Lecture 3 Allele Frequencies and HardyWeinberg Equilibrium August

Lecture 3: Allele Frequencies and Hardy-Weinberg Equilibrium August 27, 2012

Last Time u. Review of genetic variation and Mendelian Genetics u. Methods for detecting variation Ø Morphology Ø Allozymes Ø DNA Markers ä Anonymous ä Sequence-tagged

Today u. Sequence probability calculation u. Molecular markers: DNA sequencing u. Introduction to statistical distributions u. Estimating allele frequencies u. Introduction to Hardy-Weinberg Equilibrium u. Using Hardy-Weinberg: Estimating allele frequencies for dominant loci

If nucleotides occur randomly in a genome, which sequence should occur more frequently? AGTTCAGAGTAACTGATGCT What is the expected probability of each sequence to occur once? How many times would each sequence be expected to occur by chance in a 100 Mb genome?

What is the expected probability of each sequence to occur once? AGTTCAGAGT What is the sample space for the first position? A T Probability of “A” at that position? G C Probability of “A” at position 1, “G” at position 2, “T” at position 3, etc. ? AGTTCAGAGTAACTGATGCT

How many times would each sequence be expected to occur in a 100 Mb genome? AGTTCAGAGTAACTGATGCT Why is this calculation wrong?
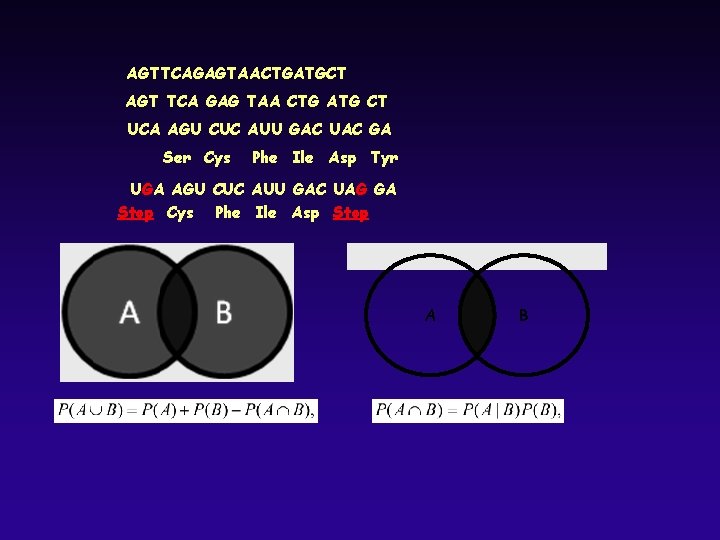
AGTTCAGAGTAACTGATGCT AGT TCA GAG TAA CTG ATG CT UCA AGU CUC AUU GAC UAC GA Ser Cys Phe Ile Asp Tyr UGA AGU CUC AUU GAC UAG GA Stop Cys Phe Ile Asp Stop A B
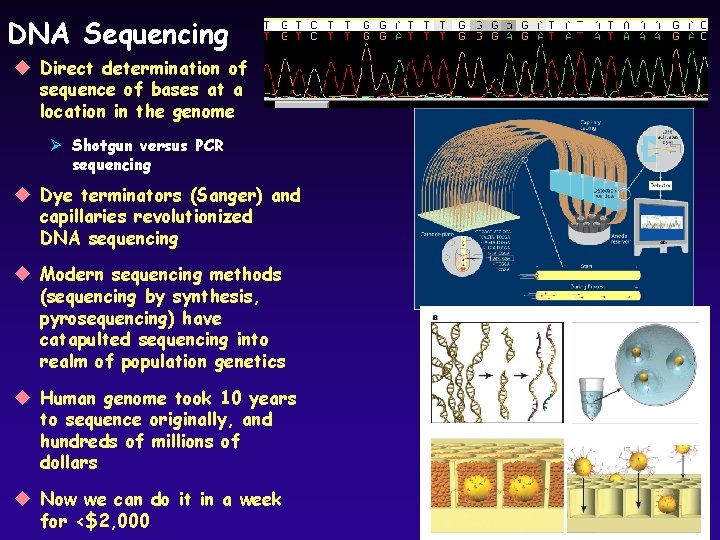
DNA Sequencing u Direct determination of sequence of bases at a location in the genome Ø Shotgun versus PCR sequencing u Dye terminators (Sanger) and capillaries revolutionized DNA sequencing u Modern sequencing methods (sequencing by synthesis, pyrosequencing) have catapulted sequencing into realm of population genetics u Human genome took 10 years to sequence originally, and hundreds of millions of dollars u Now we can do it in a week for <$2, 000
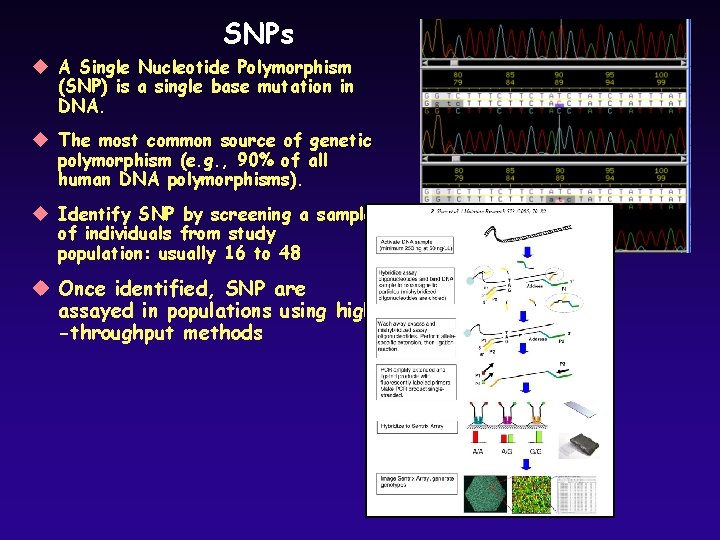
SNPs u A Single Nucleotide Polymorphism (SNP) is a single base mutation in DNA. u The most common source of genetic polymorphism (e. g. , 90% of all human DNA polymorphisms). u Identify SNP by screening a sample of individuals from study population: usually 16 to 48 u Once identified, SNP are assayed in populations using high -throughput methods
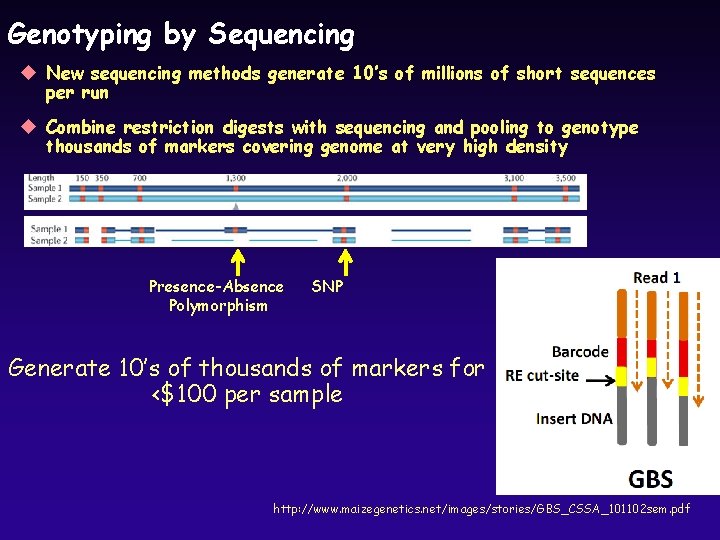
Genotyping by Sequencing u New sequencing methods generate 10’s of millions of short sequences per run u Combine restriction digests with sequencing and pooling to genotype thousands of markers covering genome at very high density Presence-Absence Polymorphism SNP Generate 10’s of thousands of markers for <$100 per sample http: //www. maizegenetics. net/images/stories/GBS_CSSA_101102 sem. pdf
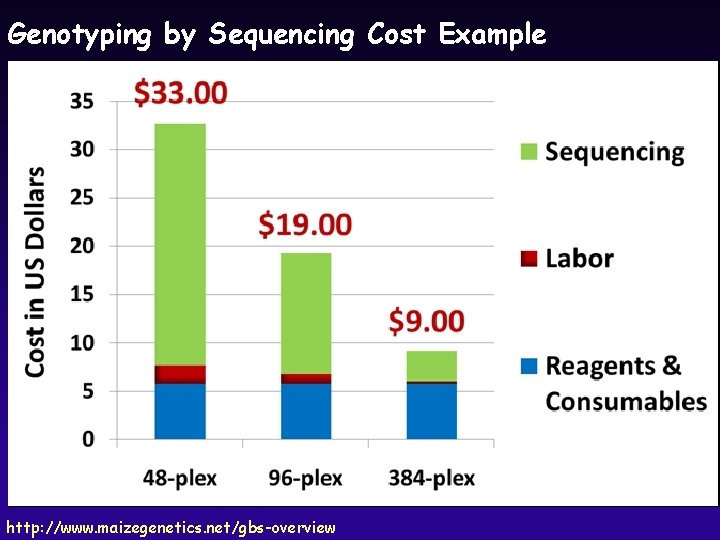
Genotyping by Sequencing Cost Example http: //www. maizegenetics. net/gbs-overview
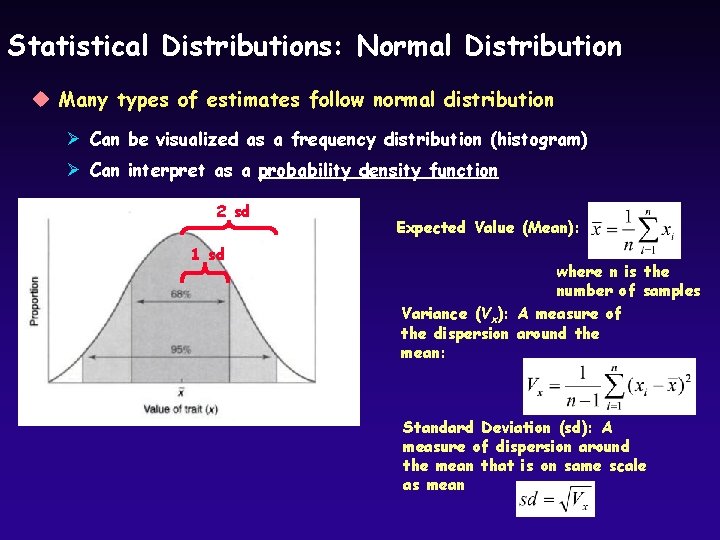
Statistical Distributions: Normal Distribution u Many types of estimates follow normal distribution Ø Can be visualized as a frequency distribution (histogram) Ø Can interpret as a probability density function 2 sd 1 sd Expected Value (Mean): where n is the number of samples Variance (Vx): A measure of the dispersion around the mean: Standard Deviation (sd): A measure of dispersion around the mean that is on same scale as mean

Standard Error of Mean u Standard Deviation is a measure of how individual points differ from the mean estimates in a single sample u Standard Error is a measure of how much the estimate differs from the true parameter value (in the case of means, μ) Ø If you repeated the experiment, how close would you expect the mean estimate to be to your previous estimate? Standard Error of the Mean (se): 95% Confidence Interval:

Estimating Allele Frequencies, Codominant Loci u Measured allele frequency is maximum likelihood estimator of the true frequency of the allele in the population (See Hedrick, pp 82 -83 for derivation) u Expected number of observations of allele A 1: E(Y)=np Ø Where n is number of samples Ø For diploid organisms, n = 2 N , where N is number of individuals sampled u Expected number of observations of allele A 1 is analogous to the mean of a sample from a normal distribution u Allele frequency can also be interpreted as an estimate of the mean

Allele Frequency Example u Assume a population of Mountain Laurel (Kalmia latifolia) at Cooper’s Rock, WV Red buds: 5000 Pink buds: 3000 White buds: 2000 A 1 A 1 A 1 A 2 A 2 u Phenotype is determined by a single, codominant locus: Anthocyanin u What is frequency of “red” alleles (A 1), and “white” alleles (A 2)? Frequency of A 1 = p Frequency of A 2 = q

Allele Frequencies are Distributed as Binomials u Based on samples from a population Ø For two-allele system, each sample is like a “trial” Ø Does the individual contain Allele A 1? Ø Remember, q=1 -p, so only one parameter is estimated u Binomials are variables that can be interpreted as the number of successes and failures in a series of trials where s is the probability of a success, and f is the probability of a failure Number of ways of observing y positive results in n trials Probability of observing y positive results in n trials once

Given the allele frequencies that you calculated earlier for Cooper’s Rock Kalmia latifolia, what is the probability of observing two “white” alleles in a sample of two plants?

Variation in Allele Frequencies, Codominant Loci u Binomial variance is pq or p(1 -p) u Variance in number of observations of A 1: V(Y) = np(1 -p) u Variance in allele frequency estimates (codominant, diploid): u Standard Error of allele frequency estimates: u Notice that estimates get better as sample size increases u Notice also that variance is maximum at intermediate allele frequencies
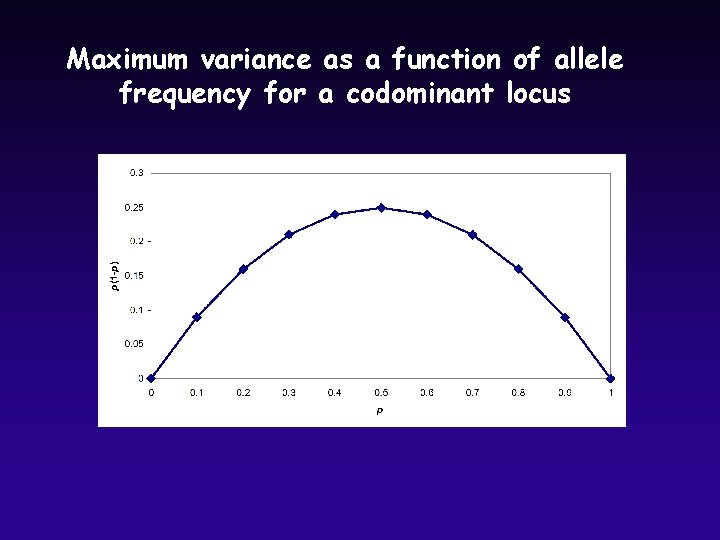
Maximum variance as a function of allele frequency for a codominant locus

Why is variance highest at intermediate allele frequencies? p = 0. 5 p = 0. 125 If this were a target, how variable would your outcome be in each case (red versus white hits)? Variance is constrained when value approaches limits (0 or 1)

What if there are more than 2 alleles? u General formula for calculating allele frequencies in multiallelic system with codominant alleles: u Variance and Standard Error of allele frequency estimates remain:

How do we estimate allele frequencies for dominant loci? Codominant locus - + A 1 A 1 A 1 A 2 A 2 A 2 Dominant locus A 1 A 1 A 1 A 2 A 2 A 2

Hardy-Weinberg Law u. After one generation of random mating, single-locus genotype frequencies can be represented by a binomial (with 2 alleles) or a multinomial function of allele frequencies Frequency of A 1 A 1 (P) Frequency of A 1 A 2 (H) Frequency of A 2 A 2 (Q)
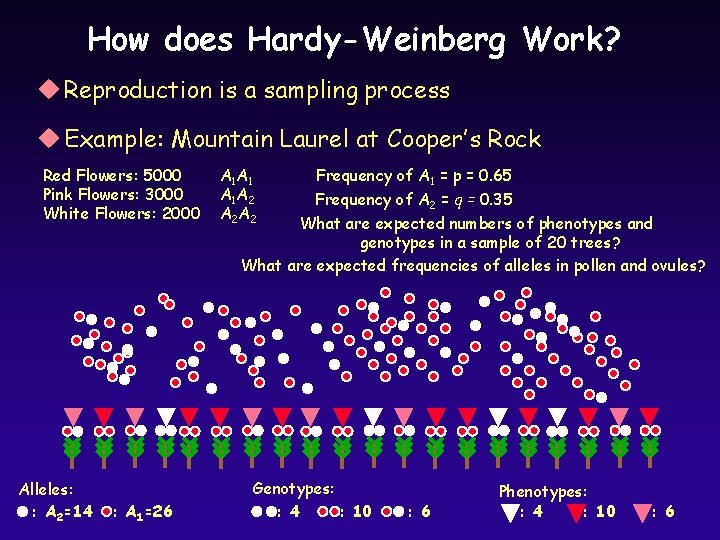
How does Hardy-Weinberg Work? u Reproduction is a sampling process u Example: Mountain Laurel at Cooper’s Rock Red Flowers: 5000 Pink Flowers: 3000 White Flowers: 2000 Alleles: : A 2=14 : A 1=26 A 1 A 1 A 1 A 2 A 2 Frequency of A 1 = p = 0. 65 Frequency of A 2 = q = 0. 35 What are expected numbers of phenotypes and genotypes in a sample of 20 trees? What are expected frequencies of alleles in pollen and ovules? Genotypes: : 4 : 10 : 6 Phenotypes: : 4 : 10 : 6

What will be the genotype and phenotype frequencies in the next generation? What assumptions must we make?

Hardy-Weinberg Equilibrium u. After one generation of random mating, genotype frequencies remain constant, as long as allele frequencies remain constant u. Provides a convenient Neutral Model to test for departures from assumptions u. Allows genotype frequencies to be represented by allele frequencies: simplification of calculations

Hardy-Weinberg Assumptions u. Diploid u. Large population u. Random Mating: equal probability of mating among genotypes u. No mutation u. No gene flow u. Equal allele frequencies between sexes u. Nonoverlapping generations
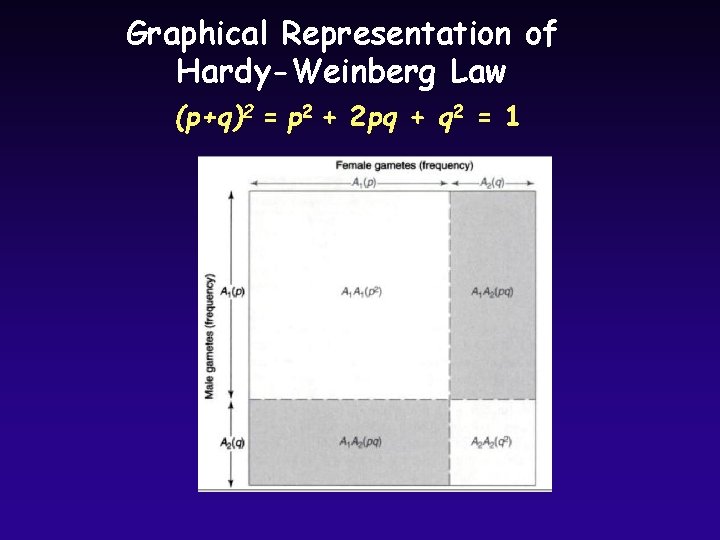
Graphical Representation of Hardy-Weinberg Law (p+q)2 = p 2 + 2 pq + q 2 = 1
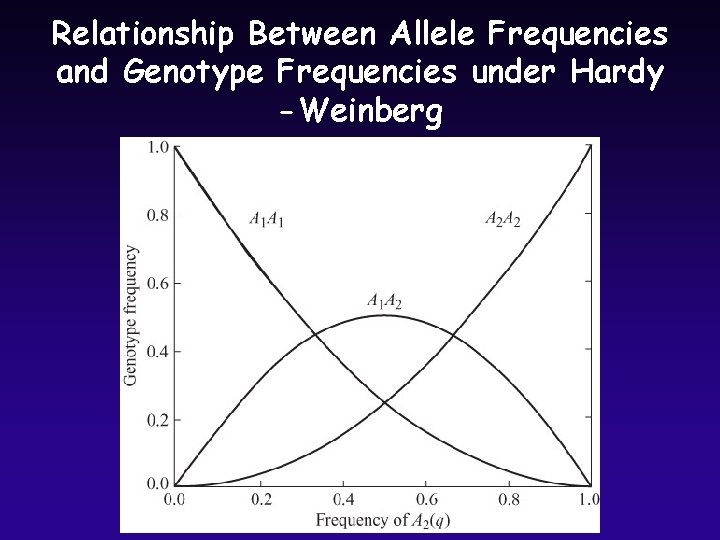
Relationship Between Allele Frequencies and Genotype Frequencies under Hardy -Weinberg
- Slides: 29