Lecture 3 January 6 2016 Biotech 3 Restriction

Lecture 3 January 6, 2016 Biotech 3

Restriction Enzyme Digest Discovery: • Late 1960 s • Werner Arber at the University of Basel, Switzerland - Showed that some E. coli strains restricted viral infection using an enzyme; called that enzyme a “restriction enzyme”. • Concurrently Hamilton Smith discovered Hind. II • Dan Nathans at Johns Hopkins University, Baltimore MD showed cleavage of Simian virus 40 using a restriction enzyme. • Received the Nobel Prize in Physiology or Medicine 1978

Restriction Enzymes as a Natural Defense • Bacteria use restriction enzymes as a defense mechanism to protect against infectious pathogens such as viruses called bacteriophage Bacterial DNA: Bacterial DNA is methylated CH 3 5’---CTAGAAGAATTCGGCAAGCT---3’ 3’---GATTCTTCTTAAGCCGTTCGA---5’ CH 3 • • Methyl groups will sterically hinder restriction sites, preventing their digestion by restriction enzymes Enzymes Dam and Dcm methylate DNA

Restriction Enzymes Cutting 5’…ACCTTGTCCCGGGTAGGCAT… 3’ 3’…TGGAACAGGGCCCATCCGTA… 5’ 5’…ACCTTGTCCC… 3’ 3’…TGGAACAGGG… 5’ 5’…GGGTAGGCAT… 3’ 3’…CCCATCCGTA… 5’ Blunt end cut 5’…ACCTTGTCTGCAGTAGGCAT… 3’ 3’…TGGAACAGACGTCATCCGTA… 5’ 5’…ACCTTGTCTGCA… 3’ 5’… GTAGGCAT… 3’ 3’…TGGAACAG… 5' 3’…ACGTCATCCGTA… 5’ Sticky end cut

Restriction Enzyme Examples Restriction Enzyme Organism Eco. RI Escherichia coli RY 13 Hind. III Haemphilus influenza Rd Pst. I Providencia stuartii Recognition sequence

Restriction Enzyme Digests • One unit is defined as the amount of enzyme required to digest 1 μg of λ DNA in 1 hour at 37°C in a total reaction volume of 50 μl • Supplied in “units” • There are various manufacturers and vendors of restriction enzymes • Popular manufacturer is New England Biolabs®, Inc. • The NEB web site has several tools that may be helpful in analyzing a DNA sequence for restriction sites and designing restriction enzyme digest.
NEB Web cutter NEB Web Cutter
NEB Web cutter NEB Web Cutter
NEB Web Cutter
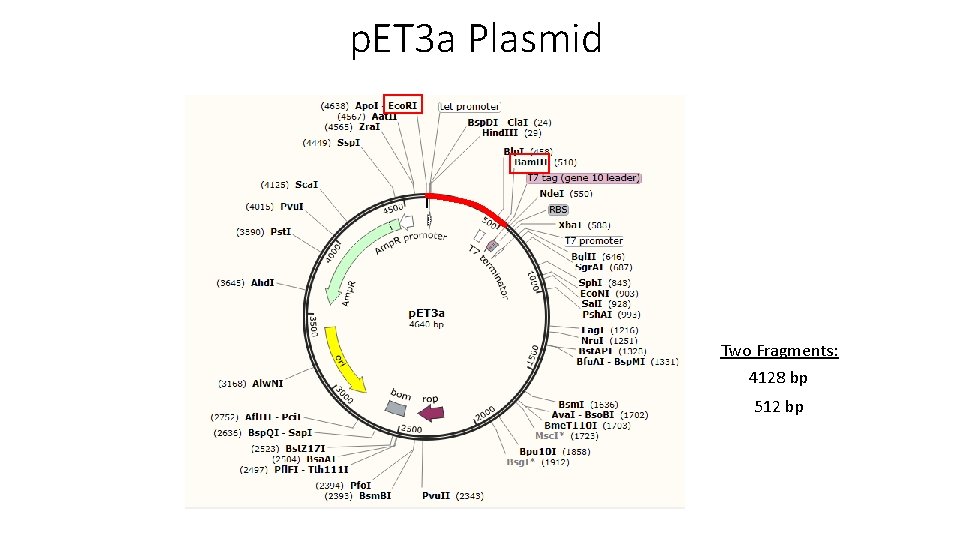
p. ET 3 a Plasmid Two Fragments: 4128 bp 512 bp
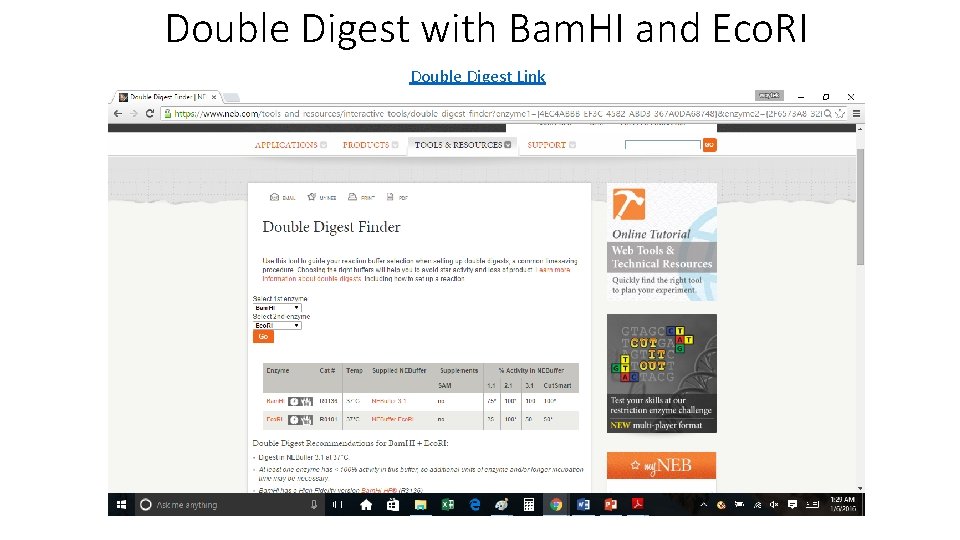
Double Digest with Bam. HI and Eco. RI Double Digest Link

Additional considerations Star Activity: Under non-standard reaction conditions, some restriction enzymes are capable of cleaving sequences which are similar, but not identical, to their defined recognition sequence. This altered specificity has been termed “star activity". Conditions that Contribute to Star Activity High glycerol concentration (> 5% v/v) High concentration of enzyme/µg of DNA ratio (varies with each enzyme, usually 100 units/µg) Non-optimal buffer Steps that can be Taken to Inhibit Star Activity Restriction enzymes are stored in 50% glycerol, therefore the amount of enzyme added should not exceed 10% of the total reaction volume. Use the standard 50 µl reaction volume to reduce evaporation during incubation. Use the fewest units possible to achieve digestion. This avoids overdigestion and reduces the final glycerol concentration in the reaction. Whenever possible, set up reactions in the recommended buffer. Buffers with differing ionic strength and p. H may contribute to star activity. Prolonged reaction time Use the minimum reaction time required for complete digestion. Prolonged incubation may result in increased star activity, as well as evaporation. Presence of organic solvents [DMSO, ethanol (4), ethylene glycol, dimethylacetamide, dimethylformamide, sulphalane Substitution of Mg 2+ with other divalent cations (Mn 2+, Cu 2+, Co 2+, Zn 2+) Make sure the reaction is free of any organic solvents, such as alcohols, which might be present in the DNA preparation. Use Mg 2+ as the divalent cation. Other divalent cations may not fit correctly into the active site of the restriction enzyme, possibly interfering with proper recognition.

Gel electrophoresis buffers
- Slides: 13