Lecture 14 February 23 2016 Biotech 3 Protein

Lecture 14 February 23, 2016 Biotech 3

Protein Purification • A process intended to isolate one or more proteins from a mixture, usually from cells or whole tissue extracts. • Exploit protein properties for purification purposes. 1. 2. 3. 4. Physio-chemical properties – charge or hydrophobicity Binding affinity – Tags Size – large or small Biological activity – Substrate analogs

Protein Purification - Preparation Secreted or not secreted? Secreted – Purify protein from cell growth cell media Not secreted (intracellular) – Extract protein from cell culture 1. Repeated freeze thaw cycles 2. Sonication 3. Homogenization by high pressure (French press 4. Homogenization by grinding (bead mill) 5. Permeabilization by detergents (e. g. Triton X-100) and enzymes (e. g. lysozyme) Centrifuge Collect pellet if insoluble Collect supernatant if soluble

Ammonium Sulfate Precipitation • • Common 1 st step in protein purification. Add increasing amounts of ammonium sulfate to protein extract. Exposes hydrophobic groups on the protein. Hydrophobic groups attract other protein hydrophobic groups. Results in aggregation of protein. Protein precipitates from solution. Inexpensive. Reversible aggregation!!
Ammonium Sulfate Precipitation Steps
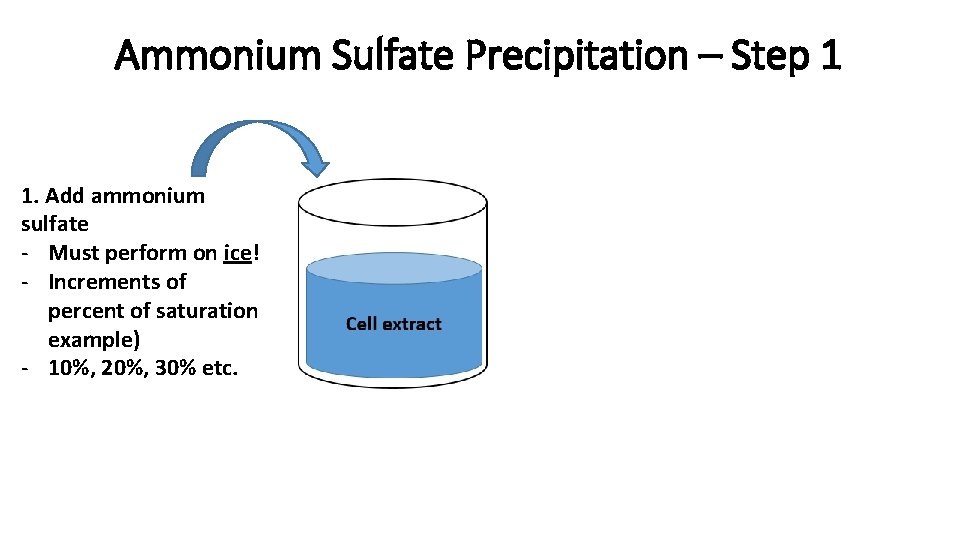
Ammonium Sulfate Precipitation – Step 1 1. Add ammonium sulfate - Must perform on ice! - Increments of percent of saturation example) - 10%, 20%, 30% etc.
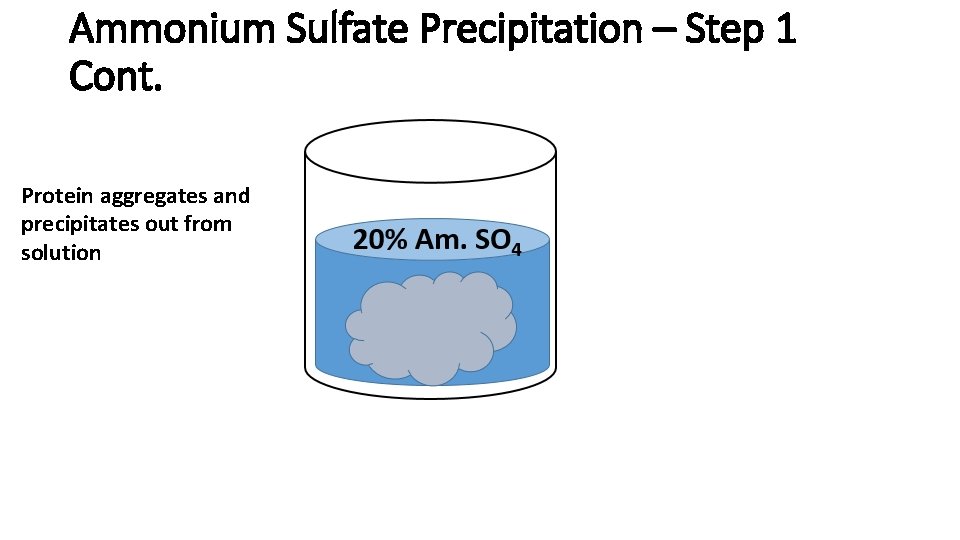
Ammonium Sulfate Precipitation – Step 1 Cont. Protein aggregates and precipitates out from solution
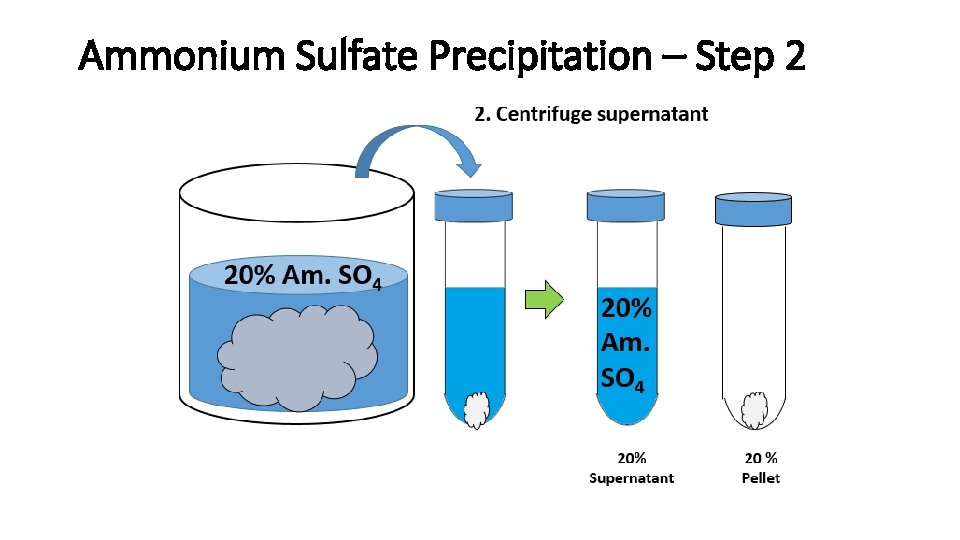
Ammonium Sulfate Precipitation – Step 2
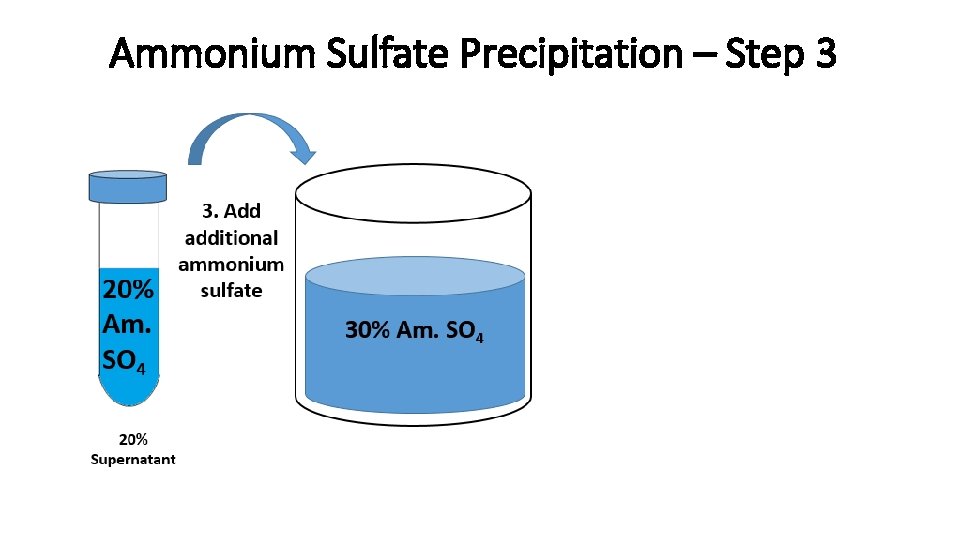
Ammonium Sulfate Precipitation – Step 3
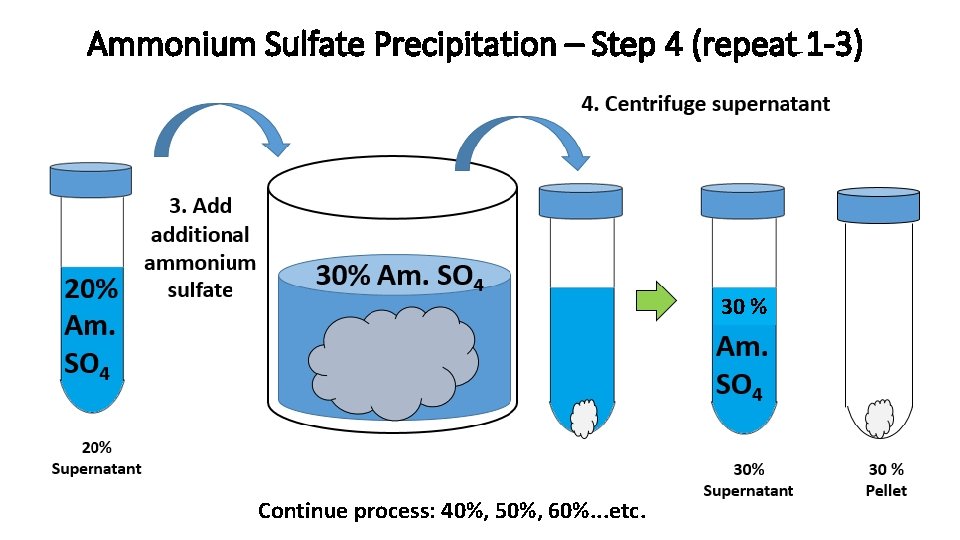
Ammonium Sulfate Precipitation – Step 4 (repeat 1 -3) 30 % Continue process: 40%, 50%, 60%. . . etc.
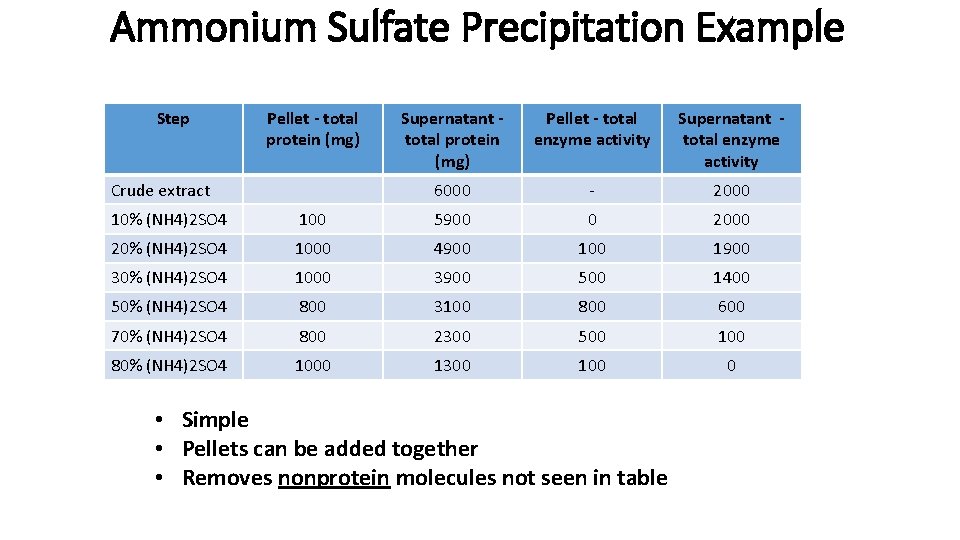
Ammonium Sulfate Precipitation Example Step Pellet - total protein (mg) Crude extract Supernatant total protein (mg) Pellet - total enzyme activity Supernatant total enzyme activity 6000 - 2000 10% (NH 4)2 SO 4 100 5900 0 2000 20% (NH 4)2 SO 4 1000 4900 1900 30% (NH 4)2 SO 4 1000 3900 500 1400 50% (NH 4)2 SO 4 800 3100 800 600 70% (NH 4)2 SO 4 800 2300 500 100 80% (NH 4)2 SO 4 1000 1300 100 0 • Simple • Pellets can be added together • Removes nonprotein molecules not seen in table

Protein Purification - Chromatography 1. Affinity 2. Ion Exchange 3. Hydrophobic interaction 4. Size exclusion 5. Reverse phase

1. Affinity Chromatography • • • Separates proteins based on specific interaction. Reversible interaction High selectivity High resolution High capacity Can obtain several 1000 -fold purity Example: Metal ion Affinity Chromatography Chelates metal ions such as Ni 2+ (Co 2+ , Cu 2+, Zn 2+)
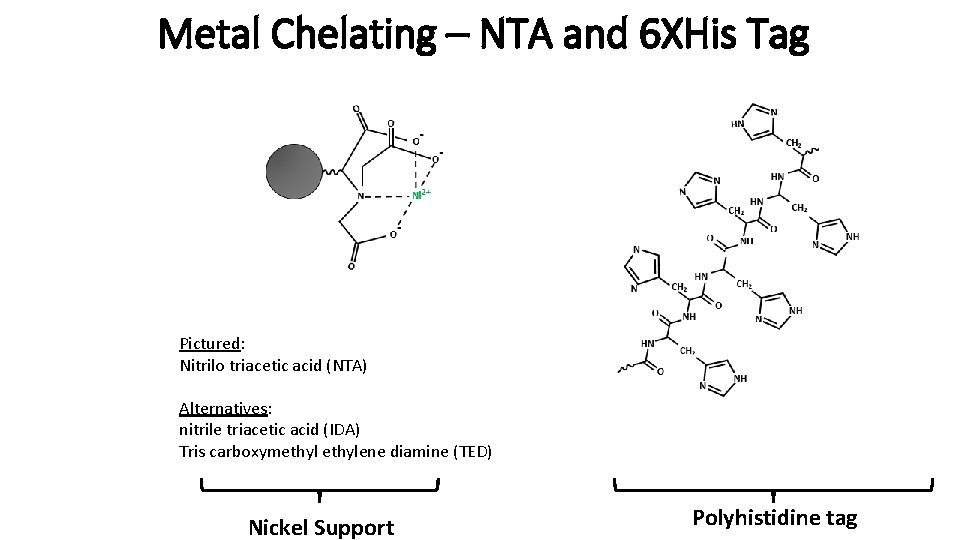
Metal Chelating – NTA and 6 XHis Tag Pictured: Nitrilo triacetic acid (NTA) Alternatives: nitrile triacetic acid (IDA) Tris carboxymethylene diamine (TED) Nickel Support Polyhistidine tag
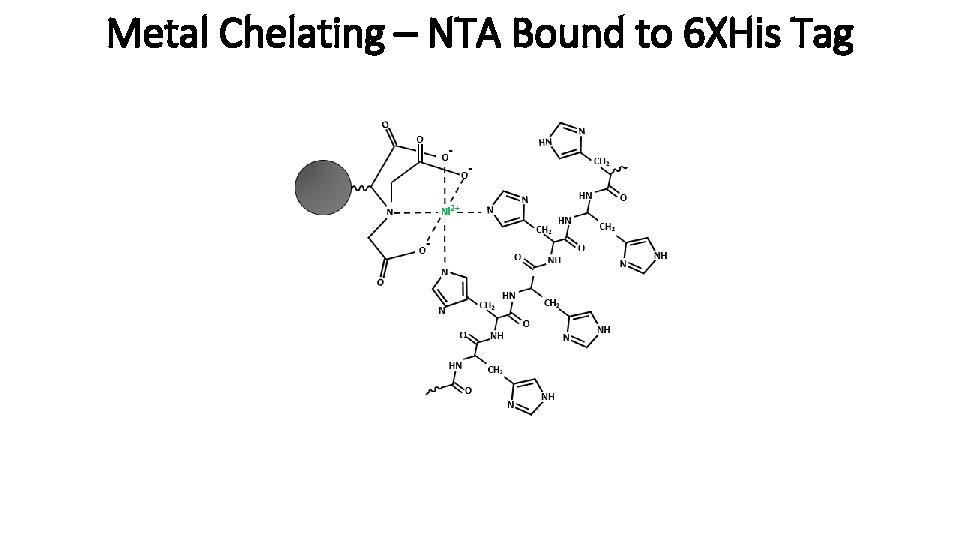
Metal Chelating – NTA Bound to 6 XHis Tag
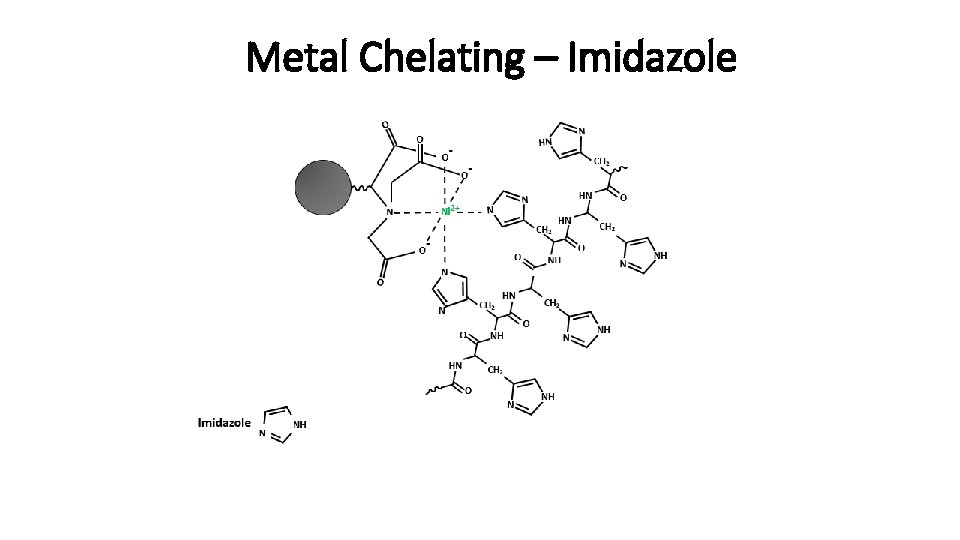
Metal Chelating – Imidazole
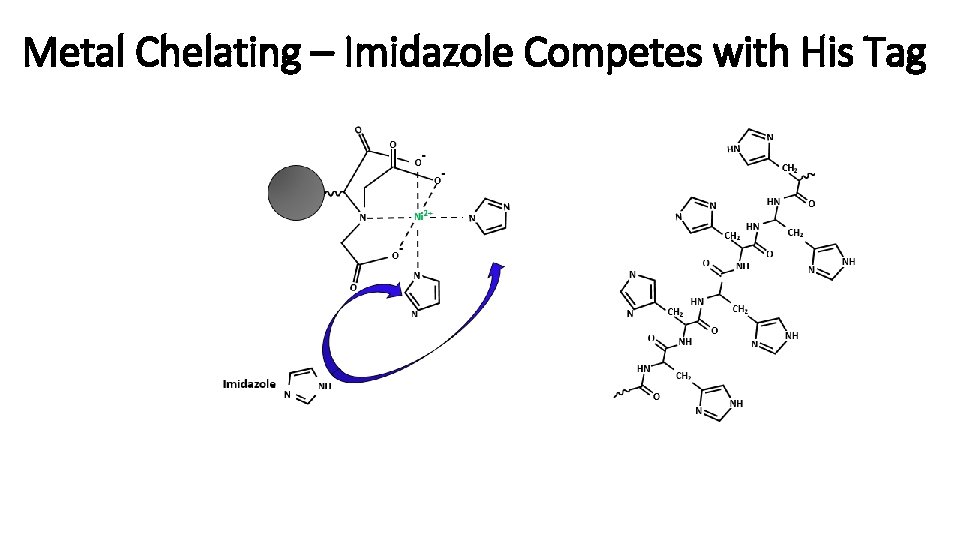
Metal Chelating – Imidazole Competes with His Tag
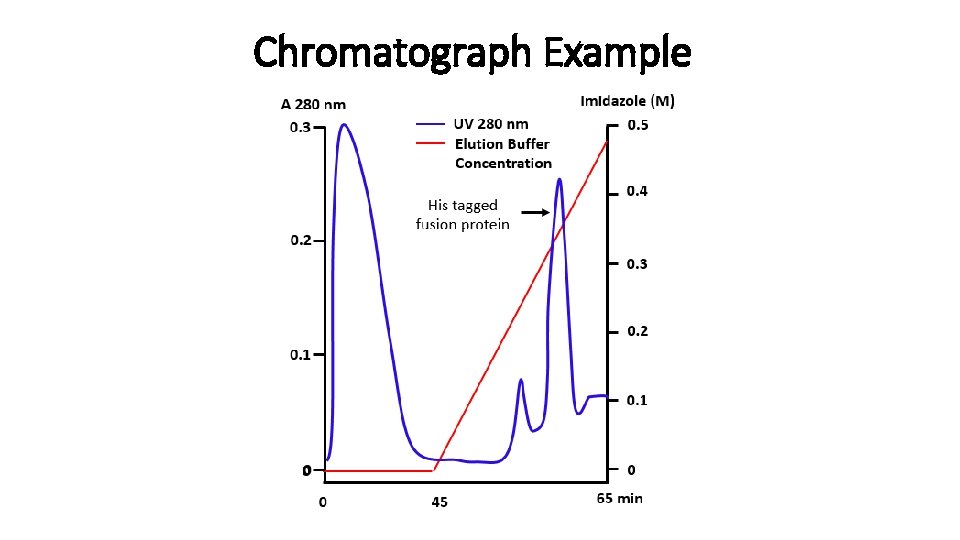
Chromatograph Example

Types of Affinity Chromatography Additional examples: a) Tags - Glutathione-S-Transferase (GST) tag - Maltose binding protein (MBP) tag b) c) Lectin affinity - Lectins are carbohydrate binding proteins - Concanavalin A is a lectin - Can purify glycosylated from non-glycosylated proteins d) Immunoaffinity (Antibodies) - Protein A e) Hormone, vitamin f) Enzyme - Substrate analogue, inhibitor, cofactor g) Nucleic acid

2. Ion Exchange Chromatography • Separates molecules based on charge • Beads of resin are modified so that they contain ac cationic or anionic functional group that can be positively or negatively charged. • Beads of resin are modified so that they contain a cationic or anionic functional group that can be positively or negatively charged. • Species of interest is applied to the column and the sample either binds to the resin or passes through the column. • A gradient of salt or p. H is used to elute the desired compound from the resin.

Cation Exchange Chromatography Cation Exchange – “Exchanging cations”, meaning the resin on the column is anionic which will binds to cations. • Strong and weak cation exchange groups. • Strength refers to extent of variation of ionization with p. H and not strength of binding. • Ionized at wide range of p. H. • Influences loading capacity of column at high or low p. H. Examples: S-Sepharose (Sulphopropyl) Strong CM-Sepharose (Carboxymethyl) Weak
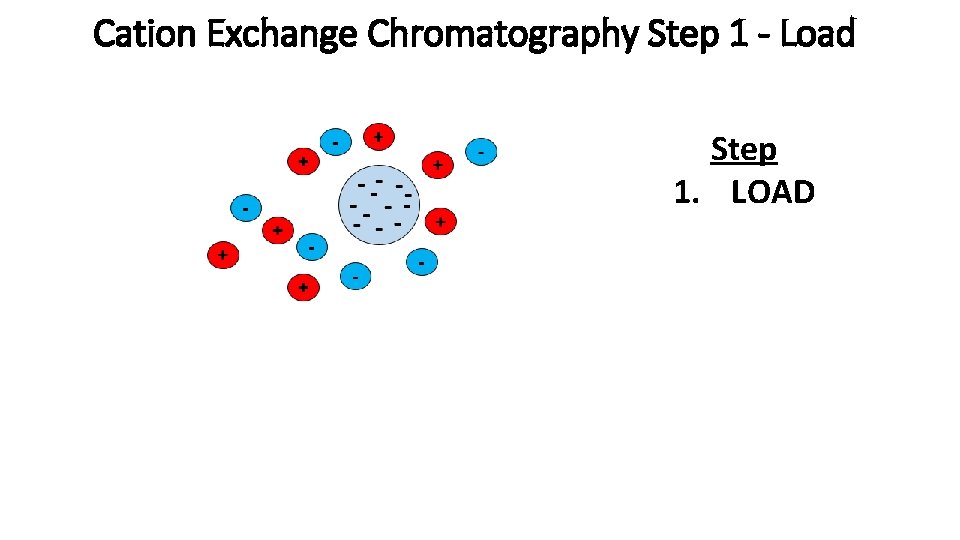
Cation Exchange Chromatography Step 1 - Load Step 1. LOAD
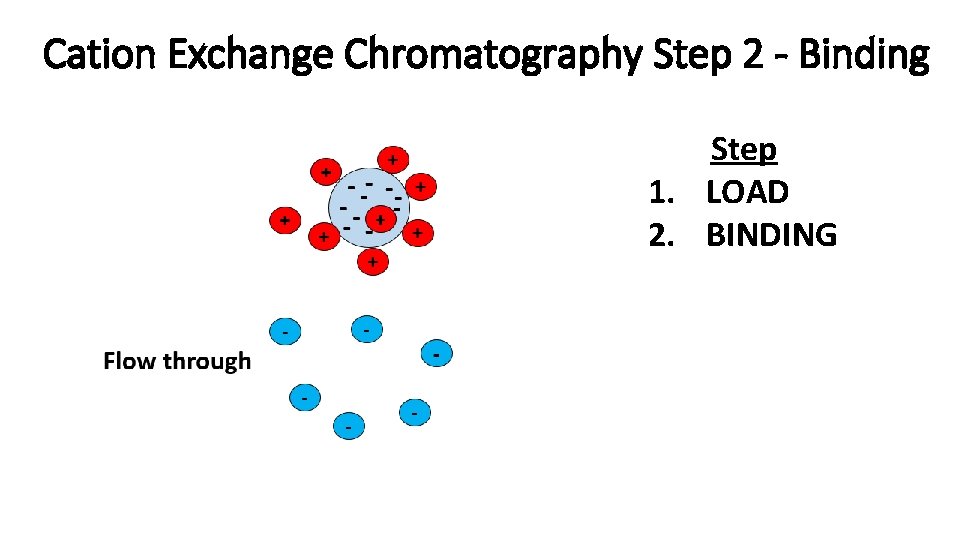
Cation Exchange Chromatography Step 2 - Binding Step 1. LOAD 2. BINDING

Cation Exchange Chromatography Step 3 - Wash Step 1. LOAD 2. BINDING 3. WASH
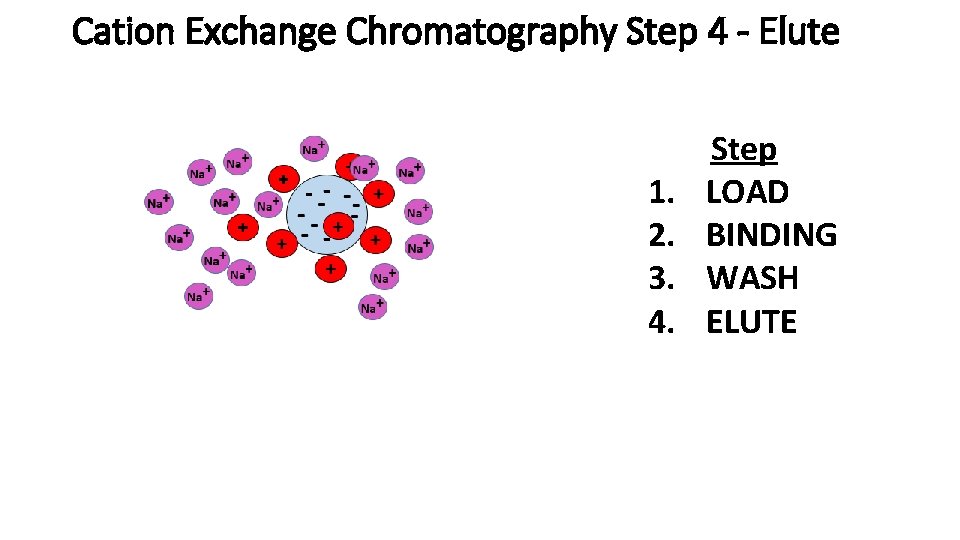
Cation Exchange Chromatography Step 4 - Elute 1. 2. 3. 4. Step LOAD BINDING WASH ELUTE
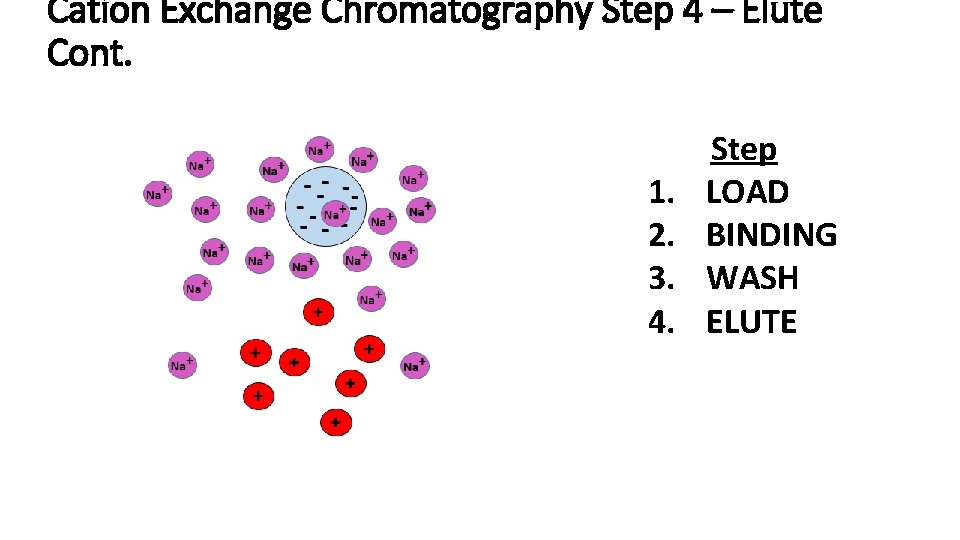
Cation Exchange Chromatography Step 4 – Elute Cont. 1. 2. 3. 4. Step LOAD BINDING WASH ELUTE
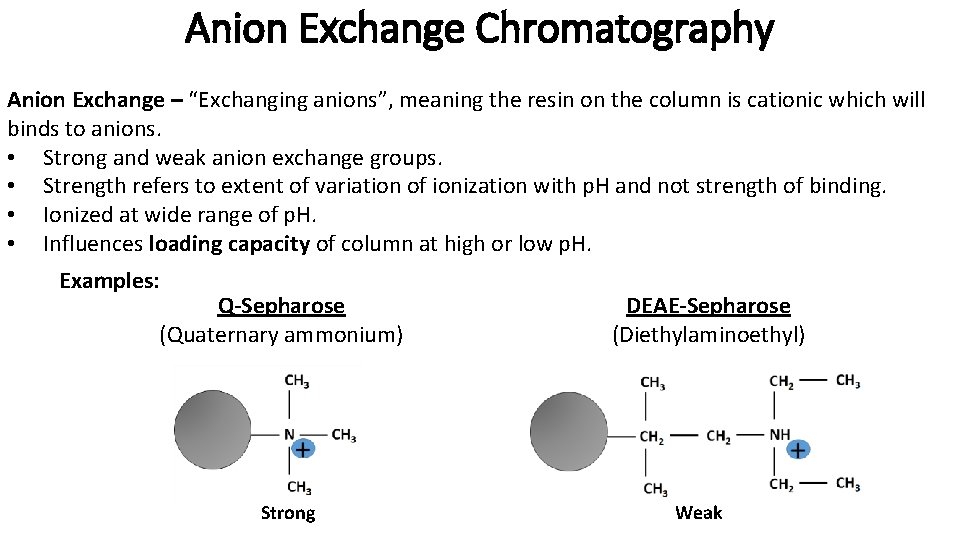
Anion Exchange Chromatography Anion Exchange – “Exchanging anions”, meaning the resin on the column is cationic which will binds to anions. • Strong and weak anion exchange groups. • Strength refers to extent of variation of ionization with p. H and not strength of binding. • Ionized at wide range of p. H. • Influences loading capacity of column at high or low p. H. Examples: Q-Sepharose (Quaternary ammonium) Strong DEAE-Sepharose (Diethylaminoethyl) Weak
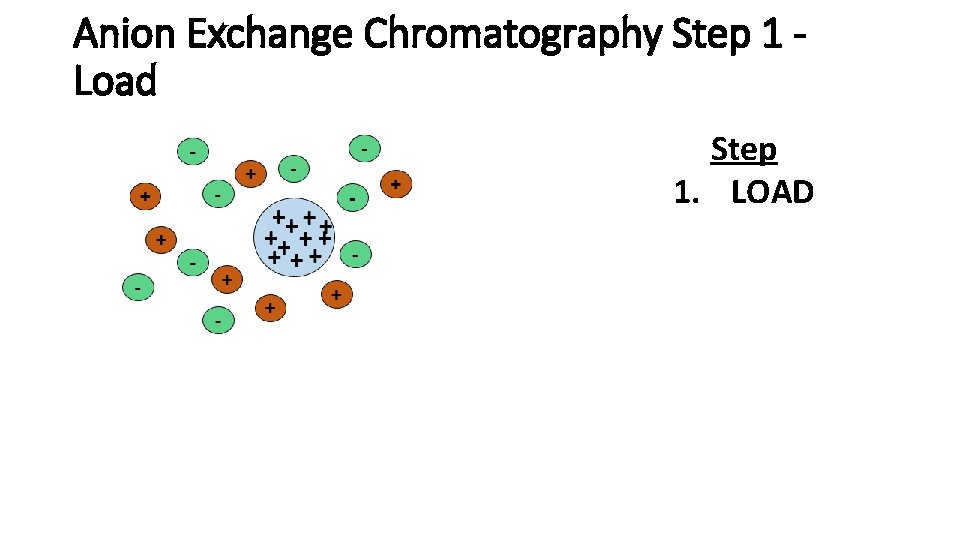
Anion Exchange Chromatography Step 1 Load Step 1. LOAD
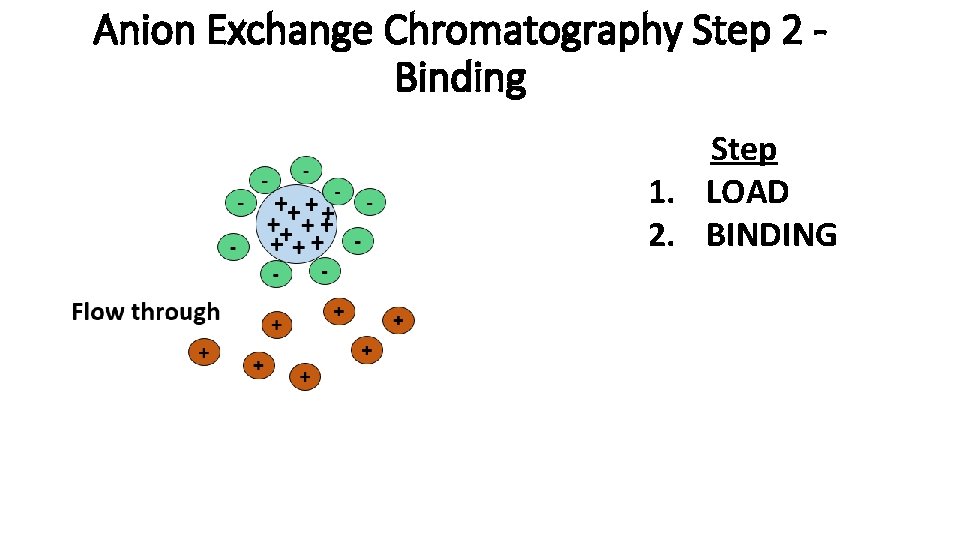
Anion Exchange Chromatography Step 2 Binding Step 1. LOAD 2. BINDING
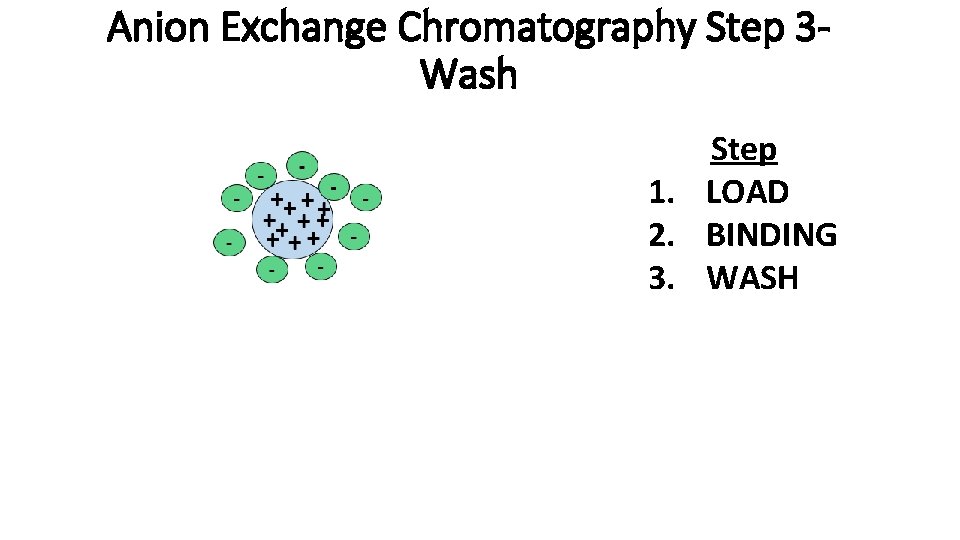
Anion Exchange Chromatography Step 3 Wash Step 1. LOAD 2. BINDING 3. WASH
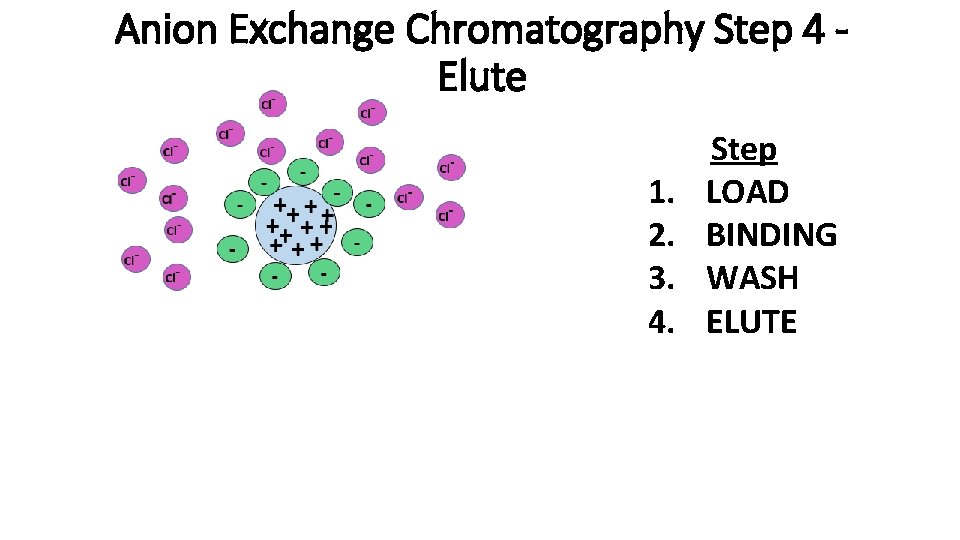
Anion Exchange Chromatography Step 4 Elute 1. 2. 3. 4. Step LOAD BINDING WASH ELUTE

Anion Exchange Chromatography Step 4 – Elute Cont. 1. 2. 3. 4. Step LOAD BINDING WASH ELUTE

Isoelectric Point (p. I) - is the p. H point where the net chare of a protein is zero. Environment p. H is lower + Environment < • Lower p. H means more protons in solution • Protein will be more protonated • Protein will carry a greater POSITIVE charge p. I < p. H is greater - • Higher p. H means less protons in solution • Protein will be more deprotonated • Protein will carry a greater NEGATIVE charge
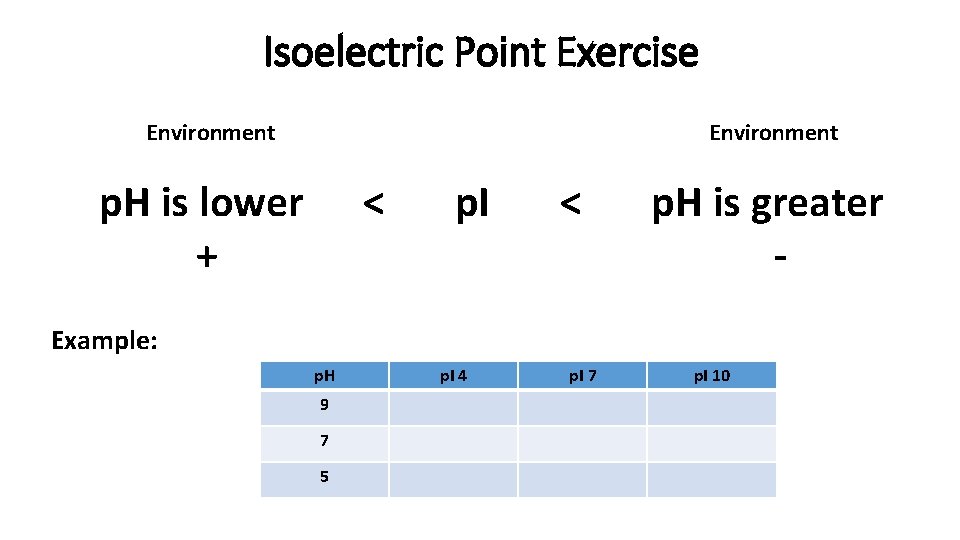
Isoelectric Point Exercise Environment p. H is lower + < p. I < p. H is greater - Example: p. H 9 7 5 p. I 4 p. I 7 p. I 10
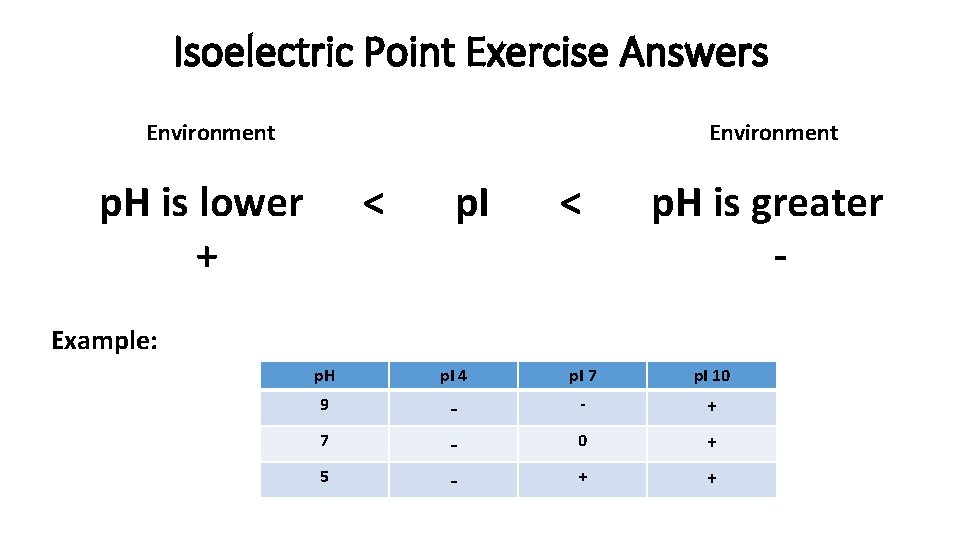
Isoelectric Point Exercise Answers Environment p. H is lower + < p. I < p. H is greater - Example: p. H p. I 4 p. I 7 p. I 10 9 - - + 0 + + + 7 5

Protein Sequence Analysis Protein p. I • How do we know the p. I of a protein? MLKRCLSPLTLVNQVALIVLLSTAIGLAGMAVSGWLVQGVQGSAHAINKAGSLRMQSYRLLAAVPLSEKDKPLIKEMEQTAFSAELTRAAER DGQLAQLQGLQDYWRNELIPALMRAQNRETVSADVSQFVAGLDQLVSGFDRTTEMRIETVVLVHRVMAVFMALLLVFTIIWLRARLLQP WRQLLAMASAVSHRDFTQRANISGRNEMAMLGTALNNMSAELAESYAVLEQRVQEKTAGLEHKNQILSFLWQANRRLHSRAPLCERLSP VLNGLQNLTLLRDIELRVYDTDDEENHQEFTCQPDMTCDDKGCQLCPRGVLPVGDRGTTLKWRLADSHTQYGILLATLPQGRHLSHDQQQ LVDTLVEQLTATLALDRHQERQQQLIVMEERATIARELHDSIAQSLSCMKMQVSCLQMQGDALPESSRELLSQIRNELNASWAQLRELLTTF RLQLTEPGLRPALEASCEEYSAKFGFPVKLDYQLPPRLVPSHQAIHLLQIAREALSNALKHSQASEVVVTVAQNDNQVKLTVQDNGCGVPEN AIRSNHYGMIIMRDRAQSLRGDCRVRRRESGGTEVVVTFIPEKTFTDVQGDTHE

Ex. PASy – Bioinformatics Resource Portal Protein p. I • How do we know the p. I of a protein? • Check on Ex. PASy – Bioinformatics Resource Portal • http: //www. expasy. org/ Sequence of interest MLKRCLSPLTLVNQVALIVLLSTAIGLAGMAVSGWLVQGVQGSAHAINK AGSLRMQSYRLLAAVPLSEKDKPLIKEMEQTAFSAELTRAAERDGQLAQ LQGLQDYWRNELIPALMRAQNRETVSADVSQFVAGLDQLVSGFDRTTE MRIETVVLVHRVMAVFMALLLVFTIIWLRARLLQPWRQLLAMASAVSH RDFTQRANISGRNEMAMLGTALNNMSAELAESYAVLEQRVQEKTAGLE HKNQILSFLWQANRRLHSRAPLCERLSPVLNGLQNLTLLRDIELRVYDTD DEENHQEFTCQPDMTCDDKGCQLCPRGVLPVGDRGTTLKWRLADSHT QYGILLATLPQGRHLSHDQQQLVDTLVEQLTATLALDRHQERQQQLIV MEERATIARELHDSIAQSLSCMKMQVSCLQMQGDALPESSRELLSQIRN ELNASWAQLRELLTTFRLQLTEPGLRPALEASCEEYSAKFGFPVKLDYQLP PRLVPSHQAIHLLQIAREALSNALKHSQASEVVVTVAQNDNQVKLTVQD NGCGVPENAIRSNHYGMIIMRDRAQSLRGDCRVRRRESGGTEVVVTFIP EKTFTDVQGDTHE Theoretical p. I = 6. 27
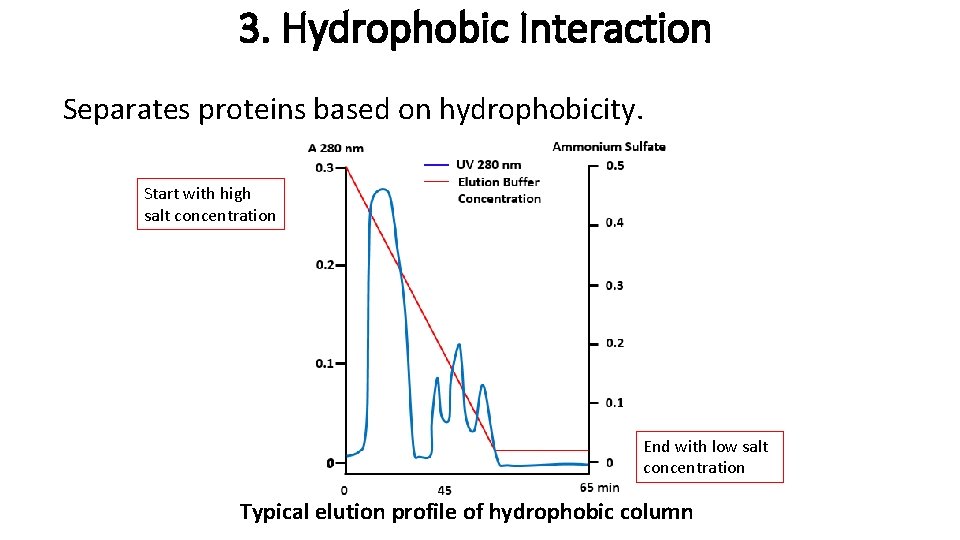
3. Hydrophobic Interaction Separates proteins based on hydrophobicity. Start with high salt concentration End with low salt concentration Typical elution profile of hydrophobic column

3. Hydrophobic Interaction – Role of Water Role of water • Good solvent for polar substances • Poor solvent for non-polar substances • Highly ordered “water shells” surround hydrophobic surfaces of ligands and proteins • Hydrophobic substance merge to minimize exposed area. • Hydrophobic proteins bind to hydrophobic column ligands. • Interaction of protein with column matrix may depend on van der Waals forces which increase as the structured order of water increases

3. Hydrophobic Interaction – Role of Salt • Principle is similar to “salting out” proteins • High salt concentrations sequester water molecules • Decrease salt concentration to elute protein. • Relative effectiveness of protein precipitation (promote hydrophobic interaction): Na 2 SO 4 > KSO 4 > (NH 4)SO 4 > Na 2 HPO 4 > Na. Cl > Li. Cl
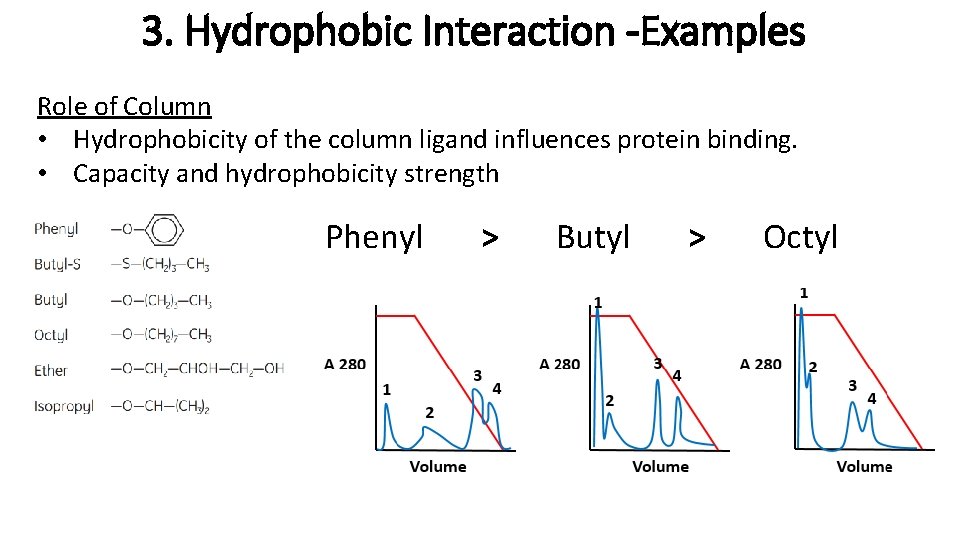
3. Hydrophobic Interaction -Examples Role of Column • Hydrophobicity of the column ligand influences protein binding. • Capacity and hydrophobicity strength Phenyl > Butyl > Octyl
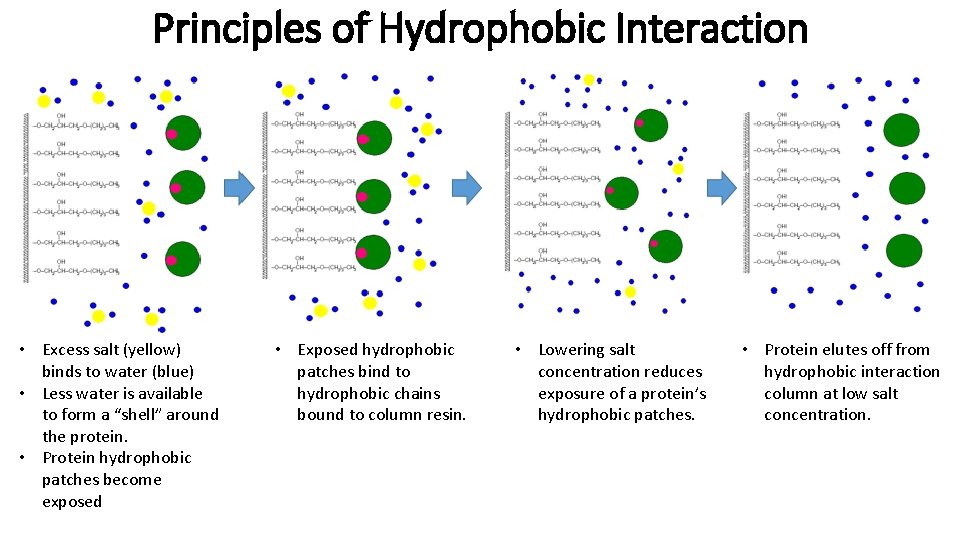
Principles of Hydrophobic Interaction • Excess salt (yellow) binds to water (blue) • Less water is available to form a “shell” around the protein. • Protein hydrophobic patches become exposed • Exposed hydrophobic patches bind to hydrophobic chains bound to column resin. • Lowering salt concentration reduces exposure of a protein’s hydrophobic patches. • Protein elutes off from hydrophobic interaction column at low salt concentration.

Protein Sequence Analysis Cont. Role of Protein • How do we know if protein of interest is hydrophobic? MLKRCLSPLTLVNQVALIVLLSTAIGLAGMAVSGWLVQGVQGSAHAINKAGSLRMQSYRLLAAVPLSEKDKPLIKEMEQTAFSAELTRAAER DGQLAQLQGLQDYWRNELIPALMRAQNRETVSADVSQFVAGLDQLVSGFDRTTEMRIETVVLVHRVMAVFMALLLVFTIIWLRARLLQP WRQLLAMASAVSHRDFTQRANISGRNEMAMLGTALNNMSAELAESYAVLEQRVQEKTAGLEHKNQILSFLWQANRRLHSRAPLCERLSP VLNGLQNLTLLRDIELRVYDTDDEENHQEFTCQPDMTCDDKGCQLCPRGVLPVGDRGTTLKWRLADSHTQYGILLATLPQGRHLSHDQQQ LVDTLVEQLTATLALDRHQERQQQLIVMEERATIARELHDSIAQSLSCMKMQVSCLQMQGDALPESSRELLSQIRNELNASWAQLRELLTTF RLQLTEPGLRPALEASCEEYSAKFGFPVKLDYQLPPRLVPSHQAIHLLQIAREALSNALKHSQASEVVVTVAQNDNQVKLTVQDNGCGVPEN AIRSNHYGMIIMRDRAQSLRGDCRVRRRESGGTEVVVTFIPEKTFTDVQGDTHE

Ex. PASy – Bioinformatics Resource Portal – Transmembrane region determination Role of Protein • How do we know if protein of interest is hydrophobic or has transmembrane regions? • Check on Ex. PASy – Bioinformatics Resource Portal • http: //www. expasy. org/ MLKRCLSPLTLVNQVALIVLLSTAIGLAGMAVSGWLVQGVQGSAHAINKAGSLRMQSYRLLAAVPLSEKDKPLIKEMEQTAFSAELTRAAER DGQLAQLQGLQDYWRNELIPALMRAQNRETVSADVSQFVAGLDQLVSGFDRTTEMRIETVVLVHRVMAVFMALLLVFTIIWLRARLLQP WRQLLAMASAVSHRDFTQRANISGRNEMAMLGTALNNMSAELAESYAVLEQRVQEKTAGLEHKNQILSFLWQANRRLHSRAPLCERLSP VLNGLQNLTLLRDIELRVYDTDDEENHQEFTCQPDMTCDDKGCQLCPRGVLPVGDRGTTLKWRLADSHTQYGILLATLPQGRHLSHDQQQ LVDTLVEQLTATLALDRHQERQQQLIVMEERATIARELHDSIAQSLSCMKMQVSCLQMQGDALPESSRELLSQIRNELNASWAQLRELLTTF RLQLTEPGLRPALEASCEEYSAKFGFPVKLDYQLPPRLVPSHQAIHLLQIAREALSNALKHSQASEVVVTVAQNDNQVKLTVQDNGCGVPEN AIRSNHYGMIIMRDRAQSLRGDCRVRRRESGGTEVVVTFIPEKTFTDVQGDTHE
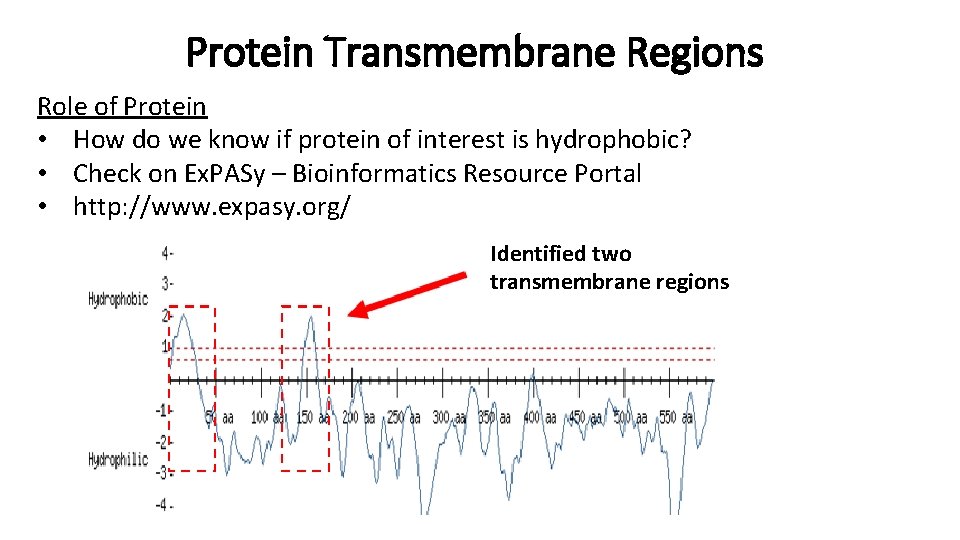
Protein Transmembrane Regions Role of Protein • How do we know if protein of interest is hydrophobic? • Check on Ex. PASy – Bioinformatics Resource Portal • http: //www. expasy. org/ Identified two transmembrane regions
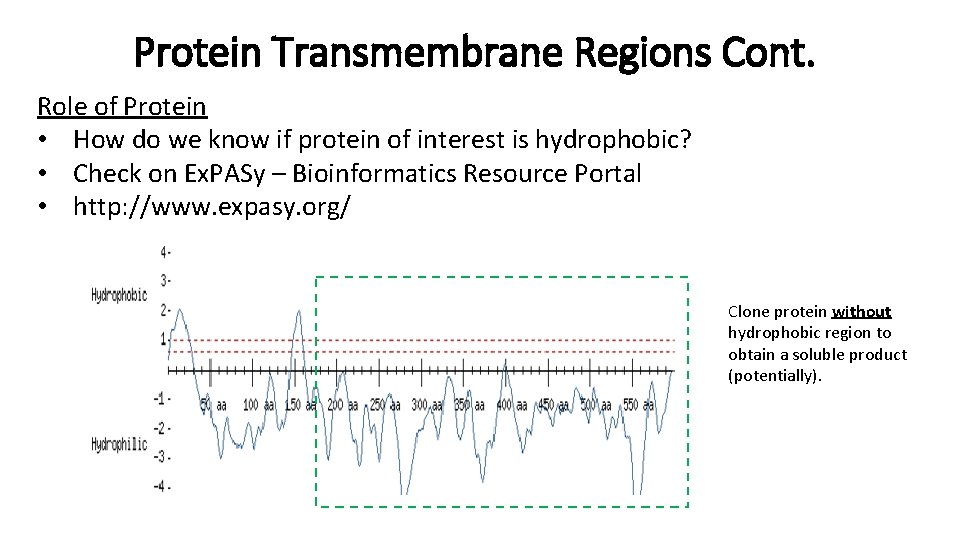
Protein Transmembrane Regions Cont. Role of Protein • How do we know if protein of interest is hydrophobic? • Check on Ex. PASy – Bioinformatics Resource Portal • http: //www. expasy. org/ Clone protein without hydrophobic region to obtain a soluble product (potentially).

Reverse Phase Chromatography • Organic solvent running buffer solutions (ex methanol, propanol, tetrahydrofuran, acetonitrile) • Nonpolar carbon chain beads (C 2 to C 18 bound to silica) • Beads are not polysacharide based • Higher pressure • Beads do not collapse • Higher resolution • Smaller particle size determination

4. Size Exclusion Chromatography Also known as “Gel Filtration Chromatography”. Separate proteins based on size. Column with resin that has small holes. Smaller proteins will enter holes more readily than larger proteins. Smaller proteins will thus be retained more than larger proteins and elute at a later volume. • Larger proteins will move more readily through column and elute at an earlier volume. • There is no competitive “elution buffer”. • • •
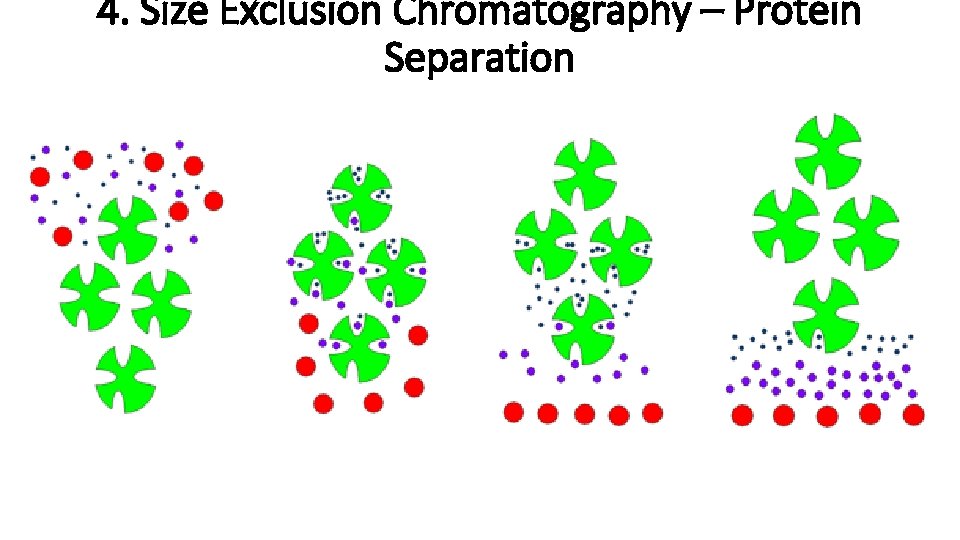
4. Size Exclusion Chromatography – Protein Separation
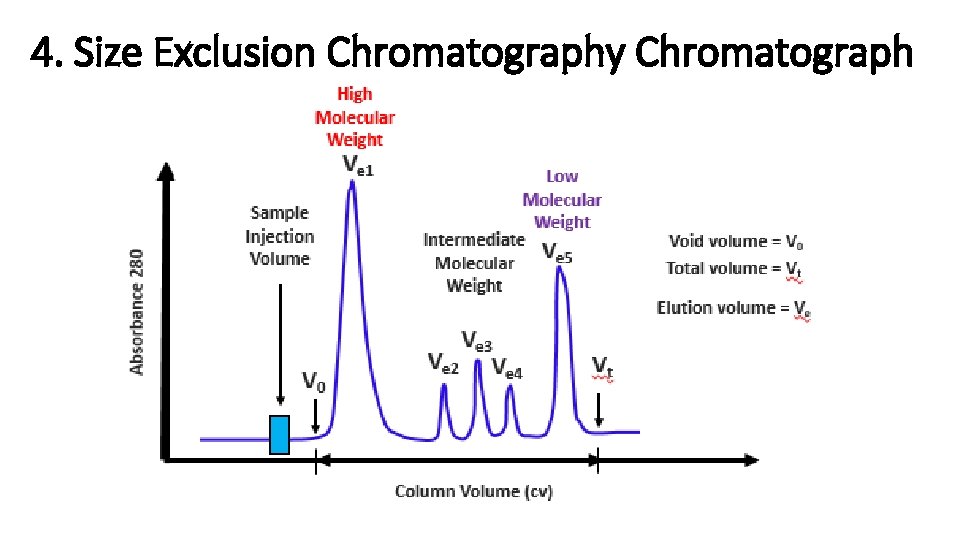
4. Size Exclusion Chromatography Chromatograph
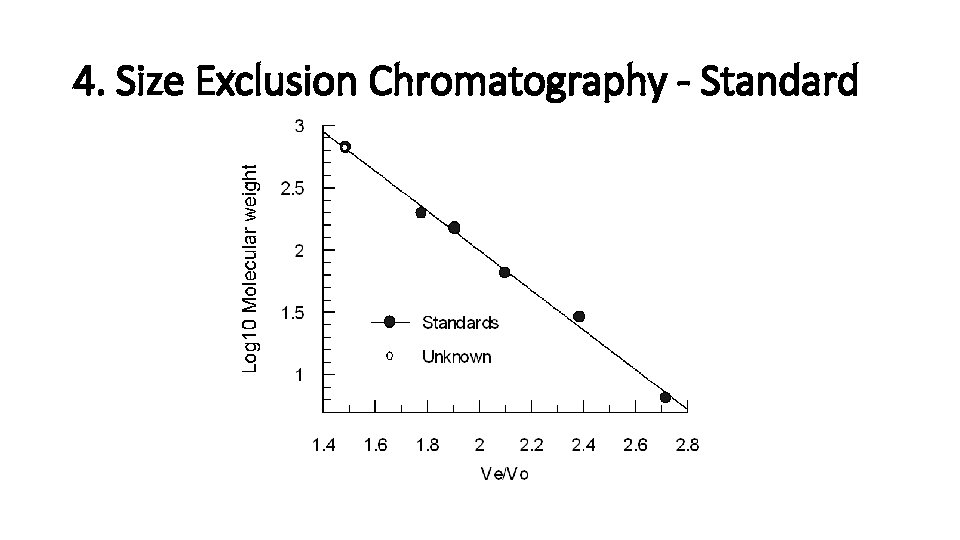
4. Size Exclusion Chromatography - Standard

4. Size Exclusion Chromatography – Why? Why do we care for the size, can’t we just look up the molecular weight online?

4. Size Exclusion Chromatography – FUN! Why do we care for the size, can’t we just look up the molecular weight online? • It’s FUN to perform protein chromatography! • But that’s not the only reason, there’s more to it than that!

4. Size Exclusion Chromatography – Analysis Method Why do we care for the size, can’t we just look up the molecular weight online? • It’s FUN to perform protein chromatography! • Can determine the Native molecular weight. • Purification step. • Analytical • Bound cofactors • Multimeric protein • Determine number of subunits (Quaternary structure)

Example Protein MLKRCLSPLTLVNQVALIVLLSTAIGLAGMAVSGWLVQGVQGSAHAINKAGSLRMQSYRLLAAVPLSEKDKPLIKEMEQTAFSAELTRAAER DGQLAQLQGLQDYWRNELIPALMRAQNRETVSADVSQFVAGLDQLVSGFDRTTEMRIETVVLVHRVMAVFMALLLVFTIIWLRARLLQP WRQLLAMASAVSHRDFTQRANISGRNEMAMLGTALNNMSAELAESYAVLEQRVQEKTAGLEHKNQILSFLWQANRRLHSRAPLCERLSP VLNGLQNLTLLRDIELRVYDTDDEENHQEFTCQPDMTCDDKGCQLCPRGVLPVGDRGTTLKWRLADSHTQYGILLATLPQGRHLSHDQQQ LVDTLVEQLTATLALDRHQERQQQLIVMEERATIARELHDSIAQSLSCMKMQVSCLQMQGDALPESSRELLSQIRNELNASWAQLRELLTTF RLQLTEPGLRPALEASCEEYSAKFGFPVKLDYQLPPRLVPSHQAIHLLQIAREALSNALKHSQASEVVVTVAQNDNQVKLTVQDNGCGVPEN AIRSNHYGMIIMRDRAQSLRGDCRVRRRESGGTEVVVTFIPEKTFTDVQGDTHE

Example Protein Characteristics Full length MW 67 KDa Truncated at Met 219 Truncated MW 44. 2 KDa (With His Tag) p. I = 5. 94 MLKRCLSPLTLVNQVALIVLLSTAIGLAGMAVSGWLVQGVQGSAHAINKAGSLRMQSYRLLAAVPLSEKDKPLIKEMEQTAFSAELTRAAER DGQLAQLQGLQDYWRNELIPALMRAQNRETVSADVSQFVAGLDQLVSGFDRTTEMRIETVVLVHRVMAVFMALLLVFTIIWLRARLLQP WRQLLAMASAVSHRDFTQRANISGRNEMAMLGTALNNMSAELAESYAVLEQRVQEKTAGLEHKNQILSFLWQANRRLHSRAPLCERLS PVLNGLQNLTLLRDIELRVYDTDDEENHQEFTCQPDMTCDDKGCQLCPRGVLPVGDRGTTLKWRLADSHTQYGILLATLPQGRHLSHDQ QQLVDTLVEQLTATLALDRHQERQQQLIVMEERATIARELHDSIAQSLSCMKMQVSCLQMQGDALPESSRELLSQIRNELNASWAQLRE LLTTFRLQLTEPGLRPALEASCEEYSAKFGFPVKLDYQLPPRLVPSHQAIHLLQIAREALSNALKHSQASEVVVTVAQNDNQVKLTVQDNG CGVPENAIRSNHYGMIIMRDRAQSLRGDCRVRRRESGGTEVVVTFIPEKTFTDVQGDTHEHHHHHH
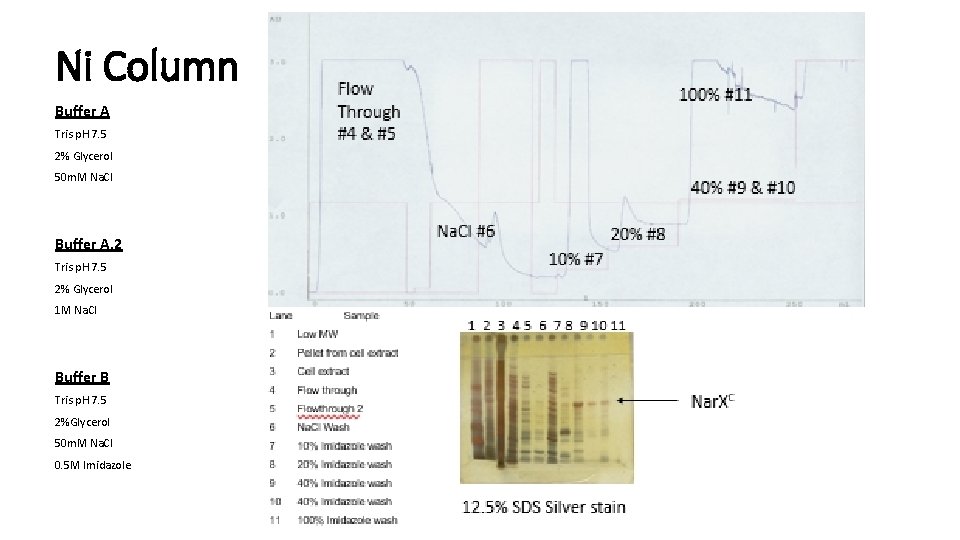
Ni Column Buffer A Tris p. H 7. 5 2% Glycerol 50 m. M Na. Cl Buffer A. 2 Tris p. H 7. 5 2% Glycerol 1 M Na. Cl Buffer B Tris p. H 7. 5 2%Glycerol 50 m. M Na. Cl 0. 5 M Imidazole
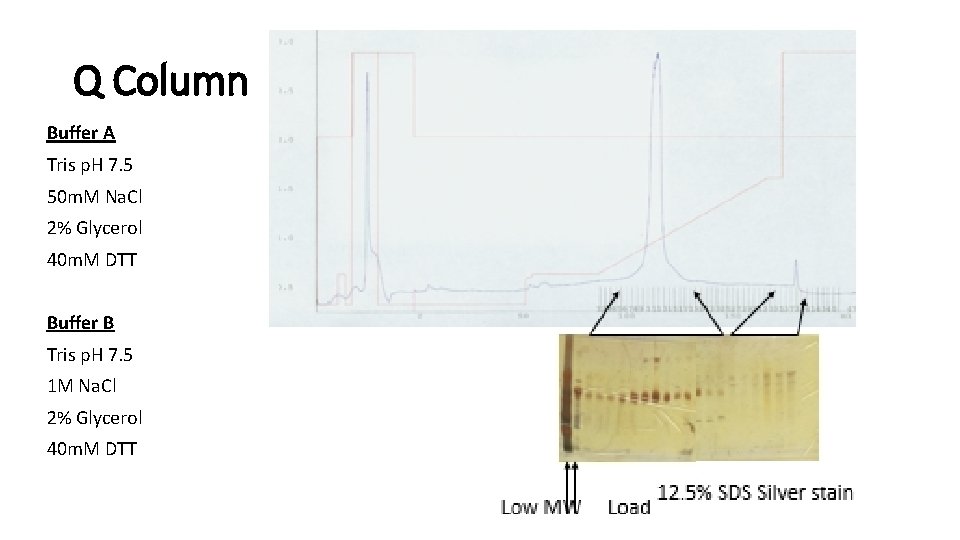
Q Column Buffer A Tris p. H 7. 5 50 m. M Na. Cl 2% Glycerol 40 m. M DTT Buffer B Tris p. H 7. 5 1 M Na. Cl 2% Glycerol 40 m. M DTT
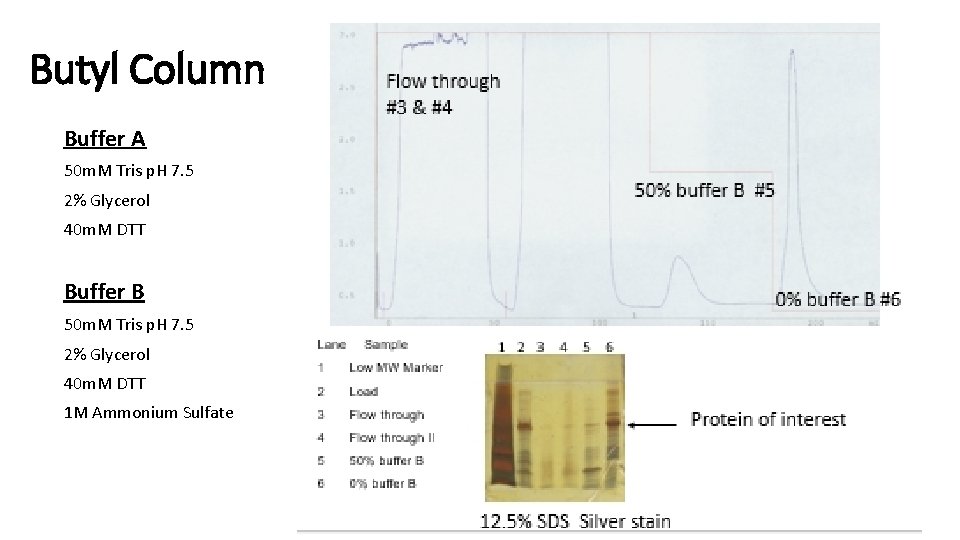
Butyl Column Buffer A 50 m. M Tris p. H 7. 5 2% Glycerol 40 m. M DTT Buffer B 50 m. M Tris p. H 7. 5 2% Glycerol 40 m. M DTT 1 M Ammonium Sulfate

Superdex 200 Superdex Buffer 50 m. M Tris p. H 7. 5 0. 3 M Na. Cl 2% Glycerol 40 m. M DTT
- Slides: 60