Lecture 26 Advanced Association Genetics December 3 2012

Lecture 26: Advanced Association Genetics December 3, 2012

Announcements § Extra credit lab this Wednesday: up to 10 points § Extra credit report due at final exam § Review session on Friday, Dec. 7 § Final exam on Monday, Dec. 10 at 11 am in computer lab ØNOT on Dec. 11 like syllabus and lecture notes say!

Last Time § Association genetics § Effects of population structure § Transmission Disequilibrium Tests

Today § Limitations of association genetics approaches § Solutions: ØImputation of genotypes ØMultiple testing corrections ØGenomic selection § The Case of the Missing Heritability
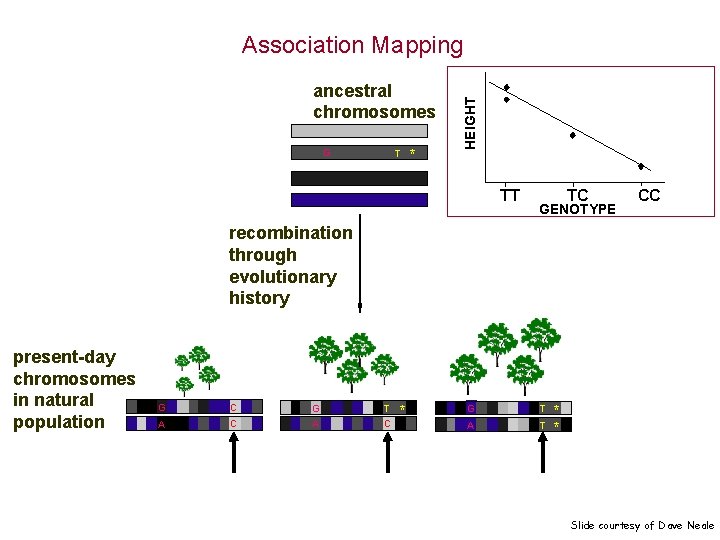
ancestral chromosomes G T * HEIGHT Association Mapping TT TC GENOTYPE CC recombination through evolutionary history present-day chromosomes in natural population G C A C G A T C * G T A T * * Slide courtesy of Dave Neale

Association Study Limitations § Population structure: differences between cases and controls § Genetic heterogeneity underlying trait § Inadequate genome coverage/Missing Genotypes § Random error/false positives § Multiple testing

Missing Genotypes §Potential source of bias in analysis Ø Some alleles under-represented Ø Problem if data gathered differently in case and control populations §Missing genotypes degrade power of analysis §More complex statistical models required §Solution: Imputation

Imputing Missing Genotypes From Isik and Wetten 2011 Workshop on Genomic Selection Typically accomplished with software such as IMPUTE, PLINK, MACH, BEAGLE, and fast. PHASE

Detecting Associations: Single SNP Tests §Contingency tests Ø Chi-square Armitage Test Ø Fisher’s Exact Test §Armitage test fits a line to relationship between genotype score (number of alleles) and “genotypic risk” §Null hypothesis: slope=0 §�Assumes additivity §Genomic control (GC): threshold of significance set by background SNPs: inflate critical value by a constant Balding 2006

Genome-Wide Association Studies and Multiple Testing § With Next-Gen sequencing, true genome-wide association studies are a reality § Millions of tests of association § How to set proper P-value cutoff? Ø With P=0. 05, expect 50, 000 type I errors per million tests § Need protection from type I error Null
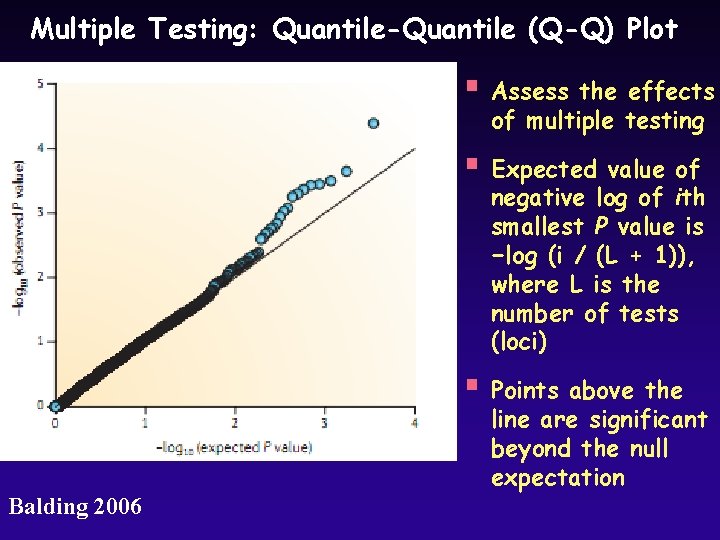
Multiple Testing: Quantile-Quantile (Q-Q) Plot § Assess the effects of multiple testing § Expected value of negative log of ith smallest P value is −log (i / (L + 1)), where L is the number of tests (loci) § Points above the line are significant beyond the null expectation Balding 2006

Corrections for Multiple Testing § Bonferoni: Where N is number of tests § Very conservative § Alternative: False Discovery Rate or Benjamani. Hochberg test Where i is the number of P-values that are less than or equal to the current P. Test is performed with smallest P first, in sorted order § P-values can also be set by permutation: randomize the phenotype data across genotypes, generate a distribution
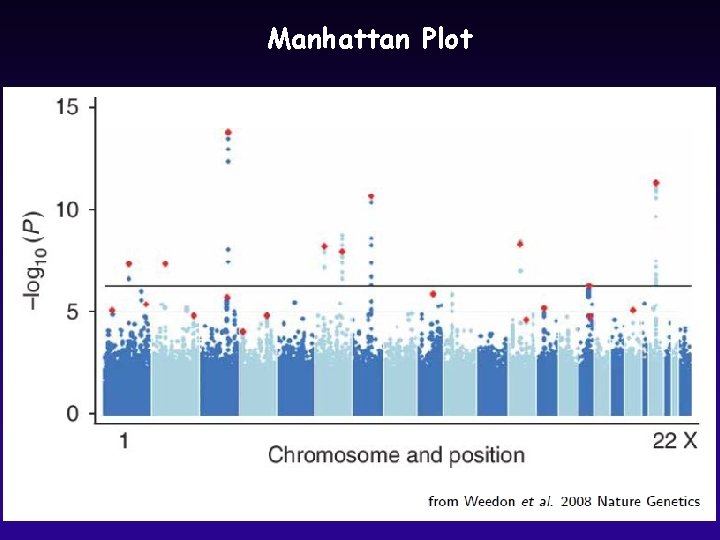
Manhattan Plot

How Successful have GWAS Been? §Thousands of associations have been identified for many different traits §Each locus explains a very small proportion of the variation in complex traits (typically <1%) §Overall percentage of variation explained is substantially less than trait heritability, even for case-control diseases: “Missing heritability” Manolio et al. 2009. Nature 461: 747– 753.

Possible Causes of Missing Heritability u Much larger numbers of common variants of smaller effect yet to be found u Gene-environment interaction u Trait heterogeneity u Rare variants (possibly with larger effects) u De novo mutations u Structural variations such as copy number variants u Gene–gene interactions, epistasis u Beyond DNA sequence: epigenetic markers 15
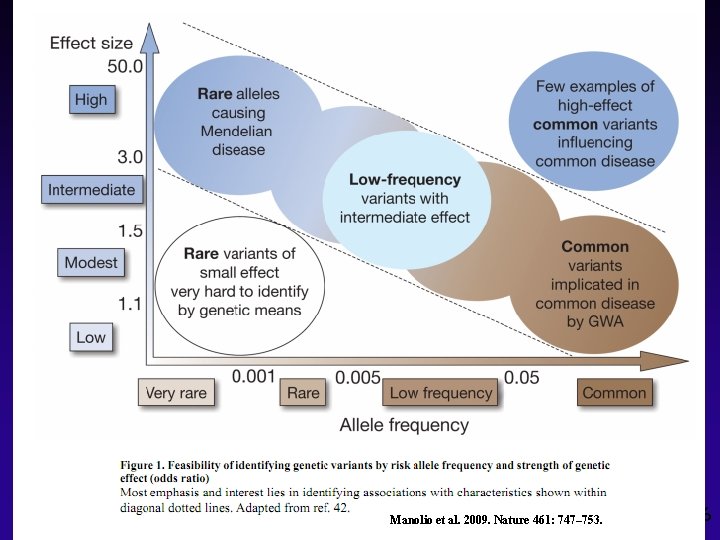
Possible Causes of Missing Heritability Manolio et al. 2009. Nature 461: 747– 753. 16
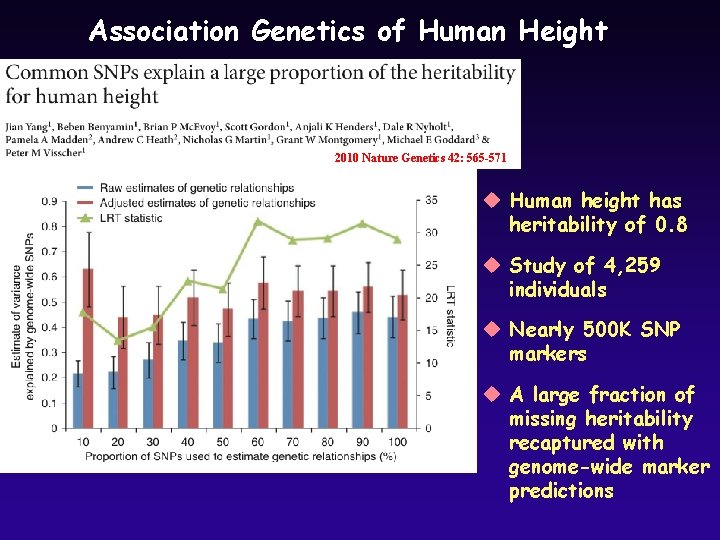
Association Genetics of Human Height 2010 Nature Genetics 42: 565 -571 u Human height has heritability of 0. 8 u Study of 4, 259 individuals u Nearly 500 K SNP markers u A large fraction of missing heritability recaptured with genome-wide marker predictions
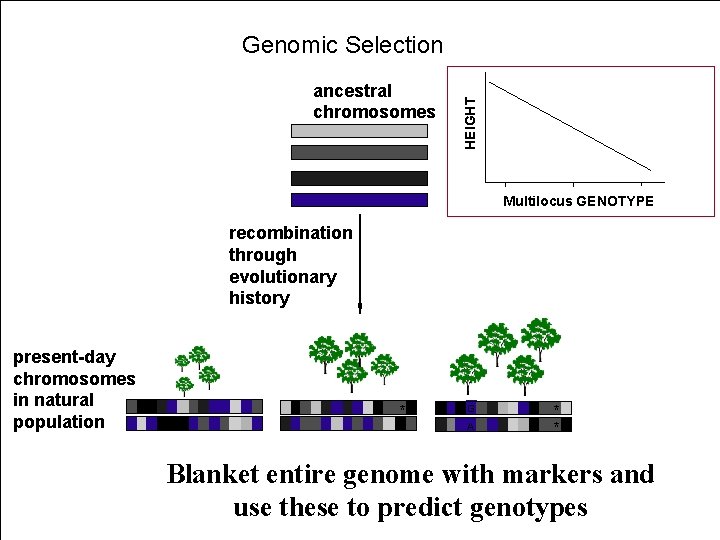
ancestral chromosomes HEIGHT Genomic Selection Multilocus GENOTYPE recombination through evolutionary history present-day chromosomes in natural population * G A * * Blanket entire genome with markers and use these to predict genotypes


Trait Heterogeneity: Height u Pygmy population has genome regions that show a high frequency of derived alleles (Ancestry-Informative Markers) and high divergence from other human populations (Locus-Specific Branch Length outliers) u Genes in these regions show association with height u Mechanisms are related to pituitary function: totally different than loci controlling height in Eurasian populations 2012 Cell 150: 457 -469 2
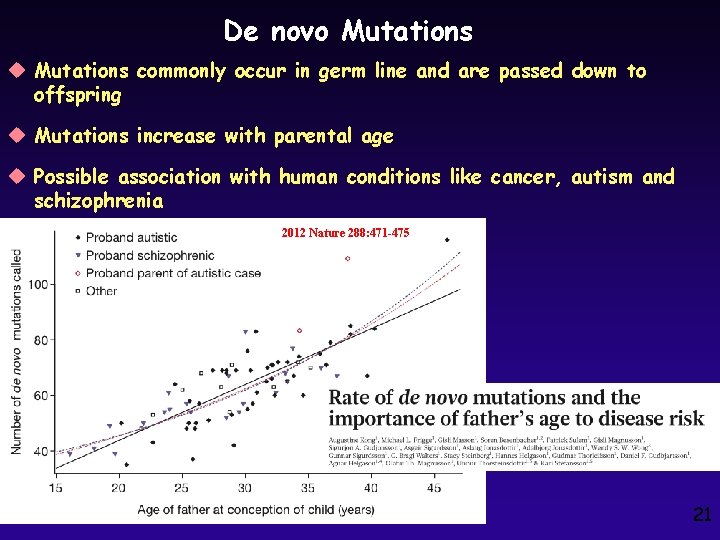
De novo Mutations u Mutations commonly occur in germ line and are passed down to offspring u Mutations increase with parental age u Possible association with human conditions like cancer, autism and schizophrenia 2012 Nature 288: 471 -475 21
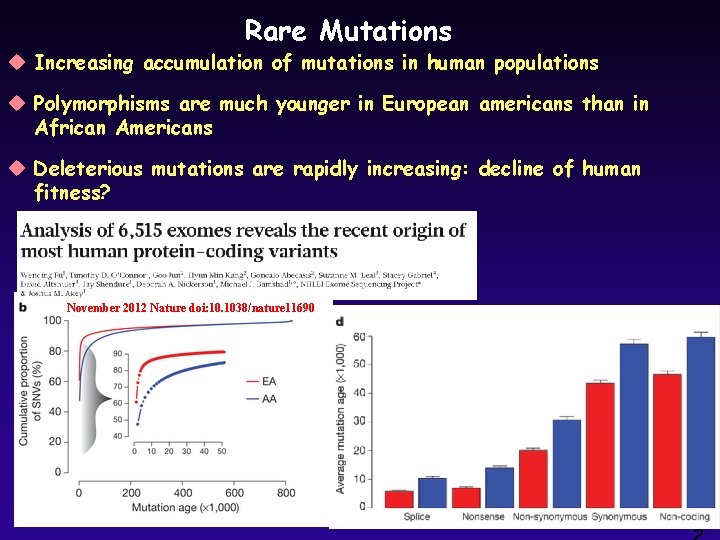
Rare Mutations u Increasing accumulation of mutations in human populations u Polymorphisms are much younger in European americans than in African Americans u Deleterious mutations are rapidly increasing: decline of human fitness? November 2012 Nature doi: 10. 1038/nature 11690 2
- Slides: 22