Lecture 6 January 12 2016 Biotech 3 Lecture

Lecture 6 January 12, 2016 Biotech 3

Lecture Topics 1. - Cloning Vectors Plasmids Bacteriophage Cosmids BACS and YACs 2. Plasmids - Origin of Replication - Selectivity Markers - Multiple Cloning Site - Promoter Sequence 3. PCR - General overview and History: 1983 Kary Mullis - Mathematical concept and cycle number - Polymerase rate and fidelity - Forward and Reverse Primer design - Affinity tags - Tm - Secondary structure - Primer design for point mutations - PCR Parameters: DNA sequence, primers, type of polymerase, melting temperature, extension time, number of cycles

A. Cloning Vectors Cloning vector: small piece of DNA that can be stably maintained in an organism and which a foreign piece of DNA can be inserted for cloning purposes. Examples: 1. Plasmids 2. Bacteriophage 3. Cosmids 4. Bacterial Artificial Chromosomes (BAC) 5. Yeast Artificial Chromosomes (YAC)

Plasmids Plasmid: Small circular DNA that can replicate independently from a host’s chromosome. Characteristics: - Smaller than chromosome (<1/20 th the size) - Contain “non-essential genes” - Size varies 1 kb – 12, 000 kb - Copy number is variable (1 – 200 copies per cell depending on the plasmid) - Some can integrate into the chromosome - Cloning vectors can carry up to ~15 kb We’ll break down the anatomy of a plasmid cloning vector shortly!
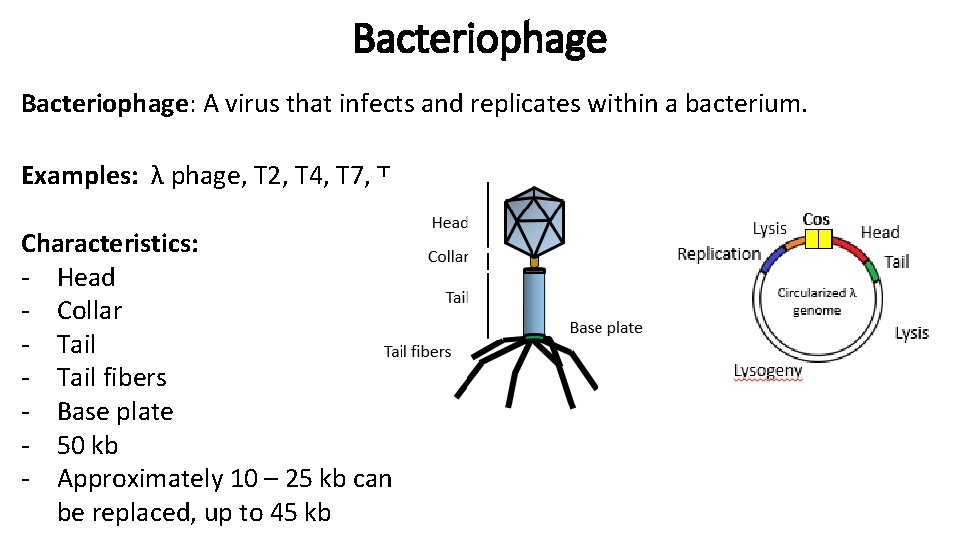
Bacteriophage: A virus that infects and replicates within a bacterium. Examples: λ phage, T 2, T 4, T 7, T Characteristics: - Head - Collar - Tail fibers - Base plate - 50 kb - Approximately 10 – 25 kb can be replaced, up to 45 kb
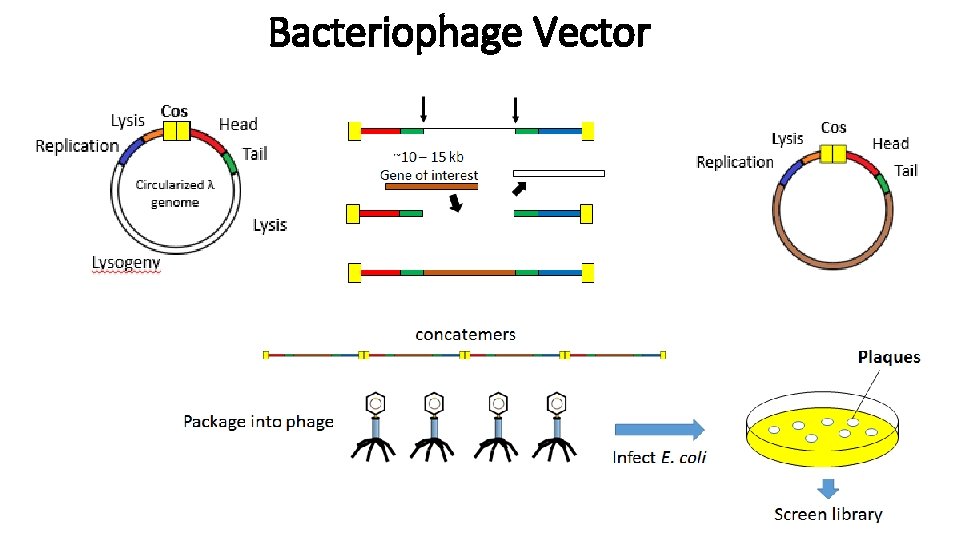
Bacteriophage Vector
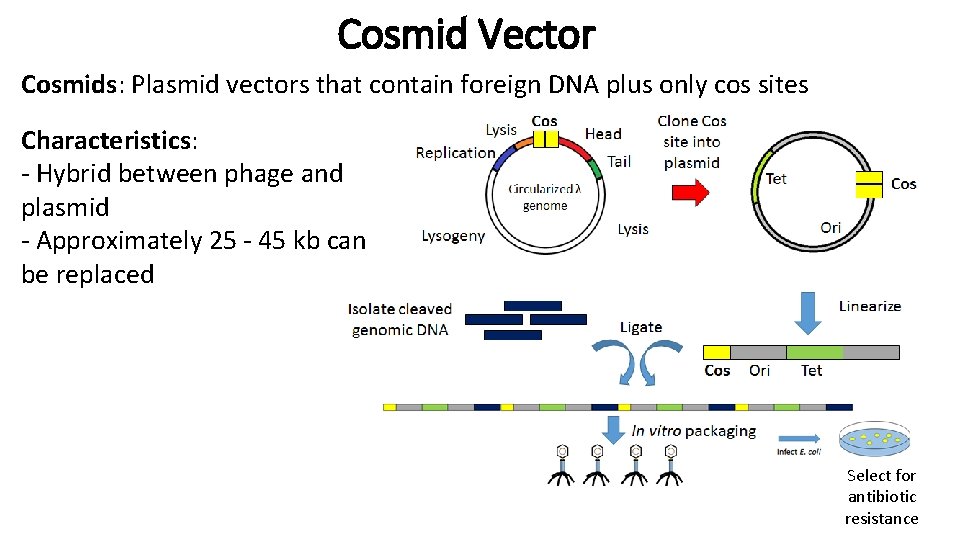
Cosmid Vector Cosmids: Plasmid vectors that contain foreign DNA plus only cos sites Characteristics: - Hybrid between phage and plasmid - Approximately 25 - 45 kb can be replaced Select for antibiotic resistance

BAC and YAC Vectors Bacterial Artificial Chromosome (BAC): - Based on the 7 kb F factor of E. coli - Can carry 100 – 300 kb Yeast Artificial Chromosome (YAC): - Very high carrying capacity; up to 1000 kb - Resemble normal yeast chromosome: Telomeres, centromere - Yeast as a host (not E. coli as previous examples)
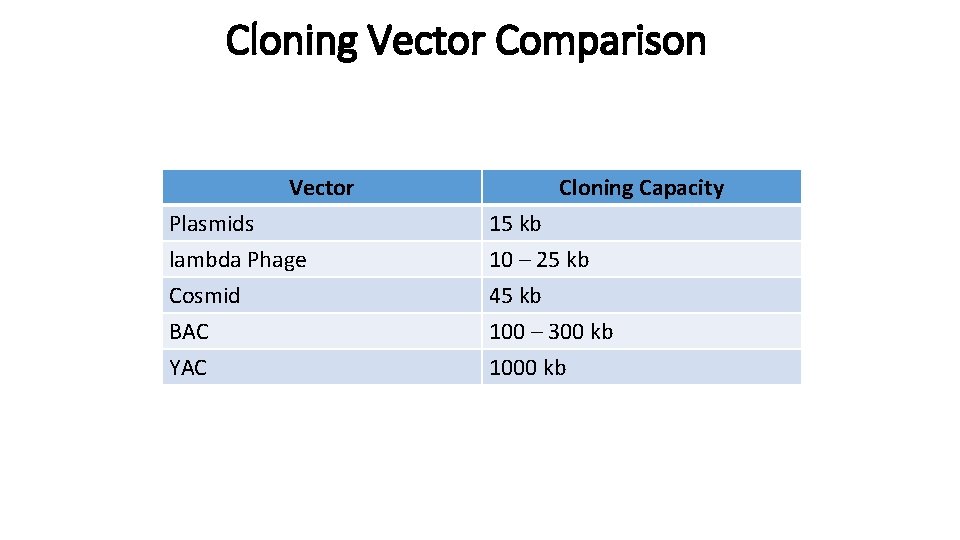
Cloning Vector Comparison Vector Cloning Capacity Plasmids lambda Phage Cosmid 15 kb 10 – 25 kb 45 kb BAC YAC 100 – 300 kb 1000 kb
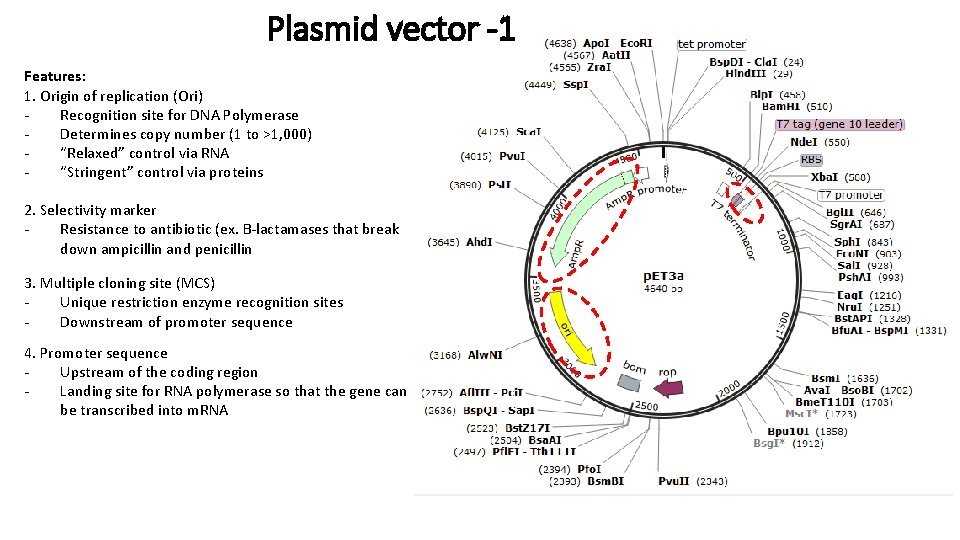
Plasmid vector -1 Features: 1. Origin of replication (Ori) Recognition site for DNA Polymerase Determines copy number (1 to >1, 000) “Relaxed” control via RNA “Stringent” control via proteins 2. Selectivity marker Resistance to antibiotic (ex. B-lactamases that break down ampicillin and penicillin 3. Multiple cloning site (MCS) Unique restriction enzyme recognition sites Downstream of promoter sequence 4. Promoter sequence Upstream of the coding region Landing site for RNA polymerase so that the gene can be transcribed into m. RNA

Features: 1. Origin of replication (Ori) Recognition site for DNA Polymerase Determines copy number (1 to >1, 000) “Relaxed” control via RNA “Stringent” control via proteins Plasmid Vector - 2 2. Selectivity marker Resistance to antibiotic (ex. B-lactamases that break down ampicillin and penicillin 3. Multiple cloning site (MCS) Unique restriction enzyme recognition sites Downstream of promoter sequence 4. Promoter sequence Upstream of the coding region Landing site for RNA polymerase so that the gene can be transcribed into m. RNA Common Vectors p. UC Copy Number+ ~500 -700 ORI Incompatibility Group Control p. MB 1 (derivative) A Relaxed p. BR 322 ~15 -20 p. MB 1 A Relaxed p. ET ~15 -20 p. BR 322 A Relaxed p. GEX ~15 -20 p. BR 322 A Relaxed p. Col. E 1 ~15 -20 Col. E 1 A Relaxed p. R 6 K ~15 -20 R 6 K B Stringent p. ACYC ~10 p 15 A B Relaxed p. SC 101 ~5 p. SC 101 C Stringent p. Bluescript ~300 -500 Col. E 1 (derivative) and F 1 A Relaxed p. GEM ~300 -500 p. UC and F 1 A Relaxed Source: http: //blog. addgene. org/plasmid-101 -origin-of-replication • • How to choose? Will plasmid be exclusively used in E. coli? Wil you have more than one plasmid? Do you need a high copy number? Will your gene product be toxic? Decision depends on copy number desired, host or hosts intended, compatibility. Plasmids from same group shouldn’t be co-transformed because they compete for the same machinery
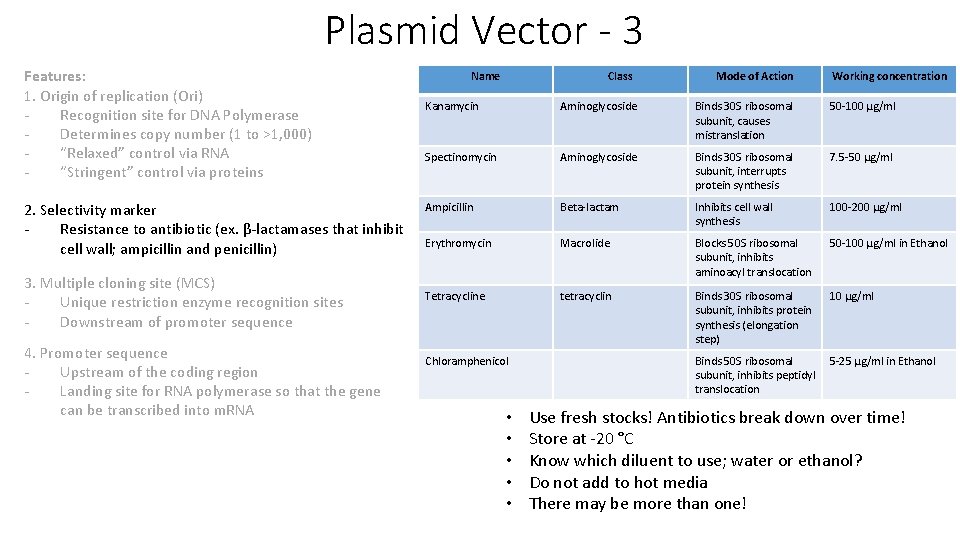
Plasmid Vector - 3 Features: 1. Origin of replication (Ori) Recognition site for DNA Polymerase Determines copy number (1 to >1, 000) “Relaxed” control via RNA “Stringent” control via proteins 2. Selectivity marker Resistance to antibiotic (ex. β-lactamases that inhibit cell wall; ampicillin and penicillin) 3. Multiple cloning site (MCS) Unique restriction enzyme recognition sites Downstream of promoter sequence 4. Promoter sequence Upstream of the coding region Landing site for RNA polymerase so that the gene can be transcribed into m. RNA Name Class Mode of Action Working concentration Kanamycin Aminoglycoside Binds 30 S ribosomal subunit, causes mistranslation 50 -100 μg/ml Spectinomycin Aminoglycoside Binds 30 S ribosomal subunit, interrupts protein synthesis 7. 5 -50 μg/ml Ampicillin Beta-lactam Inhibits cell wall synthesis 100 -200 μg/ml Erythromycin Macrolide Blocks 50 S ribosomal subunit, inhibits aminoacyl translocation 50 -100 μg/ml in Ethanol Tetracycline tetracyclin Binds 30 S ribosomal subunit, inhibits protein synthesis (elongation step) 10 μg/ml Chloramphenicol • • • Binds 50 S ribosomal 5 -25 μg/ml in Ethanol subunit, inhibits peptidyl translocation Use fresh stocks! Antibiotics break down over time! Store at -20 °C Know which diluent to use; water or ethanol? Do not add to hot media There may be more than one!
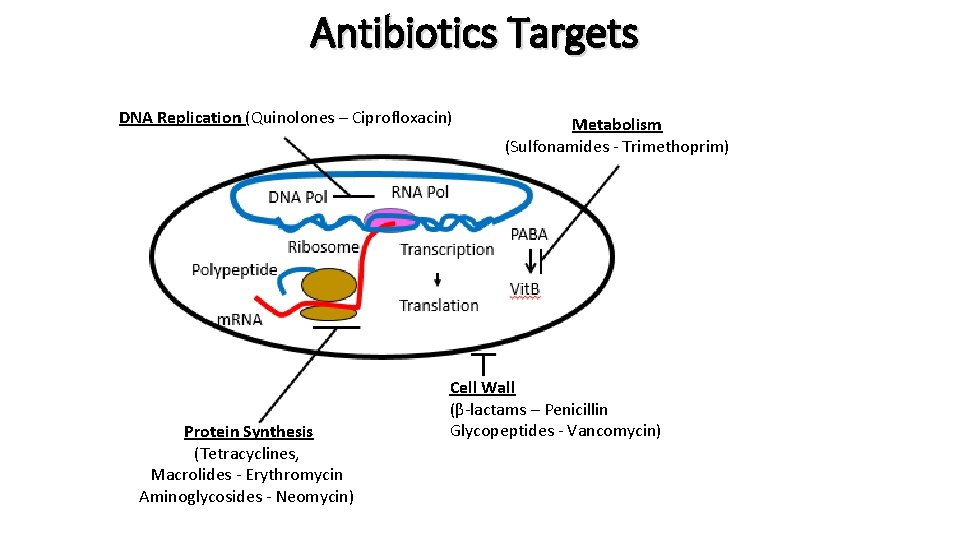
Antibiotics Targets DNA Replication (Quinolones – Ciprofloxacin) Protein Synthesis (Tetracyclines, Macrolides - Erythromycin Aminoglycosides - Neomycin) Metabolism (Sulfonamides - Trimethoprim) Cell Wall (β-lactams – Penicillin Glycopeptides - Vancomycin)
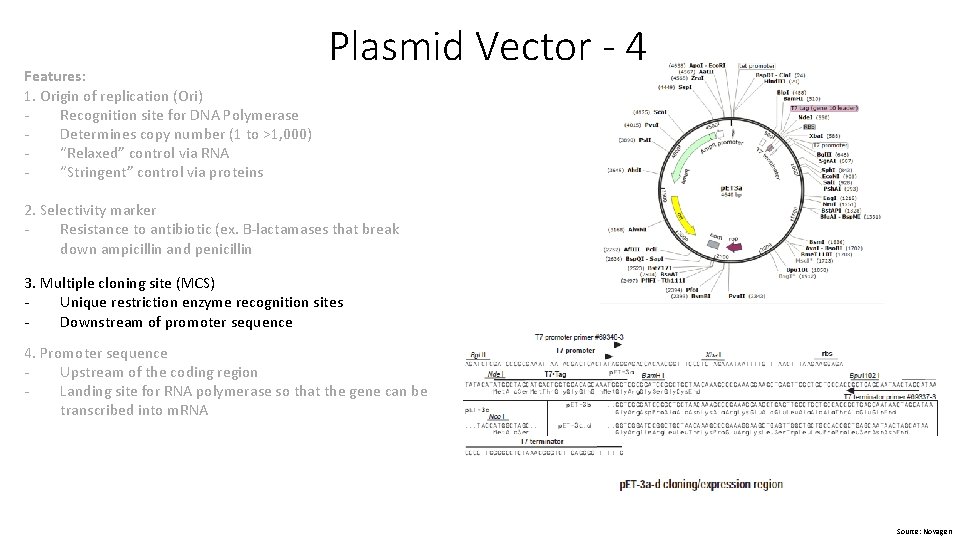
Features: 1. Origin of replication (Ori) Recognition site for DNA Polymerase Determines copy number (1 to >1, 000) “Relaxed” control via RNA “Stringent” control via proteins Plasmid Vector - 4 2. Selectivity marker Resistance to antibiotic (ex. B-lactamases that break down ampicillin and penicillin 3. Multiple cloning site (MCS) Unique restriction enzyme recognition sites Downstream of promoter sequence 4. Promoter sequence Upstream of the coding region Landing site for RNA polymerase so that the gene can be transcribed into m. RNA Source: Novagen
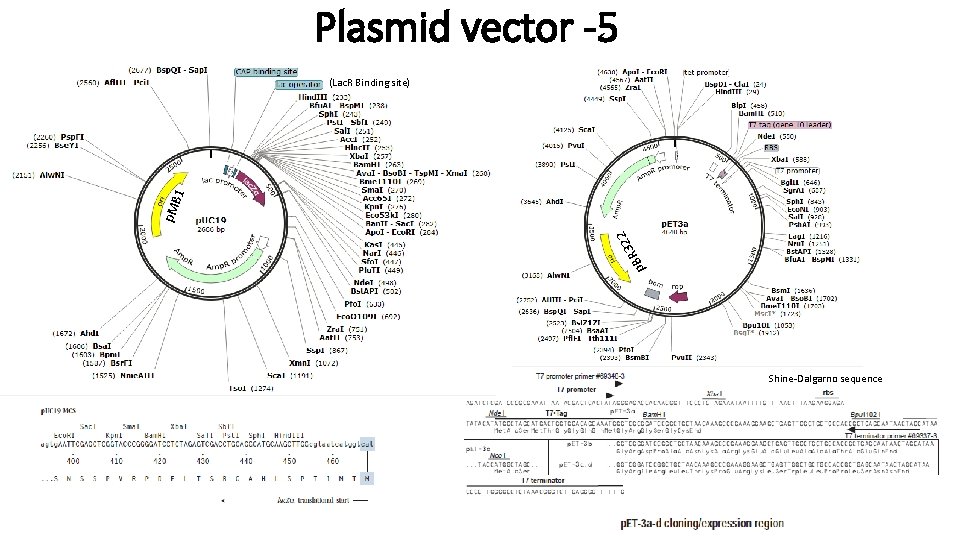
Plasmid vector -5 p 322 R B p. M B 1 (Lac. R Binding site) Shine-Dalgarno sequence

Plasmid vector - 6 Features: 1. Origin of replication (Ori) Recognition site for DNA Polymerase Determines copy number (1 to >1, 000) “Relaxed” control via RNA “Stringent” control via proteins 2. Selectivity marker Resistance to antibiotic (ex. B-lactamases that break down ampicillin and penicillin 3. Multiple cloning site (MCS) Unique restriction enzyme recognition sites Downstream of promoter sequence 4. Promoter sequence Upstream of the coding region Landing site for RNA polymerase so that the gene can be transcribed into m. RNA
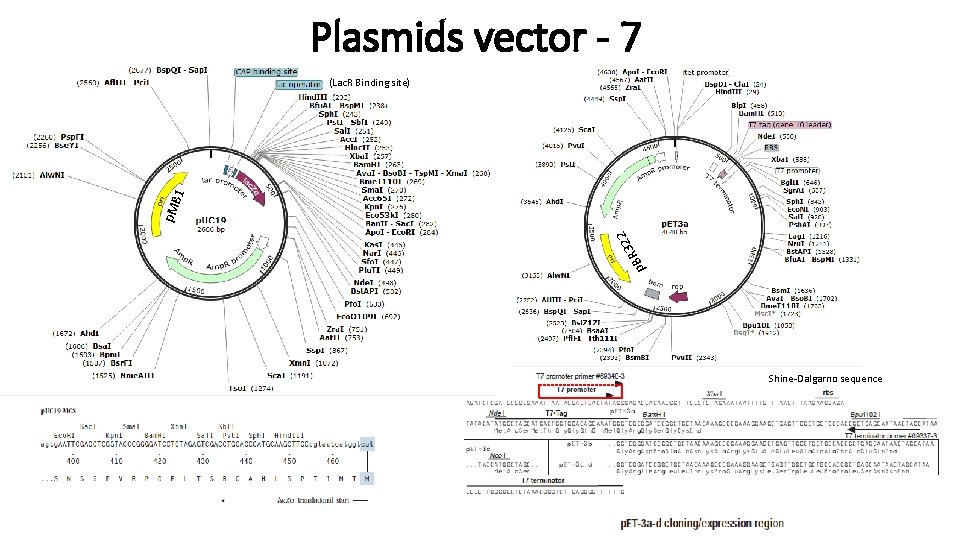
Plasmids vector - 7 p 322 R B p. M B 1 (Lac. R Binding site) Shine-Dalgarno sequence
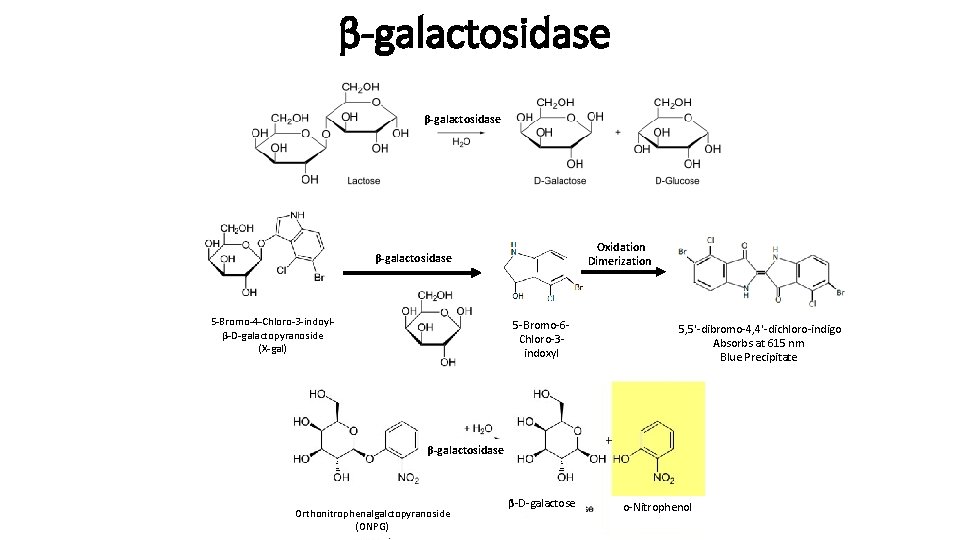
β-galactosidase Oxidation Dimerization β-galactosidase 5 -Bromo-4 -Chloro-3 -indoylβ-D-galactopyranoside (X-gal) 5 -Bromo-6 Chloro-3 indoxyl 5, 5'-dibromo-4, 4'-dichloro-indigo Absorbs at 615 nm Blue Precipitate β-galactosidase Orthonitrophenalgalctopyranoside (ONPG) β-D-galactose o-Nitrophenol
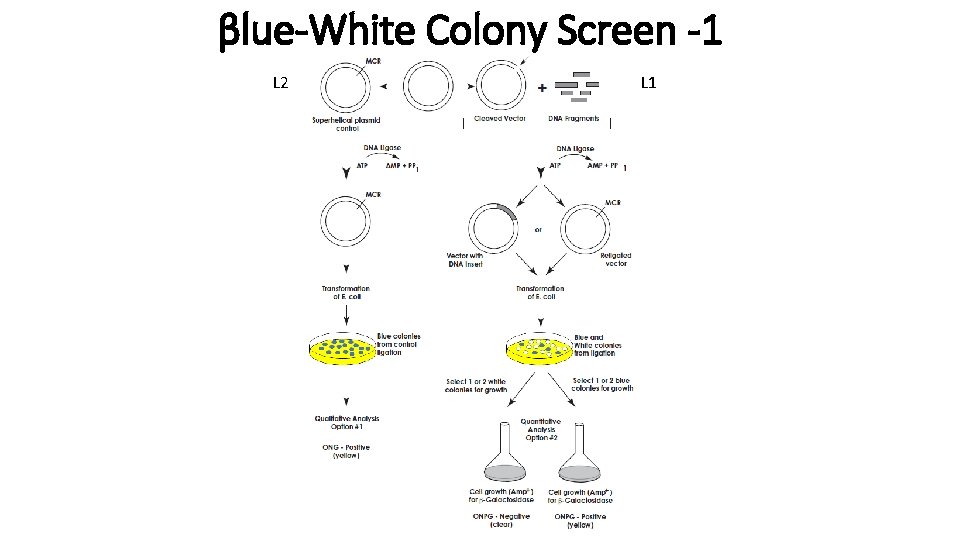
βlue-White Colony Screen -1 L 2 L 1
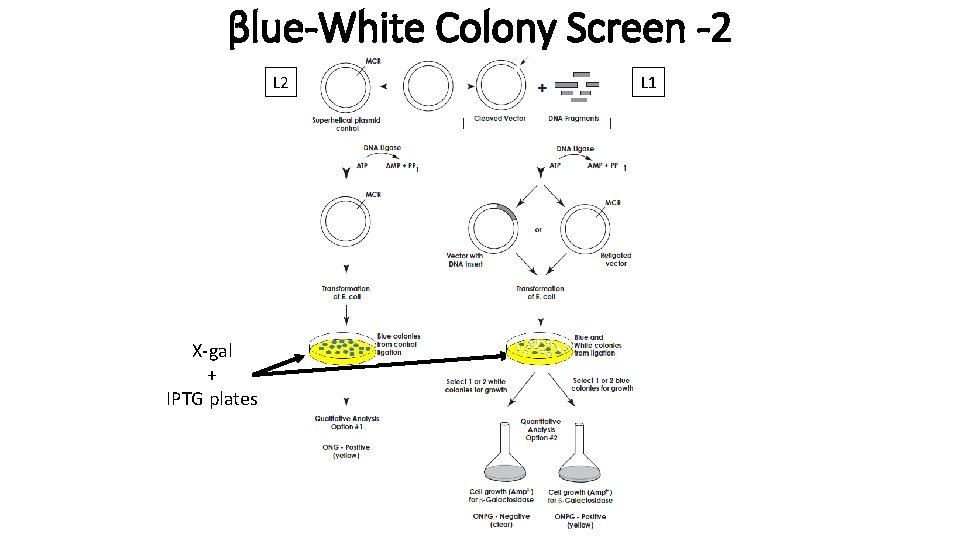
βlue-White Colony Screen -2 L 2 X-gal + IPTG plates L 1
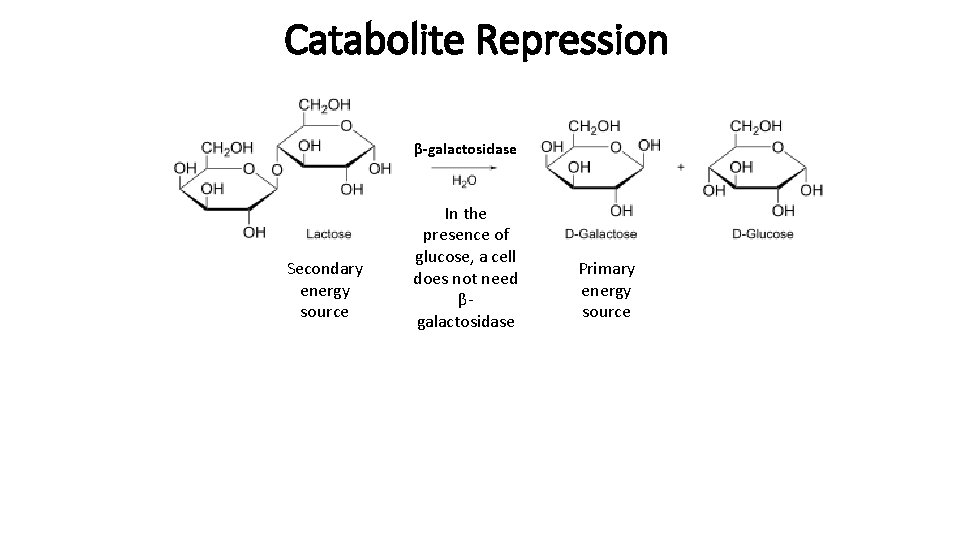
Catabolite Repression β-galactosidase Secondary energy source In the presence of glucose, a cell does not need βgalactosidase Primary energy source
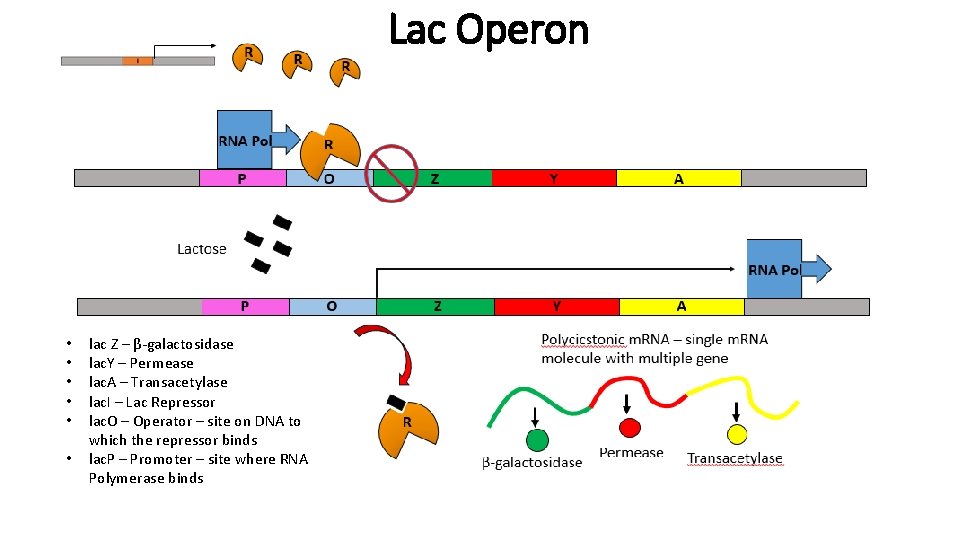
Lac Operon • • • lac Z – β-galactosidase lac. Y – Permease lac. A – Transacetylase lac. I – Lac Repressor lac. O – Operator – site on DNA to which the repressor binds lac. P – Promoter – site where RNA Polymerase binds
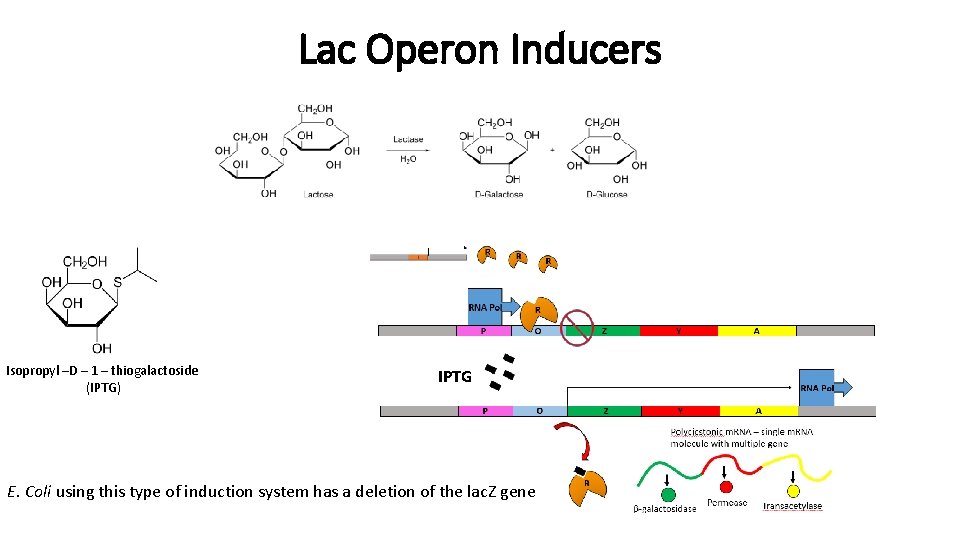
Lac Operon Inducers Isopropyl –D – 1 – thiogalactoside (IPTG) IPTG E. Coli using this type of induction system has a deletion of the lac. Z gene
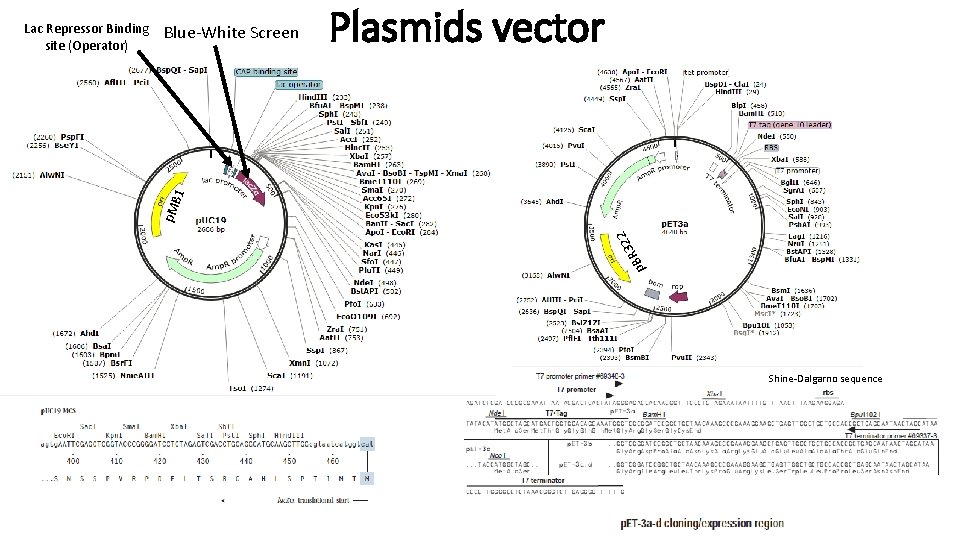
Blue-White Screen Plasmids vector p 322 R B p. M B 1 Lac Repressor Binding site (Operator) Shine-Dalgarno sequence

T 7 Promoter • • • Sequence from the T 7 bacteriophage T 7 Polymerase is under control of the lac promoter Expression is induced by the addition of IPTG Transcription of T 7 promoter is initiated by T 7 Pol T 7 DNA Polymerase necessary for expression of insert gene in plasmid How do we make T 7 Pol to express our protein? • Use strains with λ(DE 3) which means that these E. coli cells have prophage (Engineered phage) that has T 7 RNA polymerase controlled by Lac regulatory construct. Ex) BL 21 λDE 3 Which you have already used in lab!
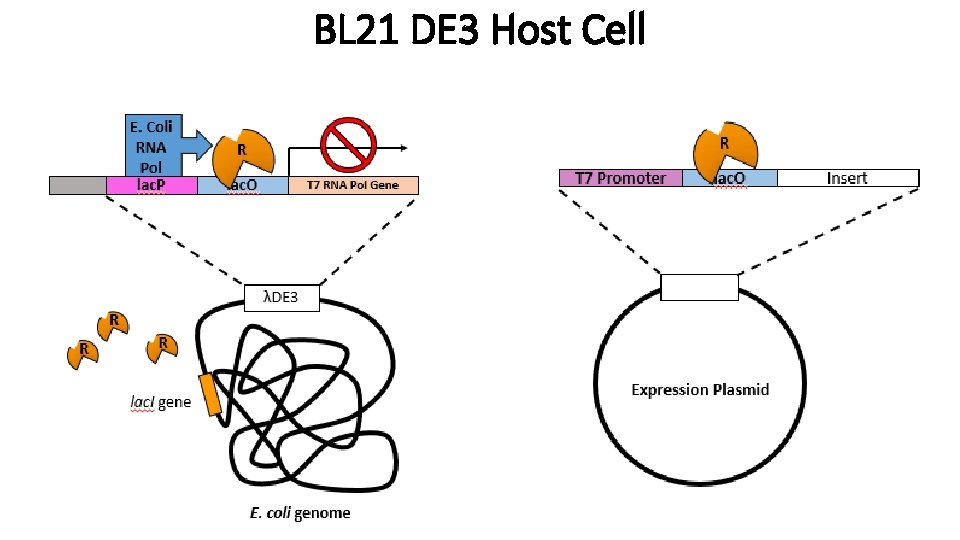
BL 21 DE 3 Host Cell
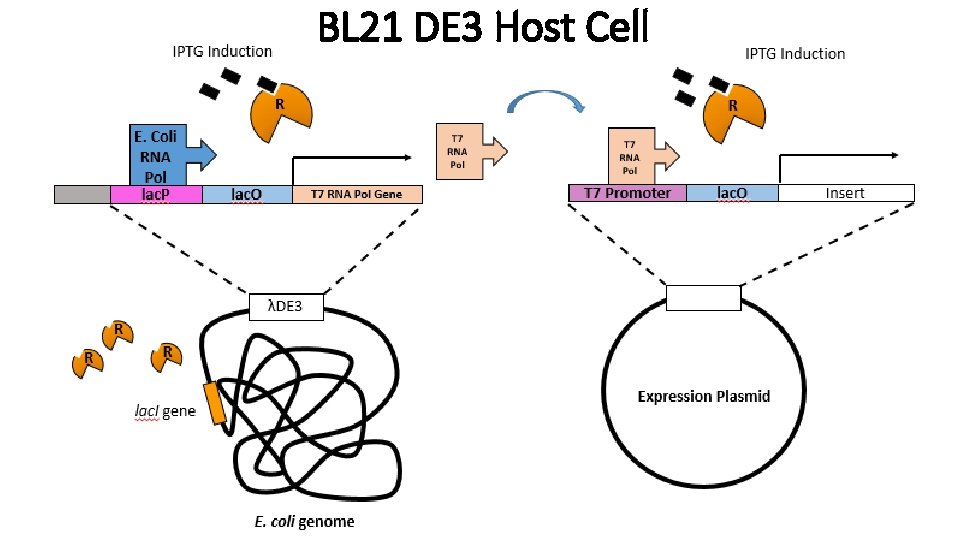
BL 21 DE 3 Host Cell
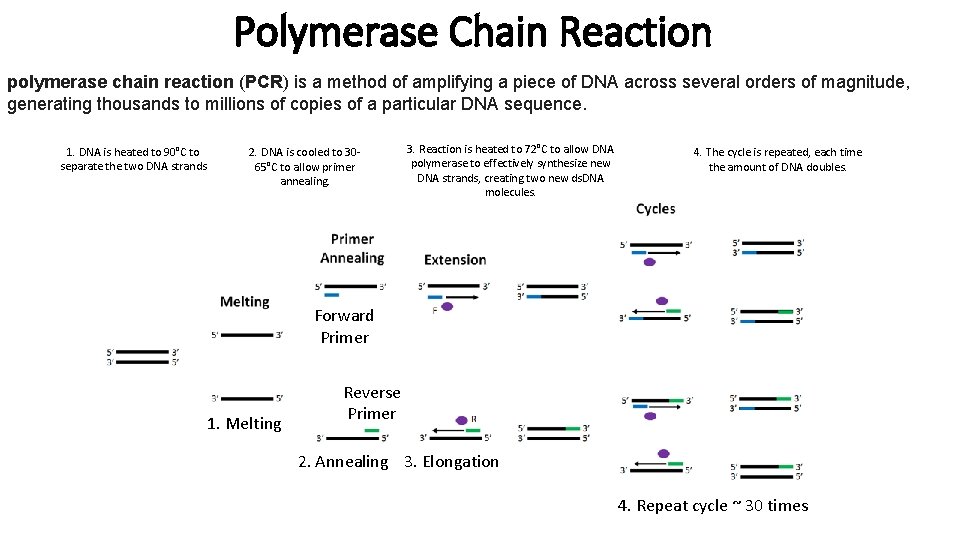
Polymerase Chain Reaction polymerase chain reaction (PCR) is a method of amplifying a piece of DNA across several orders of magnitude, generating thousands to millions of copies of a particular DNA sequence. 1. DNA is heated to 90°C to separate the two DNA strands 2. DNA is cooled to 3065°C to allow primer annealing. 3. Reaction is heated to 72°C to allow DNA polymerase to effectively synthesize new DNA strands, creating two new ds. DNA molecules. 4. The cycle is repeated, each time the amount of DNA doubles. Forward Primer 1. Melting Reverse Primer 2. Annealing 3. Elongation 4. Repeat cycle ~ 30 times
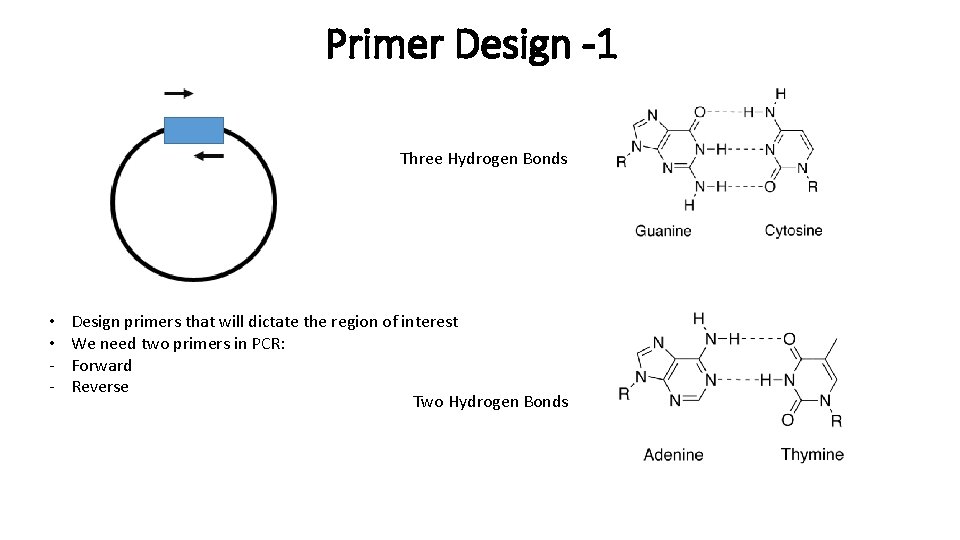
Primer Design -1 Three Hydrogen Bonds • • - Design primers that will dictate the region of interest We need two primers in PCR: Forward Reverse Two Hydrogen Bonds

Primer Design: G – C Content Factors to consider: • Primers are typically around 18 – 22 bp long • Tm – Melting Temperature • Tm between 55 – 65 °C typically works best • Tm of both primers should be within 5 °C of each other, the closer the better! • Annealing temperature (Ta) is generally 5 °C lower than Tm • Primers that have less G – C bonds will have a lower Tm than a similar length primer with more G – C bonds. • G – C content target is ~45 – 55% Three Hydrogen Bonds (Stronger bond) For sequences longer than 13 nucleotides: Tm= 64. 9 +41*(y. G+z. C-16. 4)/(w. A+x. T+y. G+z. C) Additional Factors to consider: • Primers price depends on size; 21 bp > 20 bp (but they’re cheap unless tags are included on the primer) • Primers larger than 60 bp become very expensive since synthesis becomes difficult and costs for validation increase Two Hydrogen Bonds (Weaker bond)

Primer Design - 2 5’ 3’ GCAATGCACTACGATTCGATCTCATGACAT C CGTTACGTGTATGCTAAGCTAGAGTACTGTA G 3’ 5’ DNA Sequence Complementary DNA sequence • DNA is read in the 5’ to 3’ direction (unless specified) • DNA Polymerase travels in the 3’ to the 5 ‘ direction on the template strand • New DNA strand is synthesized from 5’ to 3’
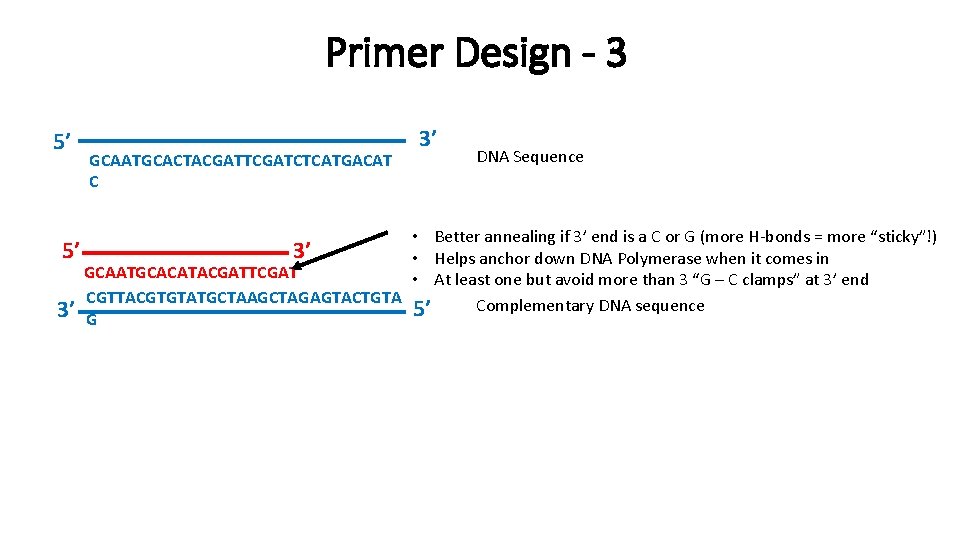
Primer Design - 3 5’ GCAATGCACTACGATTCGATCTCATGACAT C 3’ DNA Sequence • Better annealing if 3’ end is a C or G (more H-bonds = more “sticky”!) 5’ 3’ • Helps anchor down DNA Polymerase when it comes in GCAATGCACATACGATTCGAT • At least one but avoid more than 3 “G – C clamps” at 3’ end CGTTACGTGTATGCTAAGCTAGAGTACTGTA Complementary DNA sequence 5’ 3’ G
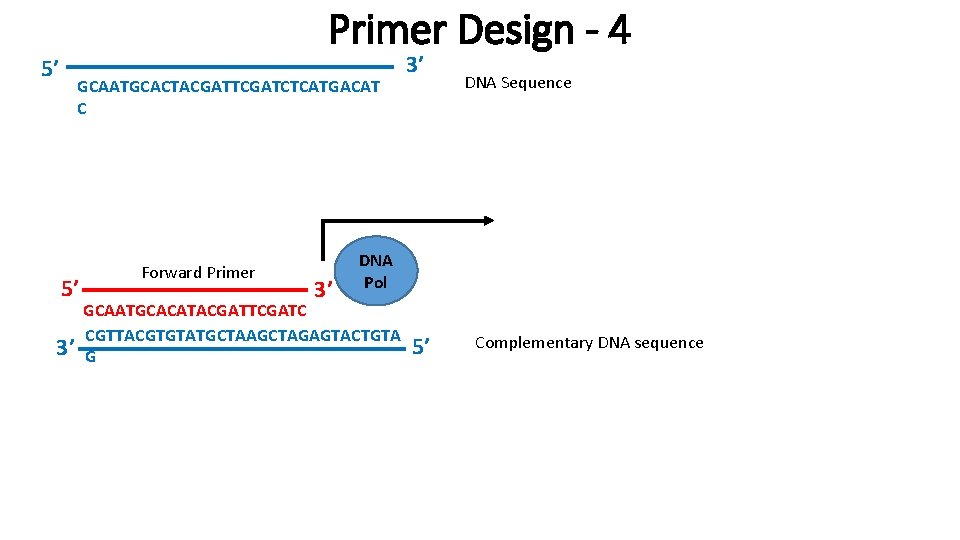
Primer Design - 4 5’ GCAATGCACTACGATTCGATCTCATGACAT C 5’ 3’ Forward Primer 3’ 3’ DNA Sequence DNA Pol GCAATGCACATACGATTCGATC CGTTACGTGTATGCTAAGCTAGAGTACTGTA G 5’ Complementary DNA sequence
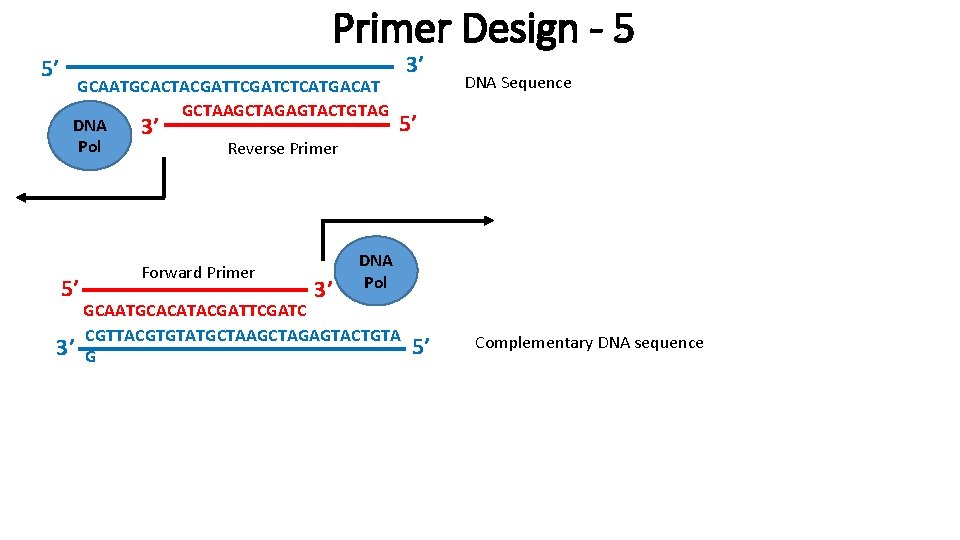
5’ Primer Design - 5 GCAATGCACTACGATTCGATCTCATGACAT C GCTAAGCTAGAGTACTGTAG DNA 3’ Pol Reverse Primer 5’ 3’ Forward Primer 3’ 3’ DNA Sequence 5’ DNA Pol GCAATGCACATACGATTCGATC CGTTACGTGTATGCTAAGCTAGAGTACTGTA G 5’ Complementary DNA sequence
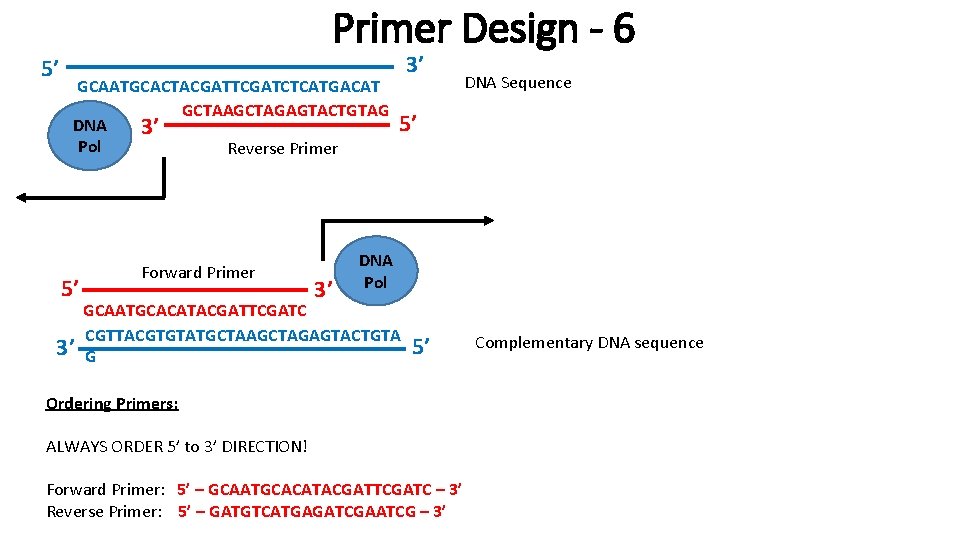
5’ Primer Design - 6 GCAATGCACTACGATTCGATCTCATGACAT C GCTAAGCTAGAGTACTGTAG DNA 3’ Pol Reverse Primer 5’ 3’ Forward Primer 3’ 3’ DNA Sequence 5’ DNA Pol GCAATGCACATACGATTCGATC CGTTACGTGTATGCTAAGCTAGAGTACTGTA G 5’ Ordering Primers: ALWAYS ORDER 5’ to 3’ DIRECTION! Forward Primer: 5’ – GCAATGCACATACGATTCGATC – 3’ Reverse Primer: 5’ – GATGTCATGAGATCGAATCG – 3’ Complementary DNA sequence
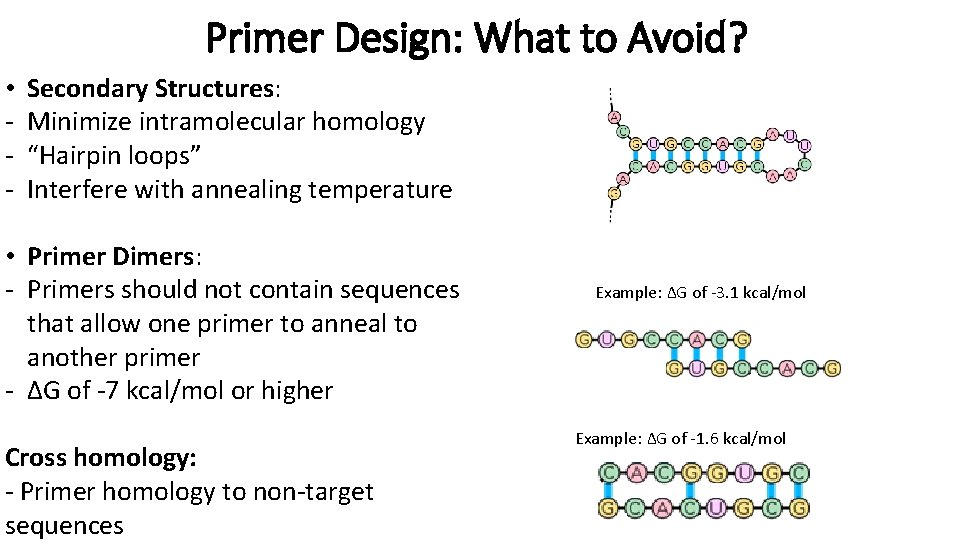
Primer Design: What to Avoid? • - Secondary Structures: Minimize intramolecular homology “Hairpin loops” Interfere with annealing temperature • Primer Dimers: - Primers should not contain sequences that allow one primer to anneal to another primer - ΔG of -7 kcal/mol or higher Cross homology: - Primer homology to non-target sequences Example: ΔG of -3. 1 kcal/mol Example: ΔG of -1. 6 kcal/mol
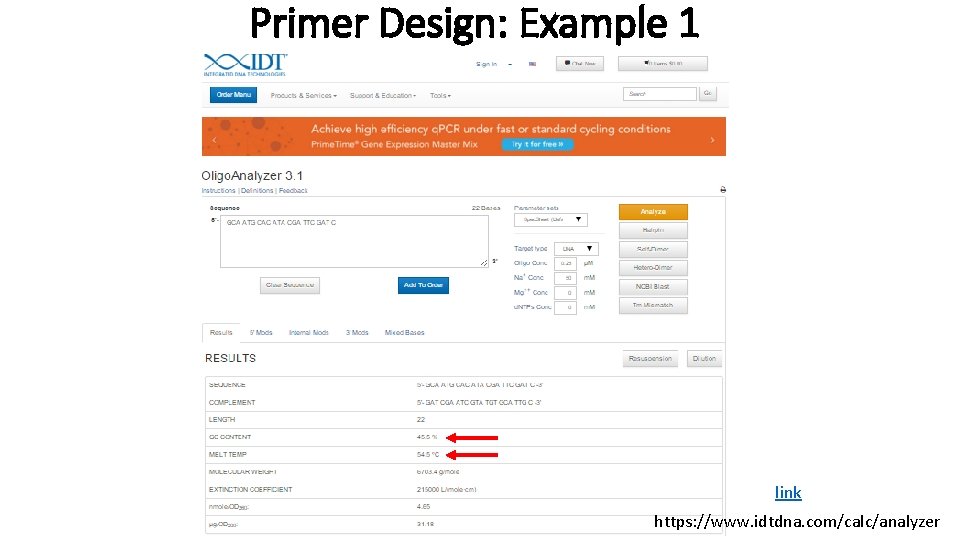
Primer Design: Example 1 link https: //www. idtdna. com/calc/analyzer
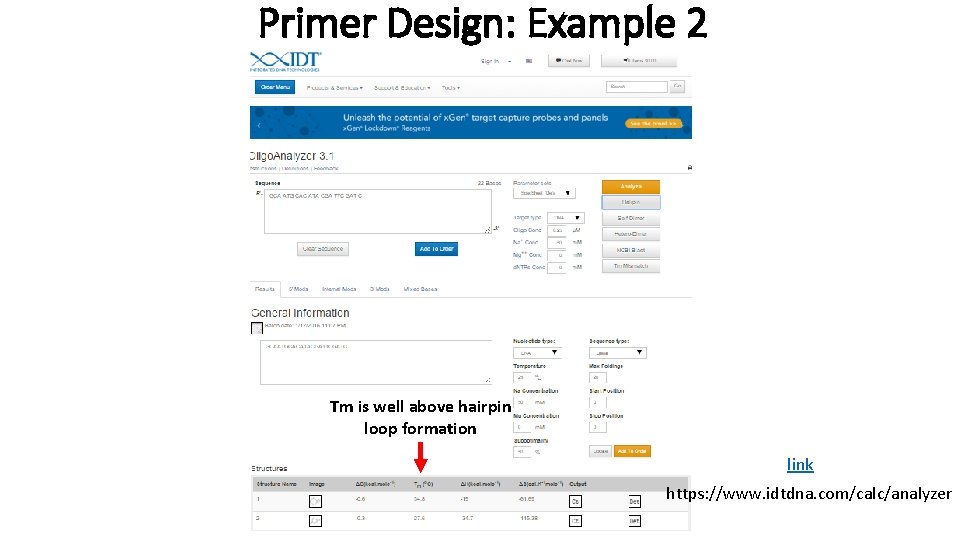
Primer Design: Example 2 Tm is well above hairpin loop formation link https: //www. idtdna. com/calc/analyzer
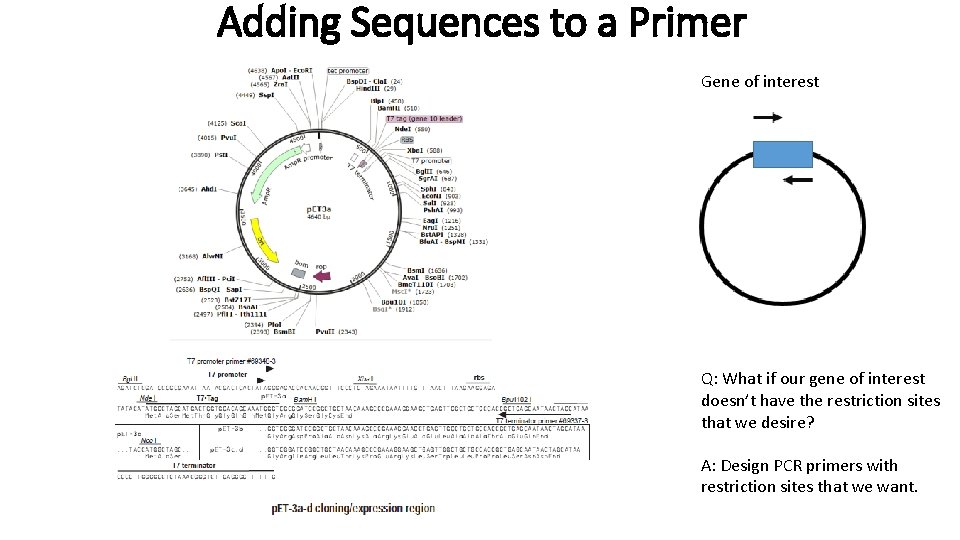
Adding Sequences to a Primer Gene of interest Q: What if our gene of interest doesn’t have the restriction sites that we desire? A: Design PCR primers with restriction sites that we want.
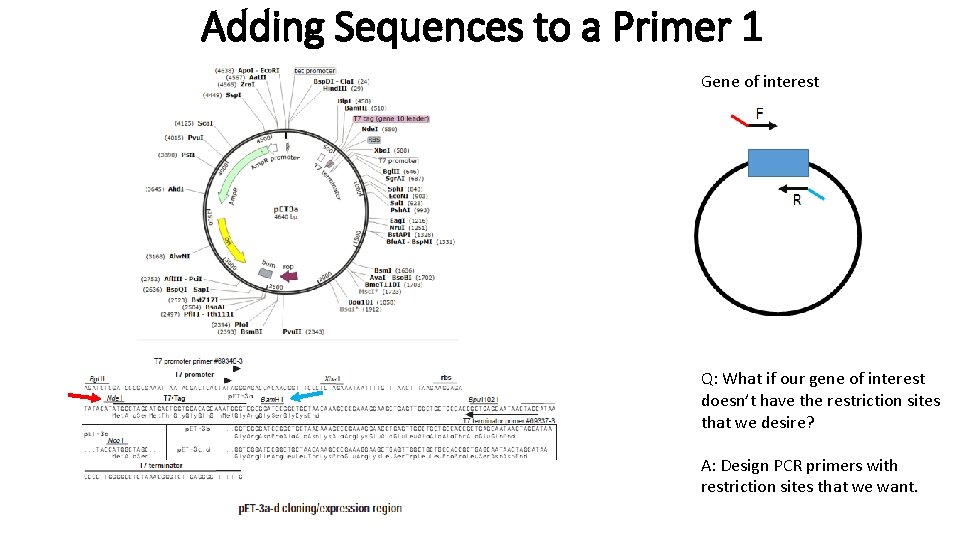
Adding Sequences to a Primer 1 Gene of interest Q: What if our gene of interest doesn’t have the restriction sites that we desire? A: Design PCR primers with restriction sites that we want.
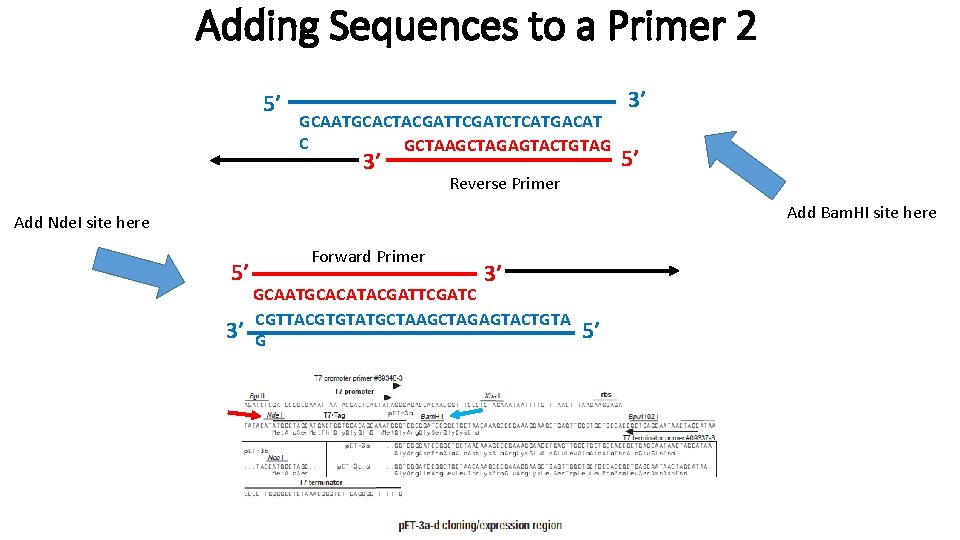
Adding Sequences to a Primer 2 5’ GCAATGCACTACGATTCGATCTCATGACAT C GCTAAGCTAGAGTACTGTAG 3’ Reverse Primer 3’ 5’ Add Bam. HI site here Add Nde. I site here 5’ 3’ Forward Primer 3’ GCAATGCACATACGATTCGATC CGTTACGTGTATGCTAAGCTAGAGTACTGTA G 5’
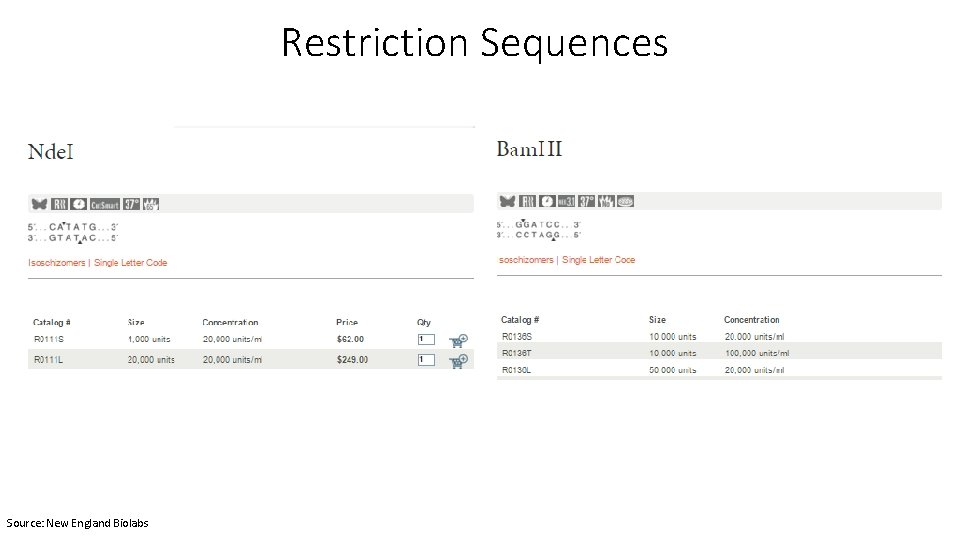
Restriction Sequences Source: New England Biolabs
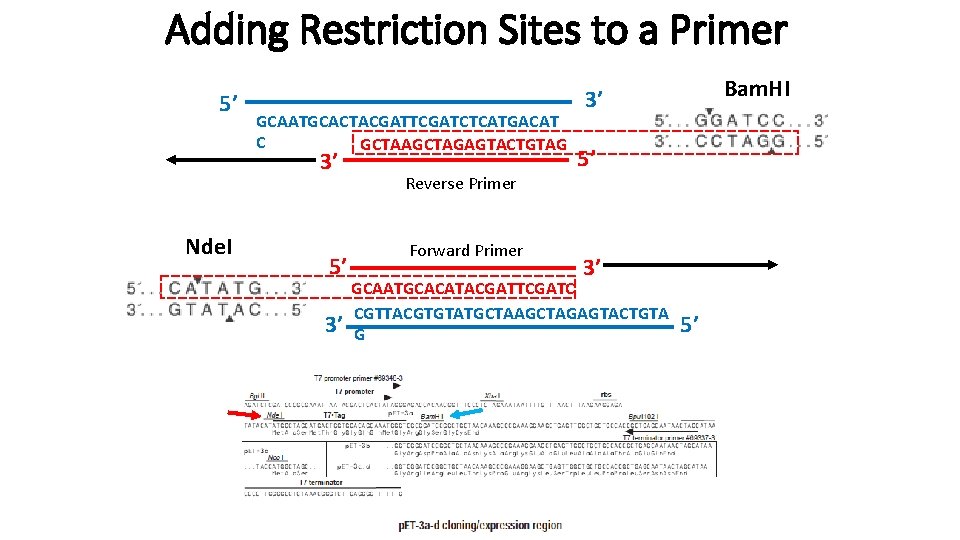
Adding Restriction Sites to a Primer 5’ GCAATGCACTACGATTCGATCTCATGACAT C GCTAAGCTAGAGTACTGTAG 3’ Nde. I 5’ 3’ Reverse Primer Forward Primer Bam. HI 3’ 5’ 3’ GCAATGCACATACGATTCGATC CGTTACGTGTATGCTAAGCTAGAGTACTGTA G 5’

Adding Restriction Sites to a Primer 3 5’ GCAATGCACTACGATTCGATCTCATGACAT C GCTAAGCTAGAGTACTGTAG 3’ Nde. I 5’ 3’ Reverse Primer Forward Primer Bam. HI 3’ 5’ 3’ GCAATGCACATACGATTCGATC CGTTACGTGTATGCTAAGCTAGAGTACTGTA G 5’ • Many restriction enzymes need additional nucleotides at the terminal ends of a DNA fragment to “sit” properly before cutting at their recognition site • The number of nucleotides necessary depends on the enzyme

Cleavage Close to DNA Fragment • May add on arbitrary sequence to the ends of your primers • Stay within optimal G-C content!
- Slides: 45