Lecture 2 Basic Population and Quantitative Genetics Allele

Lecture 2: Basic Population and Quantitative Genetics

Allele and Genotype Frequencies Given genotype frequencies, we can always compute allele frequencies, e. g. , The converse is not true: given allele frequencies we cannot uniquely determine the genotype frequencies For n alleles, there are n(n+1)/2 genotypes If we are willing to assume random mating, Hardy-Weinberg proportions

Hardy-Weinberg • Prediction of genotype frequencies from allele freqs • Allele frequencies remain unchanged over generations, provided: • Infinite population size (no genetic drift) • No mutation • No selection • No migration • Under HW conditions, a single generation of random mating gives genotype frequencies in Hardy-Weinberg proportions, and they remain forever in these proportions

Gametes and Gamete Frequencies When we consider two (or more) loci, we follow gametes Under random mating, gametes combine at random, e. g. Major complication: Even under HW conditions, gamete frequencies can change over time
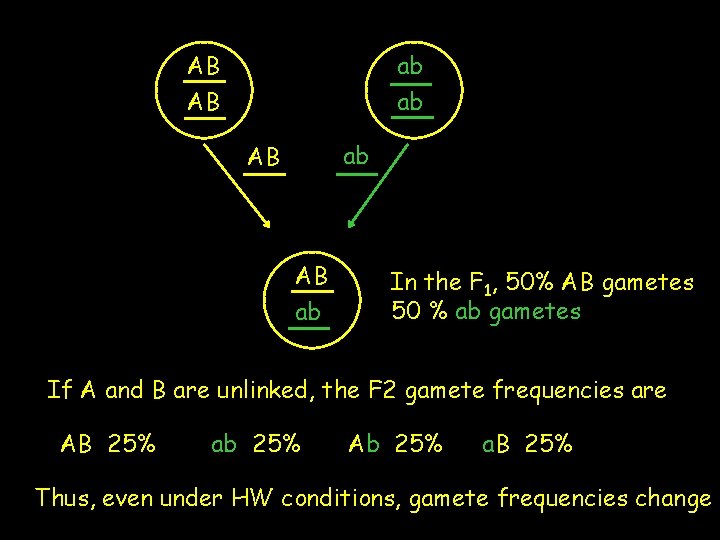
AB AB ab ab ab AB AB ab In the F 1, 50% AB gametes 50 % ab gametes If A and B are unlinked, the F 2 gamete frequencies are AB 25% ab 25% Ab 25% a. B 25% Thus, even under HW conditions, gamete frequencies change

Linkage disequilibrium Random mating and recombination eventually changes gamete frequencies so that they are in linkage equilibrium (LE). once in LE, gamete frequencies do not change (unless acted on by other forces) At LE, alleles in gametes are independent of each other: When linkage disequilibrium (LD) present, alleles are no longer independent --- knowing that one allele is in the gamete provides information on alleles at other loci The disequilibrium between alleles A and B is given by

The Decay of Linkage Disequilibrium The frequency of the AB gamete is given by Departure from If recombination frequency between the. LE A and B loci LE value is c, the disequilibrium in generation t is value the approach can be Note that D(t) ->Initial zero, LD although slow when c is very small

Contribution of a locus to a trait Basic model: P = G + E Genotypic Phenotypic Environmental value -- we will value occasionally also use z value for this G = average phenotypic forvalue that genotype if we are able to replicate it over the universe of environmental values G - E covariance -- higher performing animals may be disproportionately rewarded G x E interaction --- G values are different across environments. Basic model now becomes P = G + E + GE

Alternative parameterizations of Genotypic values Q 1 Q 1 Q 2 Q 2 C C C -a C + a(1+k) C+a+d C + 2 a C+a d measures dominance, with d. G(Q =+ 0 G(Q if) the heterozygote d = ak =G(Q ) - [G(Q Q ) ]/2 2 a =2 G(Q Q ) Q 1 Q 2 2 1 1 is exactly intermediate to the two homozygotes k = d/a is a scaled measure of the dominance

Example: Booroola (B) gene Genotype Average Litter size bb Bb 1. 48 2. 17 BB 2. 66 2 a = G(BB) - G(bb) = 2. 66 -1. 46 --> a = 0. 59 ak =d = G(Bb) - [ G(BB)+G(bb)]/2 = 0. 10 k = d/a = 0. 17

Fisher’s Decomposition of G One of Fisher’s key insights was that the genotypic value consists of a fraction that can be passed from parent to offspring and a fraction that cannot. Dominance deviations --the difference (for genotype Average contribution to genotypic value for allele i The Since genotypic parents value pass predicted along single from alleles theto individual their Mean value, with Aallelic Aj) between the genotypic value predicted from the ioffspring, effects the isathus i (the average effect of allele i) two single alleles the actual genotypic value, represent theseand contributions

Fisher’s decomposition is a Regression Predicted value. Residual A notational change clearly shows this is a error regression, Independent. Intercept (predictor)Regression variable Nslope =Regression # of Q 2 alleles residual
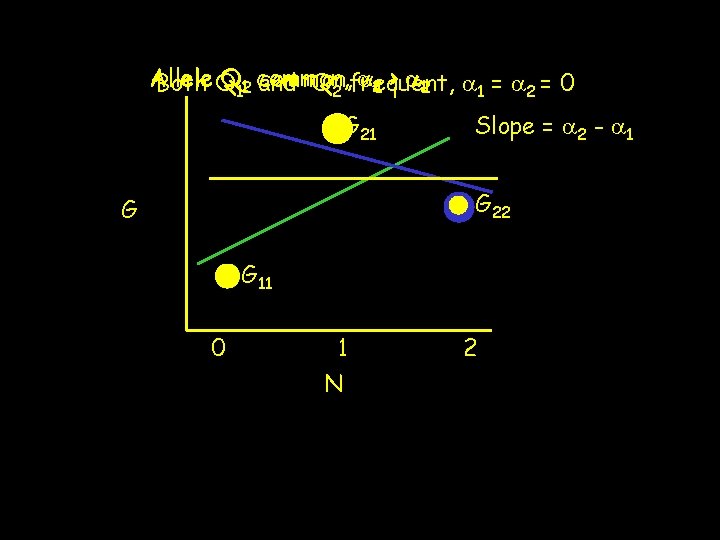
Allele Q 112 common, a 21 > a 12 a 1 = a 2 = 0 Both Q and Q 2 frequent, G 21 Slope = a 2 - a 1 G 22 G G 11 0 1 N 2

Consider a diallelic locus, where p 1 = freq(Q 1) Genotype Q 1 Q 1 Q 2 Q 2 Genotypic value 0 a(1+k) 2 a Mean Allelic effects Dominance deviations

Average effects and Breeding Values The a values are the average effects of an allele Breeders focus on breeding value (BV) ( ) Why all the fuss over the BV? Consider the offspring of a Qx. Qy sire mated to a random dam. What is the expected value of the offspring?

The expected value of an offspring is the expected value of Hence, For random and z alleles, this has an 0 For a random dam, w these have expected value of zero

We can thus estimate the BV for a sire by twice the deviation of his offspring from the pop mean, More generally, the expected value of an offspring is the average breeding value of its parents,
Genetic Variances As Cov(a, d) = 0 Dominance Genetic Variance Additive Genetic Variance (or simply. Variance) dominance variance) (or simply Additive
![Q 1 Q 1 Q 1 Q 2 Q 2 Q 2 Since E[a] Q 1 Q 1 Q 1 Q 2 Q 2 Q 2 Since E[a]](http://slidetodoc.com/presentation_image_h/0cf64a671fb32a0330f7651360172c96/image-19.jpg)
Q 1 Q 1 Q 1 Q 2 Q 2 Q 2 Since E[a] = 0, Var(a)0= E[(aa(1+k) -ma)2] = E[a 22 a ] One locus, 2 alleles: When dominance Dominancepresent, effects asymmetric function of allele additive variance Frequencies One locus, 2 alleles: Equals zero if k = of 0 This is a symmetric function allele frequencies

Additive variance, VA, with no dominance (k = 0) VA Allele frequency, p

Complete dominance (k = 1) VA VD Allele frequency, p

Overdominance (k = 2) VA VD Zero additive variance Allele frequency, p

Epistasis Dominance Additive Dominance xx Additive x. Dominant dominance value to -interactions interaction ------Breeding value These components are defined beinteraction uncorrelated, the interactions between interaction between two alleles aansingle the allele atdominance aallele at locus one (or orthogonal), so that the deviation at locus onewith locus atthe one with genotype locus a single with at allele the another, dominance at another e. g. deviation allele Ai and at another. genotype Bkj
- Slides: 23