Lecture 36 Cloning and Sequencing Genes Lecture Outline









- Slides: 28

Lecture 36: Cloning and Sequencing Genes

Lecture Outline, 12/5/05 • Case Study: BRCA 1, continued – Cloning DNA fragments into plasmids • other vectors • “Libraries” of DNA – Di-deoxy Sequencing – Polymerase chain reaction (PCR)

Finding the Cancer Gene BRCA 1 • 1980’s: found several families that were predisposed to breast cancer • Studied 23 breast cancer families – Early onset – Frequent bilateral disease – Male relatives with breast cancer • 1990: linked the disease to a marker on Chromosome 17 q 21 – D 17 S 74 - 183 rd marker used! – Initial candidate region spanned half the chromosome (hundreds of possible genes. . . )

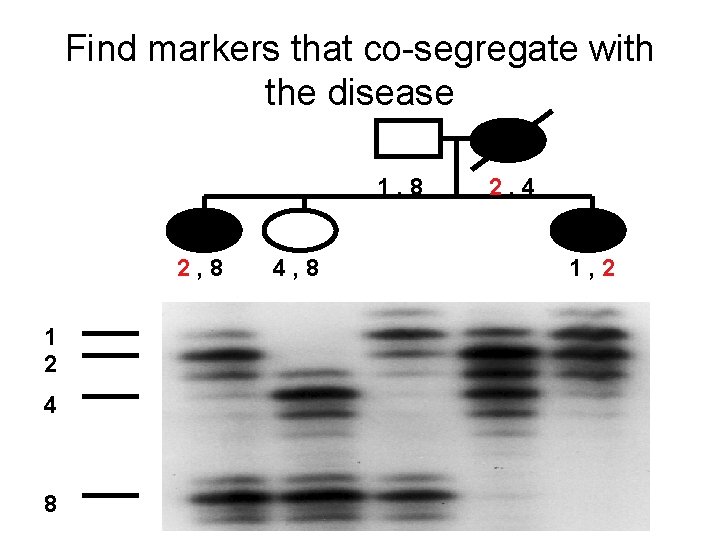
Find markers that co-segregate with the disease 1, 8 2, 8 1 2 4 8 4, 8 2, 4 1, 2

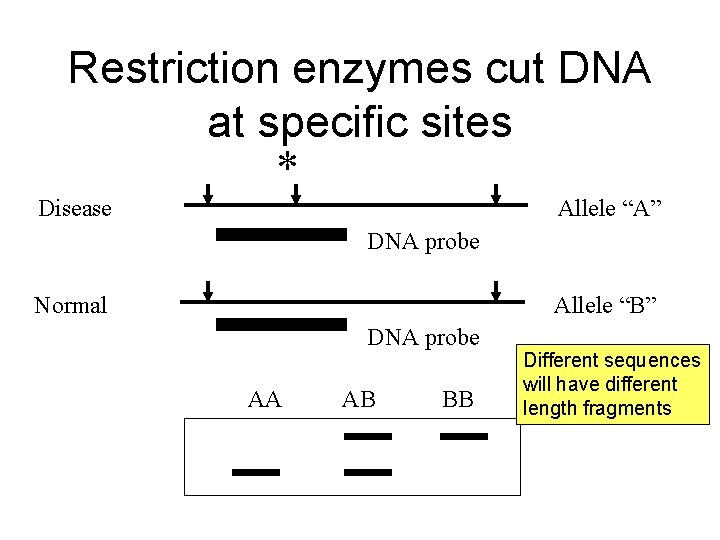
Restriction enzymes cut DNA at specific sites Disease * Allele “A” DNA probe Normal Allele “B” DNA probe AA AB BB Different sequences will have different length fragments

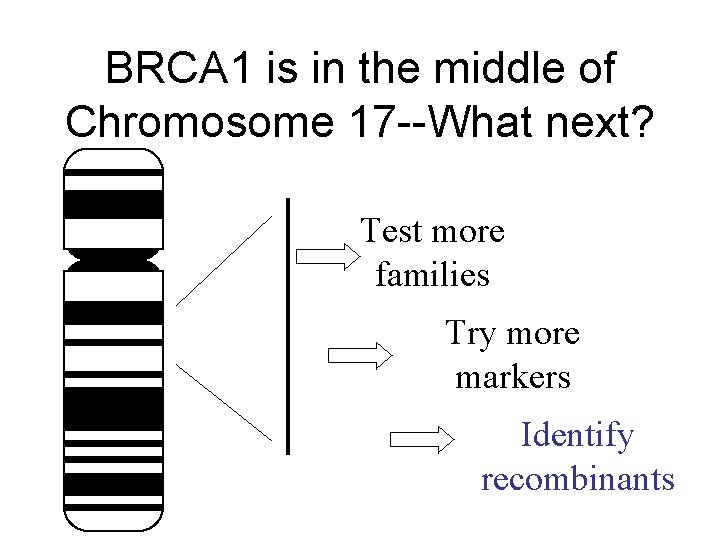
BRCA 1 is in the middle of Chromosome 17 --What next? Test more families Try more markers Identify recombinants

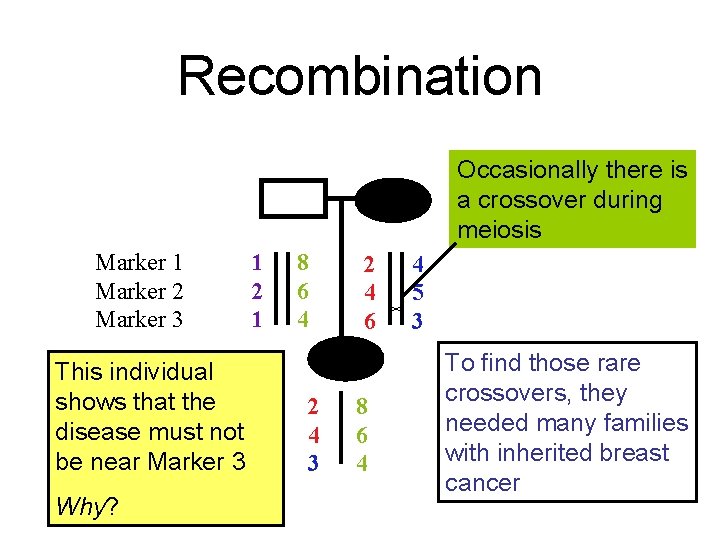
Recombination Occasionally there is a crossover during meiosis Marker 1 Marker 2 Marker 3 This individual shows that the disease must not be near Marker 3 Why? 1 2 1 8 6 4 2 4 3 2 4 6 8 6 4 4 5 3 To find those rare crossovers, they needed many families with inherited breast cancer

Mapping BRCA 1 • Larger study • 214 breast cancer families Chromosome 17 – Location narrowed to 8 c. M • But that was still a 600, 000 nucleotide region! • Step 2: Positional cloning to find the actual gene – – Make a “library” of cloned fragments Order those fragments Find fragments that contain coding sequences Sequence those fragments

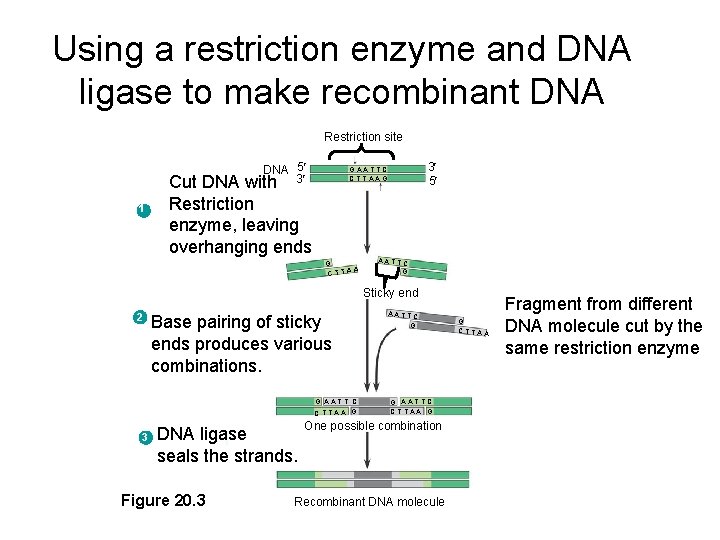
Using a restriction enzyme and DNA ligase to make recombinant DNA Restriction site DNA 5 with 3 1 3 5 GAATTC CTTAAG Cut DNA Restriction enzyme, leaving overhanging ends G CTTAA AATTC G Sticky end 2 Base pairing of sticky ends produces various combinations. G AATT C C TTAA G 3 DNA ligase seals the strands. Figure 20. 3 AATTC G G AATTC CTTAA G One possible combination Recombinant DNA molecule G CTTAA Fragment from different DNA molecule cut by the same restriction enzyme

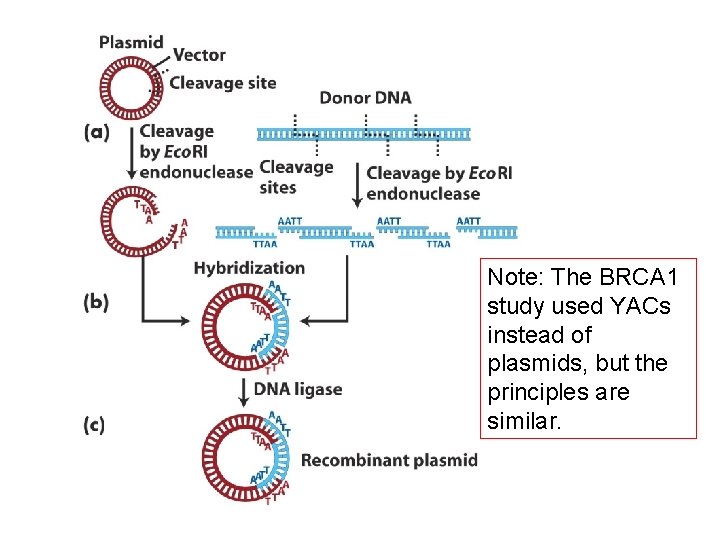
Note: The BRCA 1 study used YACs instead of plasmids, but the principles are similar.

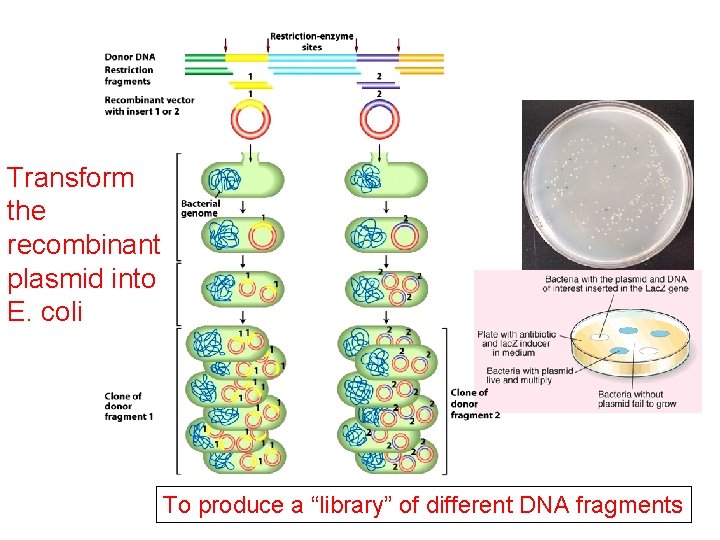
Transform the recombinant plasmid into E. coli To produce a “library” of different DNA fragments

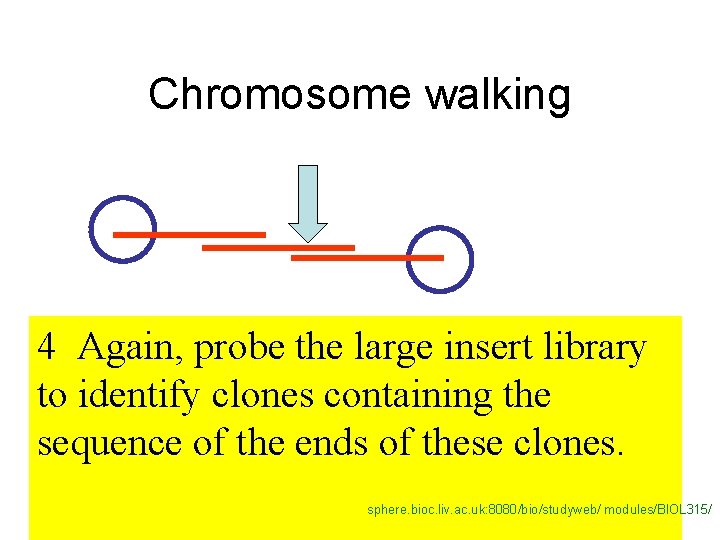
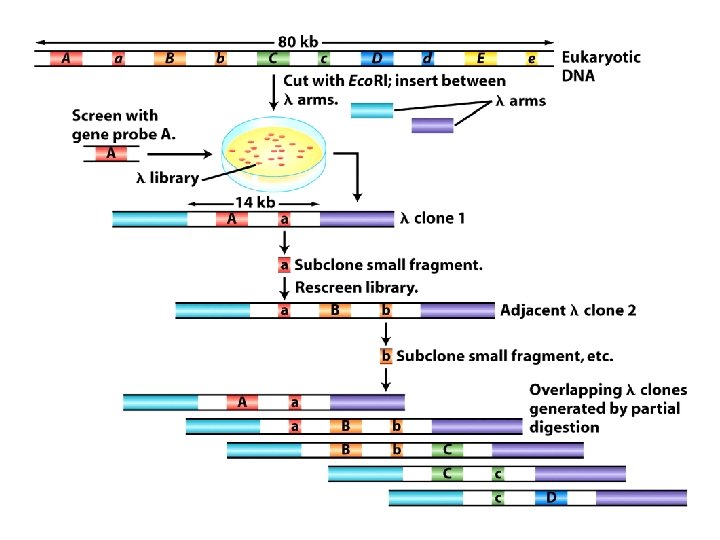
One of the clones in the library should contain the gene, but which one? 1. Probe a large insert library to identify a clone containing the marker linked to the trait. 1 b. Sequence the ends of that fragment. sphere. bioc. liv. ac. uk: 8080/bio/studyweb/ modules/BIOL 315/

Chromosome walking 2. Probe again to identify clones containing the end sequence of the first clone sphere. bioc. liv. ac. uk: 8080/bio/studyweb/ modules/BIOL 315/

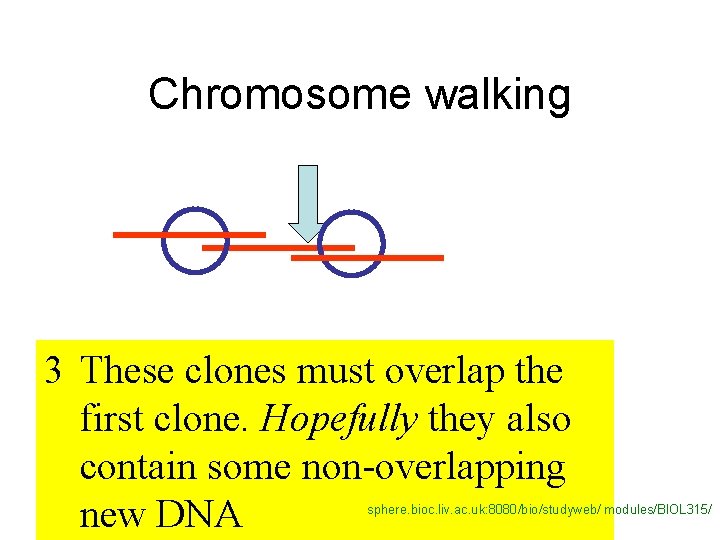
Chromosome walking 3 These clones must overlap the first clone. Hopefully they also contain some non-overlapping new DNA sphere. bioc. liv. ac. uk: 8080/bio/studyweb/ modules/BIOL 315/

Chromosome walking 4 Again, probe the large insert library to identify clones containing the sequence of the ends of these clones. sphere. bioc. liv. ac. uk: 8080/bio/studyweb/ modules/BIOL 315/

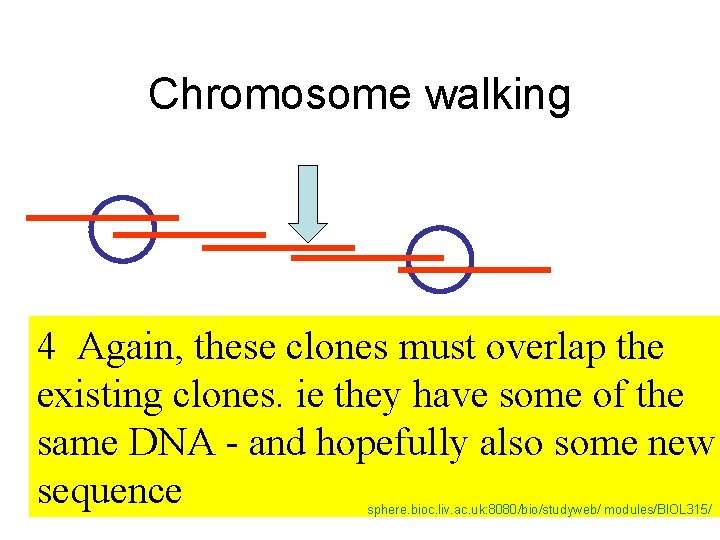
Chromosome walking 4 Again, these clones must overlap the existing clones. ie they have some of the same DNA - and hopefully also some new sequence sphere. bioc. liv. ac. uk: 8080/bio/studyweb/ modules/BIOL 315/

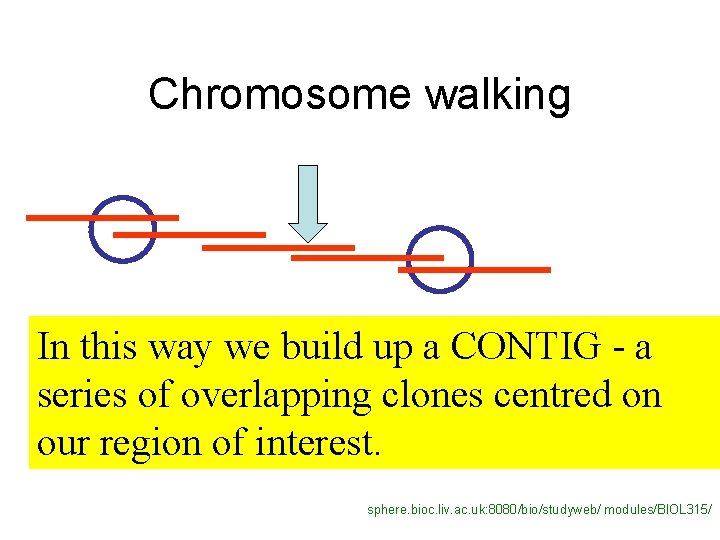
Chromosome walking In this way we build up a CONTIG - a series of overlapping clones centred on our region of interest. sphere. bioc. liv. ac. uk: 8080/bio/studyweb/ modules/BIOL 315/


• 8 cm may have many genes, but also lots of non-coding DNA • Kinds of DNA sequences: – Coding, SSR, pseudogenes, transposons – Limit sequencing only to coding sequences – All coding sequences make m. RNA

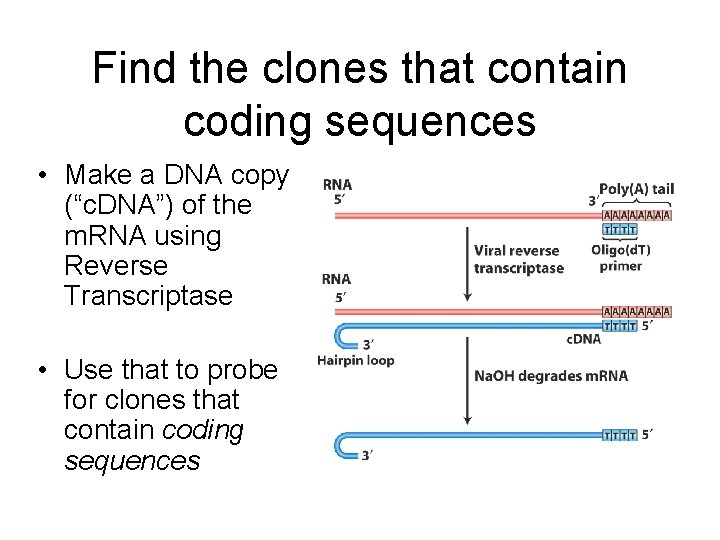
Find the clones that contain coding sequences • Make a DNA copy (“c. DNA”) of the m. RNA using Reverse Transcriptase • Use that to probe for clones that contain coding sequences

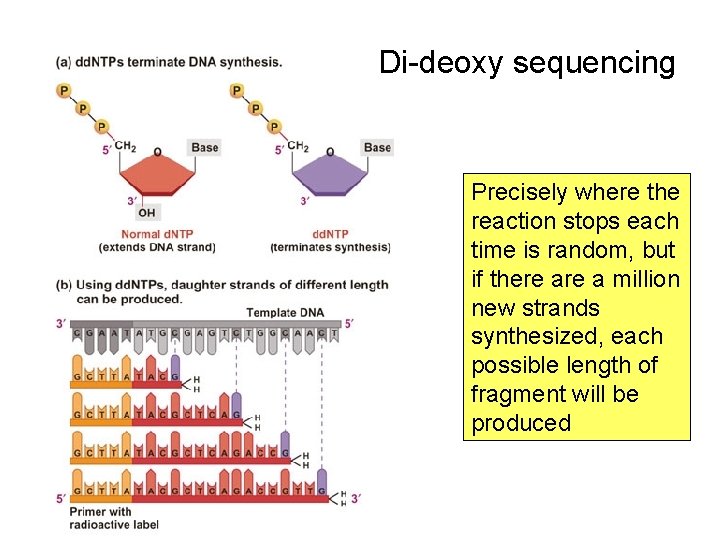
Determining the Nucleotide Sequence Ingredients to synthesize DNA in vitro: – Template DNA – DNA polymerase – A, C, G, T nucleotide triphosphates – Buffer (incl. salts and Mg. Cl) Then “poison” this + One more critical ingredient: Primer with 3’ OH recipe with small amounts of dideoxy nucleotides to stop the reaction

Di-deoxy sequencing Precisely where the reaction stops each time is random, but if there a million new strands synthesized, each possible length of fragment will be produced

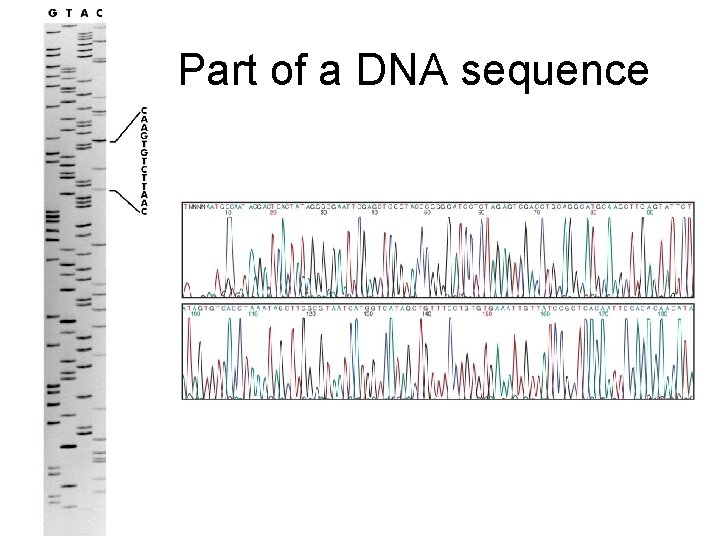
Part of a DNA sequence

BRCA 1 found in 1994 Science. 1994 Oct 7; 266(5182): 66 -71. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA 1. Miki Y, Swensen J, Shattuck-Eidens D, Futreal PA, Harshman K, Tavtigian S, Liu Q, Cochran C, Bennett LM, Ding W, et al. Department of Medical Informatics, University of Utah Medical Center, Salt Lake City 84132. A strong candidate for the 17 q-linked BRCA 1 gene, which influences susceptibility to breast and ovarian cancer, has been identified by positional cloning methods. Probable predisposing mutations have been detected in five of eight kindreds presumed to segregate BRCA 1 susceptibility alleles.

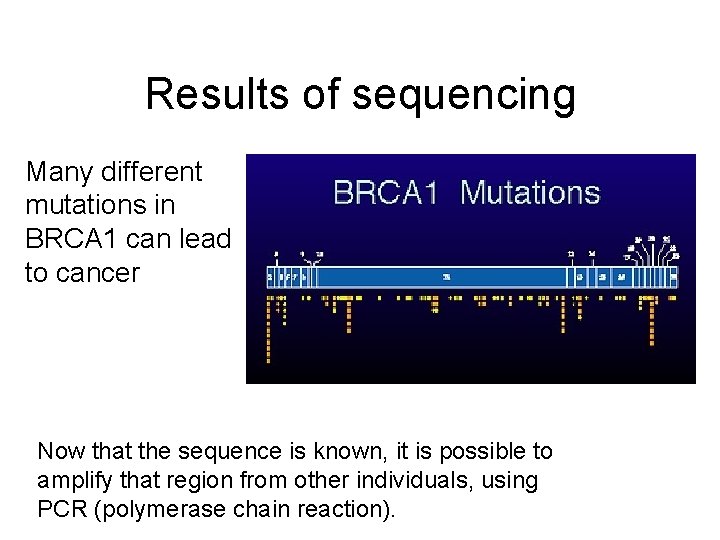
Results of sequencing Many different mutations in BRCA 1 can lead to cancer Now that the sequence is known, it is possible to amplify that region from other individuals, using PCR (polymerase chain reaction).

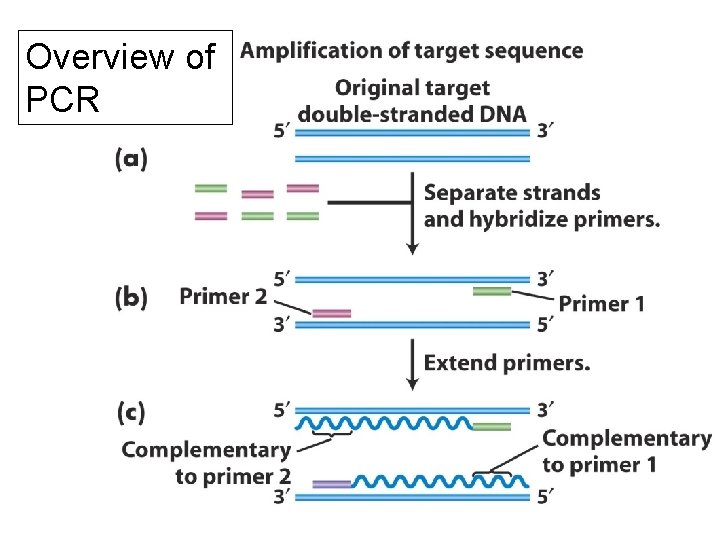
Overview of PCR

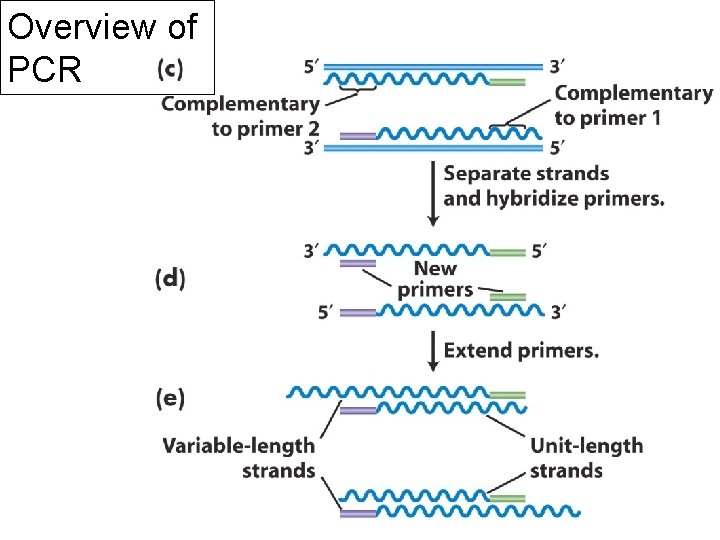
Overview of PCR

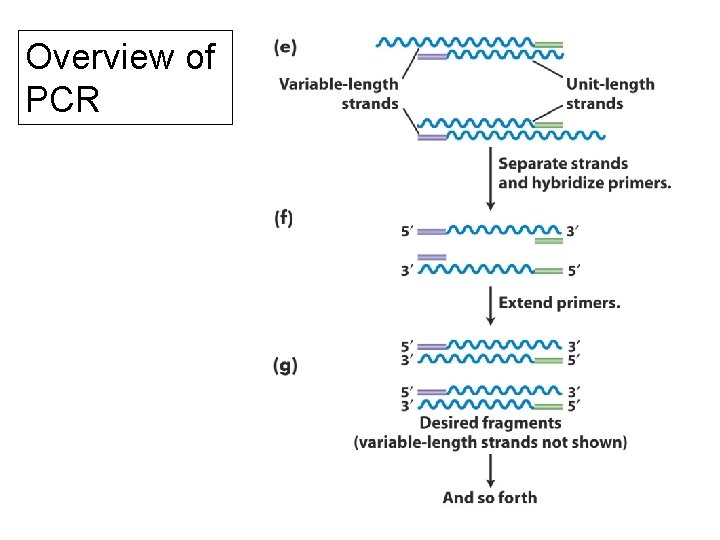
Overview of PCR