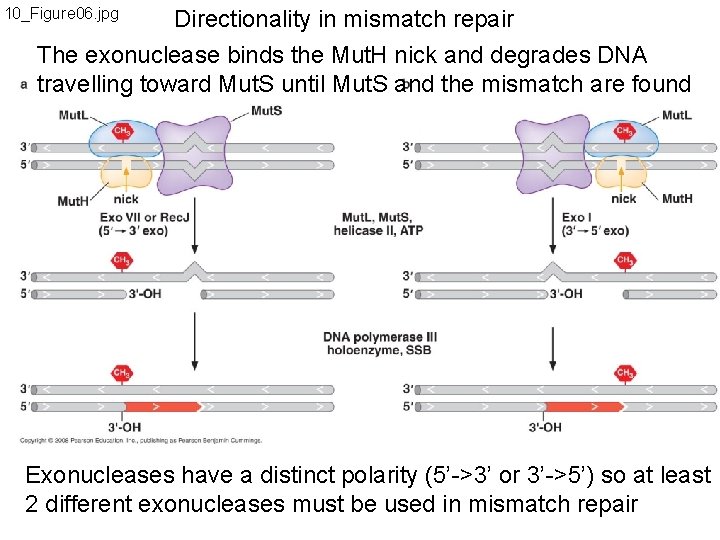
10_Figure 06. jpg Directionality in mismatch repair The exonuclease binds the Mut. H nick and degrades DNA travelling toward Mut. S until Mut. S and the mismatch are found Exonucleases have a distinct polarity (5’->3’ or 3’->5’) so at least 2 different exonucleases must be used in mismatch repair

Eukaryotic cells lack a Mut. H homolog Then how do the mismatch repair enzymes recognize the newly synthesized strand of DNA? Nicks present in unligated Okazaki fragments may play this role following lagging strand DNA synthesis. But how nicks get introduced in the leading strand DNA is not quite clear.
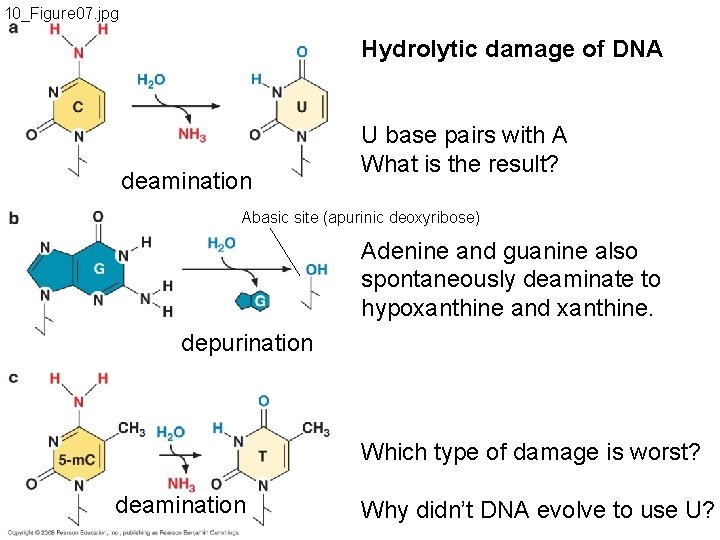
10_Figure 07. jpg Hydrolytic damage of DNA deamination U base pairs with A What is the result? Abasic site (apurinic deoxyribose) Adenine and guanine also spontaneously deaminate to hypoxanthine and xanthine. depurination Which type of damage is worst? deamination Why didn’t DNA evolve to use U?
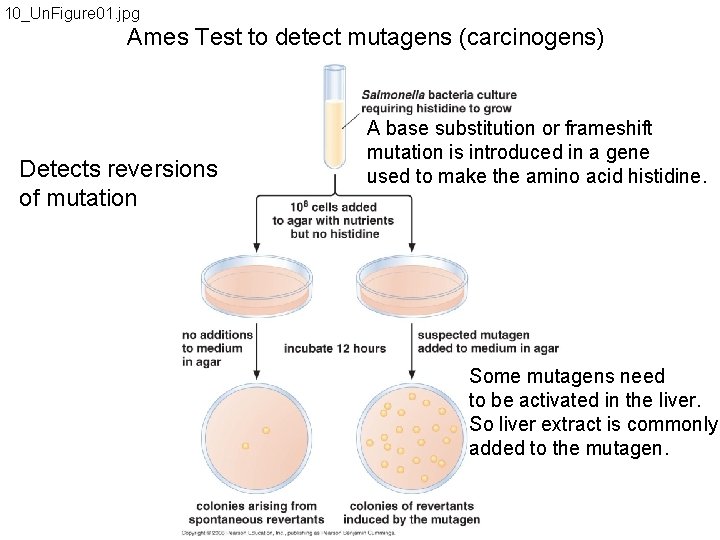
10_Un. Figure 01. jpg Ames Test to detect mutagens (carcinogens) Detects reversions of mutation A base substitution or frameshift mutation is introduced in a gene used to make the amino acid histidine. Some mutagens need to be activated in the liver. So liver extract is commonly added to the mutagen.
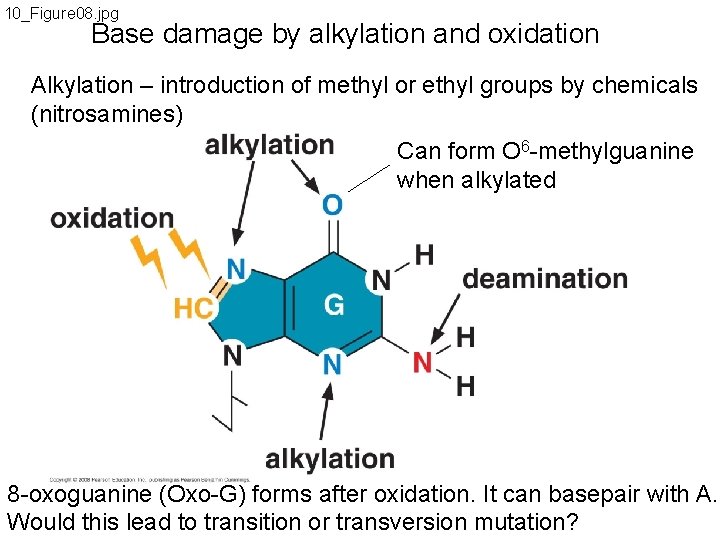
10_Figure 08. jpg Base damage by alkylation and oxidation Alkylation – introduction of methyl or ethyl groups by chemicals (nitrosamines) Can form O 6 -methylguanine when alkylated 8 -oxoguanine (Oxo-G) forms after oxidation. It can basepair with A. Would this lead to transition or transversion mutation?
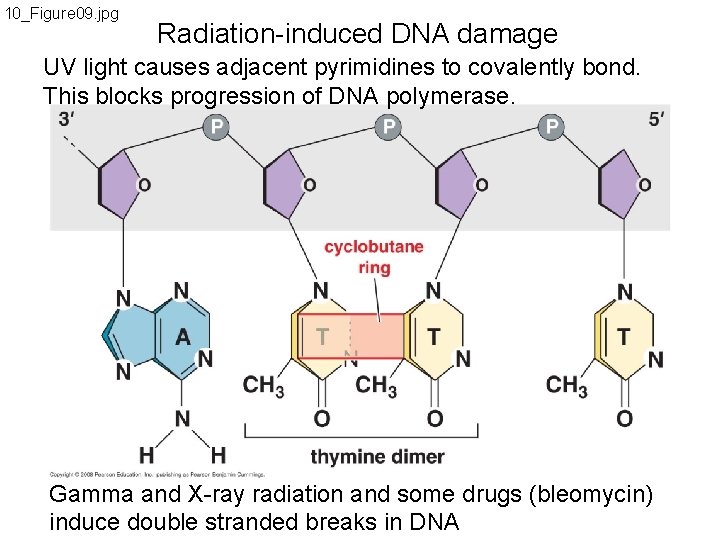
10_Figure 09. jpg Radiation-induced DNA damage UV light causes adjacent pyrimidines to covalently bond. This blocks progression of DNA polymerase. Gamma and X-ray radiation and some drugs (bleomycin) induce double stranded breaks in DNA
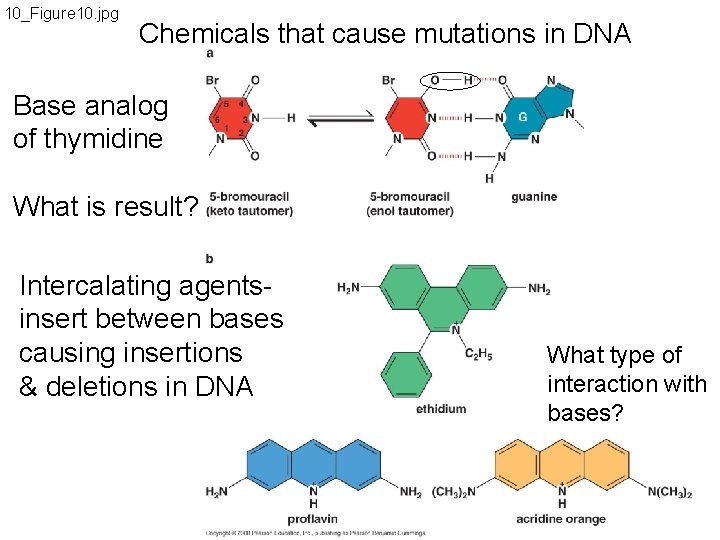
10_Figure 10. jpg Chemicals that cause mutations in DNA Base analog of thymidine What is result? Intercalating agentsinsert between bases causing insertions & deletions in DNA What type of interaction with bases?

10_Table 01. jpg DNA repair systems 10
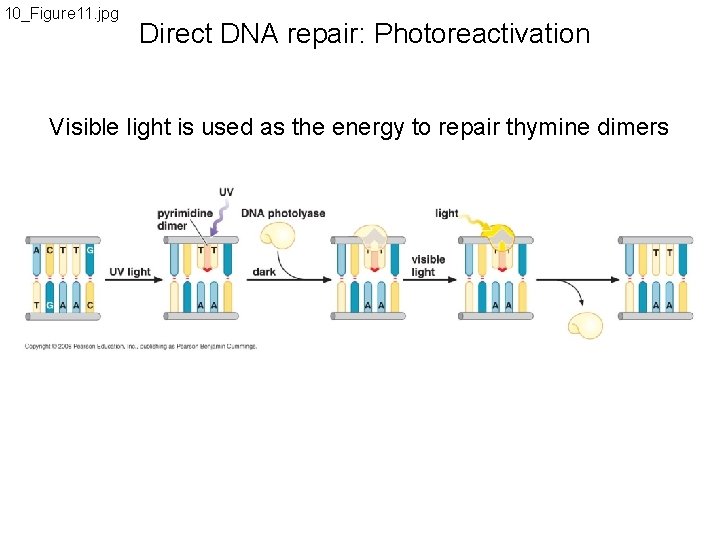
10_Figure 11. jpg Direct DNA repair: Photoreactivation Visible light is used as the energy to repair thymine dimers
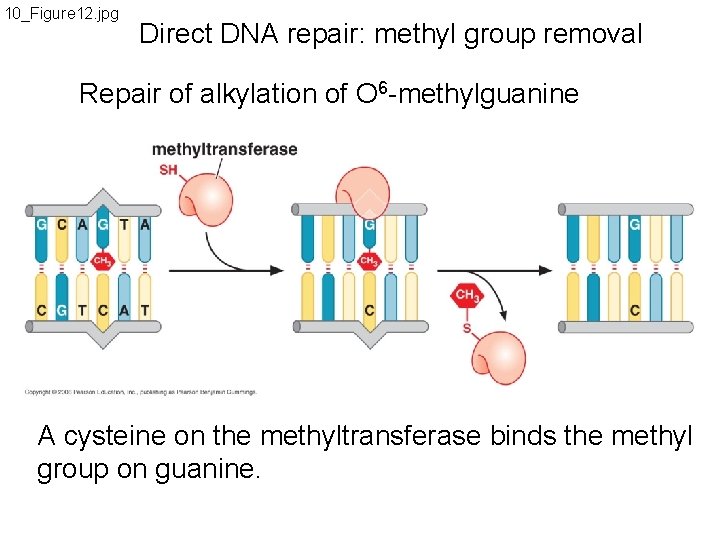
10_Figure 12. jpg Direct DNA repair: methyl group removal Repair of alkylation of O 6 -methylguanine A cysteine on the methyltransferase binds the methyl group on guanine.
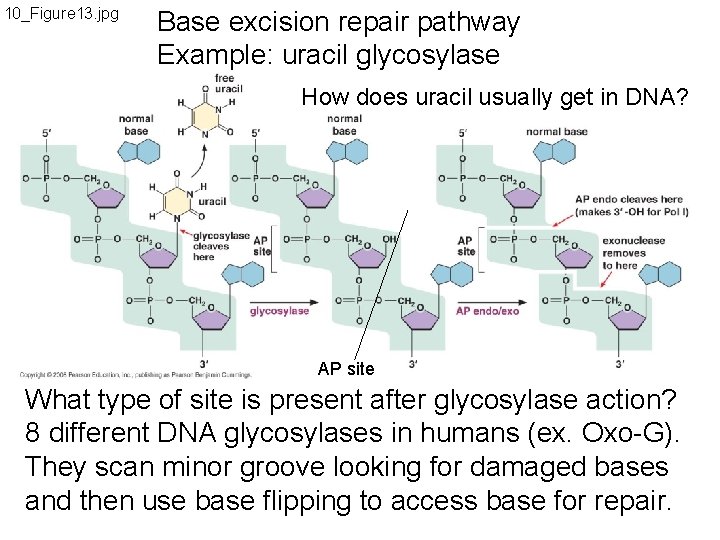
10_Figure 13. jpg Base excision repair pathway Example: uracil glycosylase How does uracil usually get in DNA? AP site What type of site is present after glycosylase action? 8 different DNA glycosylases in humans (ex. Oxo-G). They scan minor groove looking for damaged bases and then use base flipping to access base for repair.
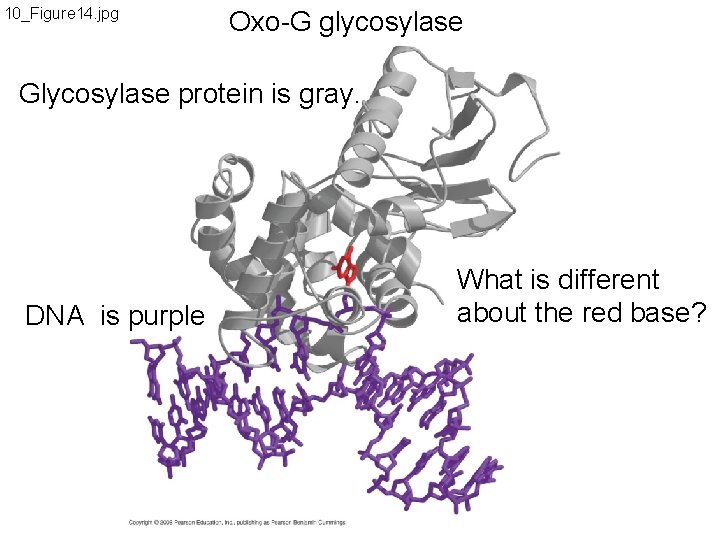
10_Figure 14. jpg Oxo-G glycosylase Glycosylase protein is gray. DNA is purple What is different about the red base?
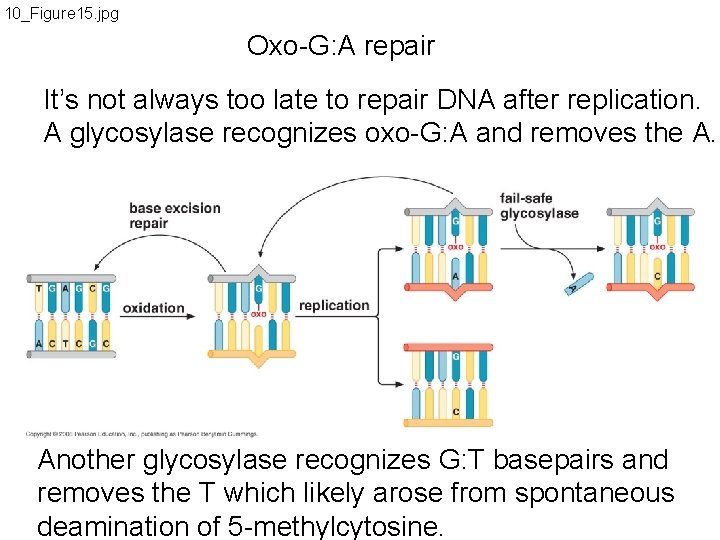
10_Figure 15. jpg Oxo-G: A repair It’s not always too late to repair DNA after replication. A glycosylase recognizes oxo-G: A and removes the A. Another glycosylase recognizes G: T basepairs and removes the T which likely arose from spontaneous deamination of 5 -methylcytosine.
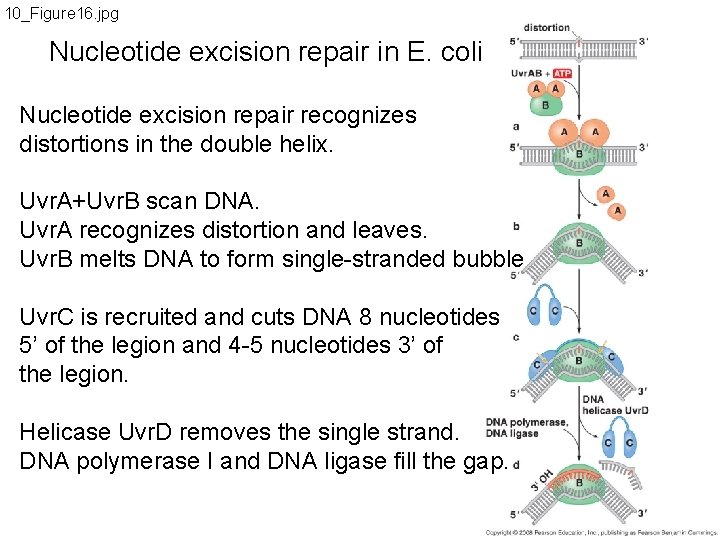
10_Figure 16. jpg Nucleotide excision repair in E. coli Nucleotide excision repair recognizes distortions in the double helix. Uvr. A+Uvr. B scan DNA. Uvr. A recognizes distortion and leaves. Uvr. B melts DNA to form single-stranded bubble Uvr. C is recruited and cuts DNA 8 nucleotides 5’ of the legion and 4 -5 nucleotides 3’ of the legion. Helicase Uvr. D removes the single strand. DNA polymerase I and DNA ligase fill the gap.

Nucleotide excision repair (NER) in humans Similar but more complex than NER in E. Coli. Mammalian NER uses around 25 proteins. XPC recognizes distortions (like Uvr. A in E. coli). XPA and XPD helicases melt DNA (like Uvr. B in E. coli). Single stranded binding protein RPA binds DNA. 5’-cleavage site cut by ERCC 1 -XPF nuclease and 3’-cleavage site cut by XPG nuclease (similar to Uvr. C in E. coli) 24 -32 nucleotide long DNA strand is released that is filled in by a polymerase and sealed by DNA ligase. Xeroderma pigmentosum disease caused by mutations in XP_ (NER) genes. Patients are susceptiple to cancer from UV light.
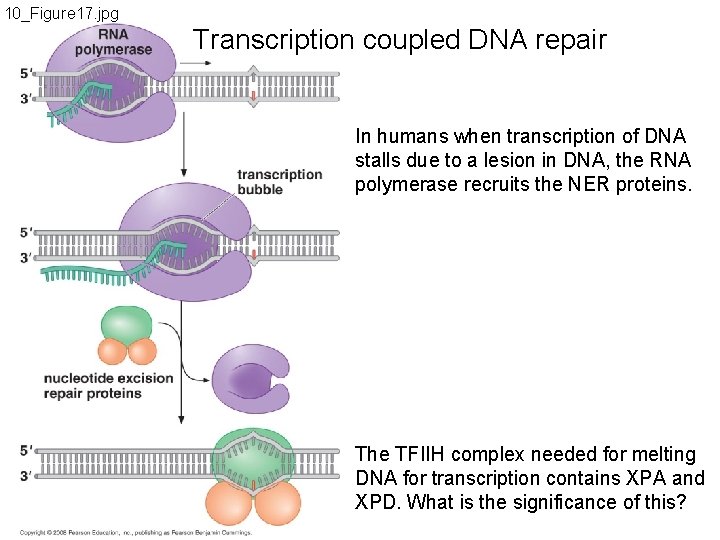
10_Figure 17. jpg Transcription coupled DNA repair In humans when transcription of DNA stalls due to a lesion in DNA, the RNA polymerase recruits the NER proteins. The TFIIH complex needed for melting DNA for transcription contains XPA and XPD. What is the significance of this?
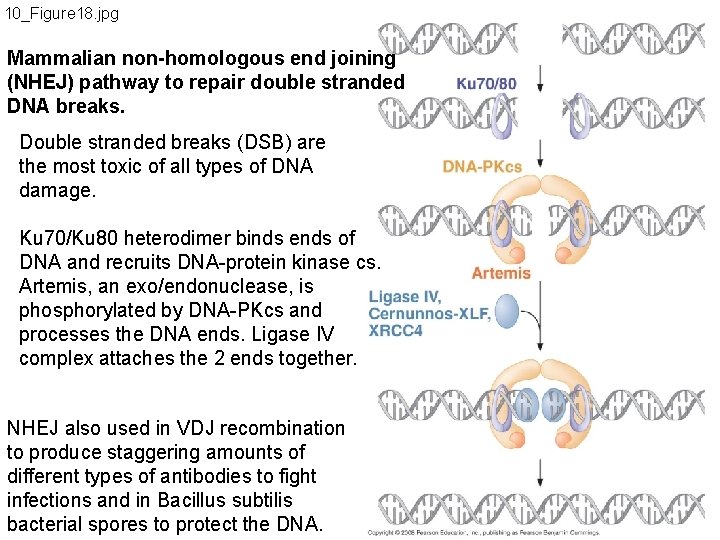
10_Figure 18. jpg Mammalian non-homologous end joining (NHEJ) pathway to repair double stranded DNA breaks. Double stranded breaks (DSB) are the most toxic of all types of DNA damage. Ku 70/Ku 80 heterodimer binds ends of DNA and recruits DNA-protein kinase cs. Artemis, an exo/endonuclease, is phosphorylated by DNA-PKcs and processes the DNA ends. Ligase IV complex attaches the 2 ends together. NHEJ also used in VDJ recombination to produce staggering amounts of different types of antibodies to fight infections and in Bacillus subtilis bacterial spores to protect the DNA.
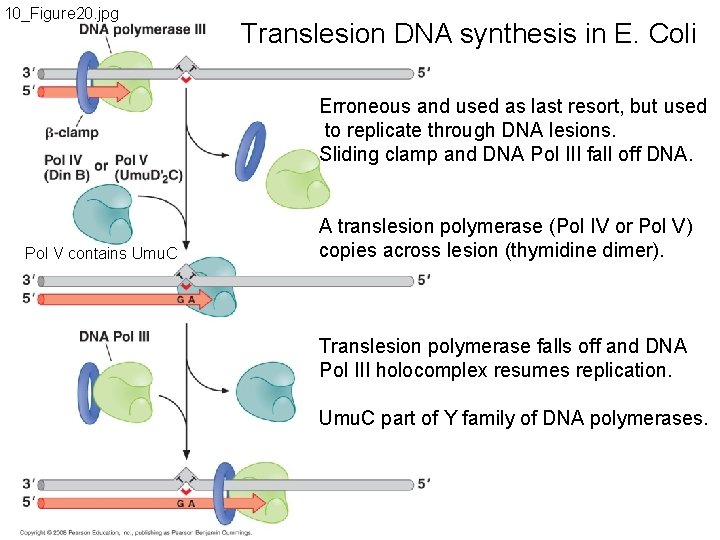
10_Figure 20. jpg Translesion DNA synthesis in E. Coli Erroneous and used as last resort, but used to replicate through DNA lesions. Sliding clamp and DNA Pol III fall off DNA. Pol V contains Umu. C A translesion polymerase (Pol IV or Pol V) copies across lesion (thymidine dimer). Translesion polymerase falls off and DNA Pol III holocomplex resumes replication. Umu. C part of Y family of DNA polymerases.
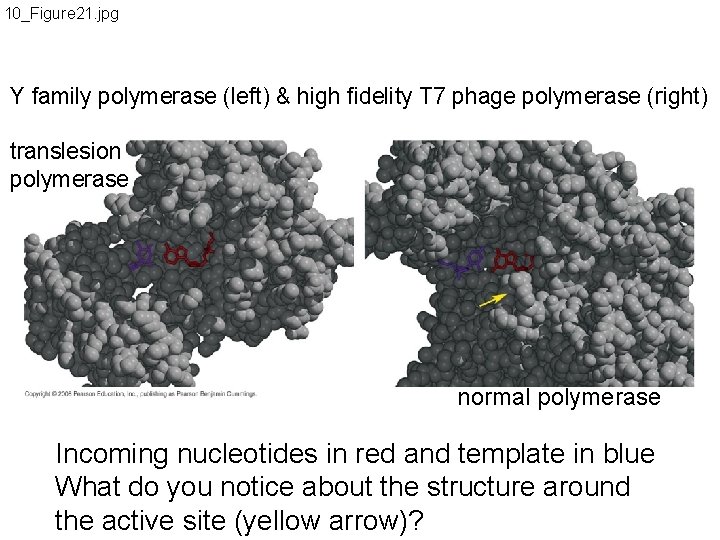
10_Figure 21. jpg Y family polymerase (left) & high fidelity T 7 phage polymerase (right) translesion polymerase normal polymerase Incoming nucleotides in red and template in blue What do you notice about the structure around the active site (yellow arrow)?
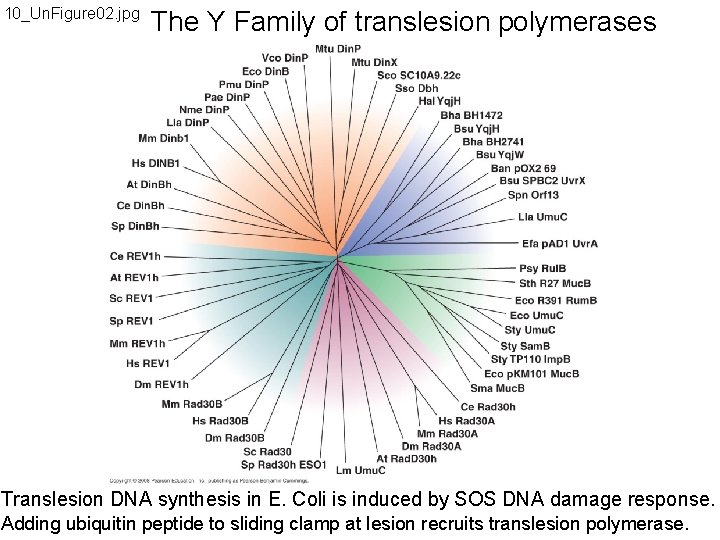
10_Un. Figure 02. jpg The Y Family of translesion polymerases Translesion DNA synthesis in E. Coli is induced by SOS DNA damage response. Adding ubiquitin peptide to sliding clamp at lesion recruits translesion polymerase.

10_Table 01. jpg DNA repair systems 10

Double strand break repair pathway (homologous recombination) Uses information from homologous sister chromosome and will be discussed next lecture (chapter 11)
- Slides: 28