Lecture 13 Regulation of Gene Expression Part 2






- Slides: 29

Lecture #13: Regulation of Gene Expression (Part 2)

Lecture #13 Reading Assignment Physiology & Biochemistry of Prokaryotes 3 rd Ed. : Ch. 10, pp. 291; Ch. 18, pp. 496 -503, 554 -561, 509 -518 Brock Biology of Microorganisms 11 th Ed: Ch. 8, pp. 221 -227 Brock Biology of Microorganisms 12 th ed. : Ch. 9, pp. 231 -234, 244 -248 Article(s) to be discussed Majdalani Review (Bacterial Small RNA regulators) Small Things Considered Essay for Lecture #13

Topics: § Transcriptional control by attenuation § Two-component regulatory systems § Quorum sensing § Bacterial small RNA regulators

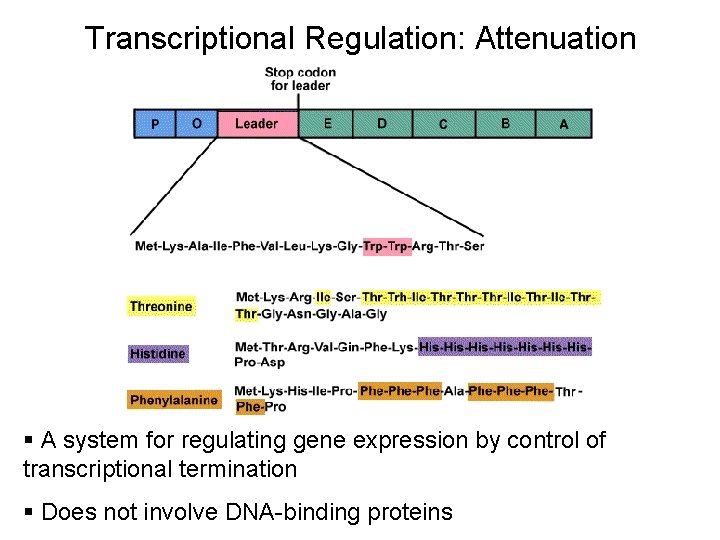
Transcriptional Regulation: Attenuation § A system for regulating gene expression by control of transcriptional termination § Does not involve DNA-binding proteins

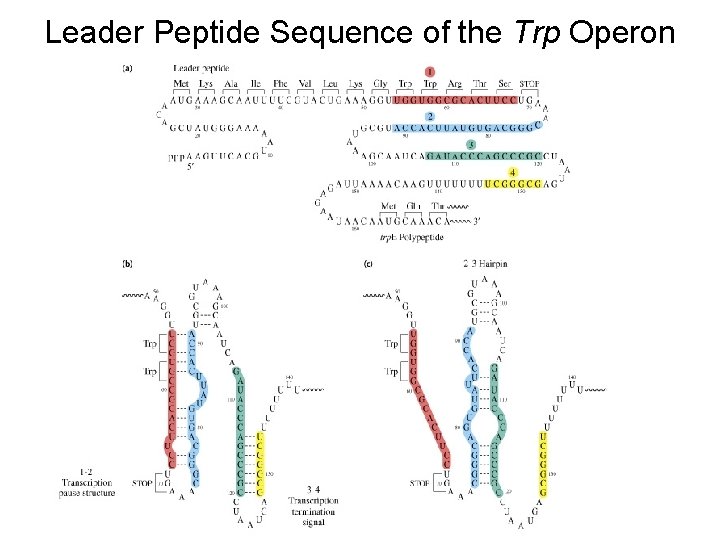
Leader Peptide Sequence of the Trp Operon

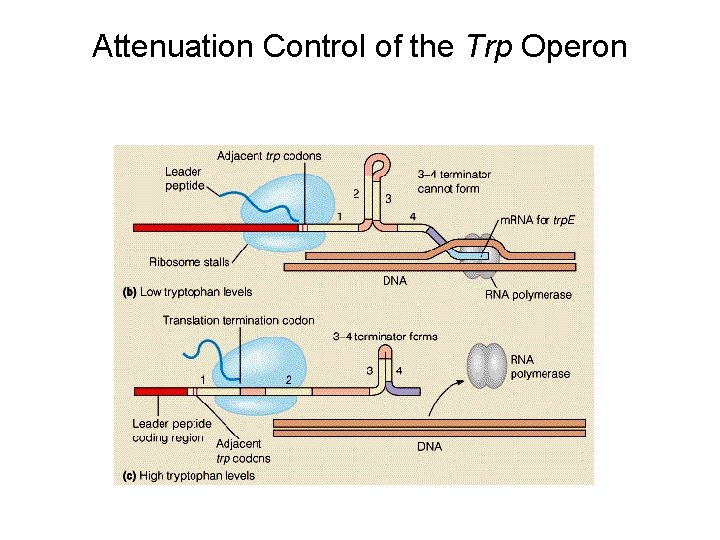
Attenuation Control of the Trp Operon

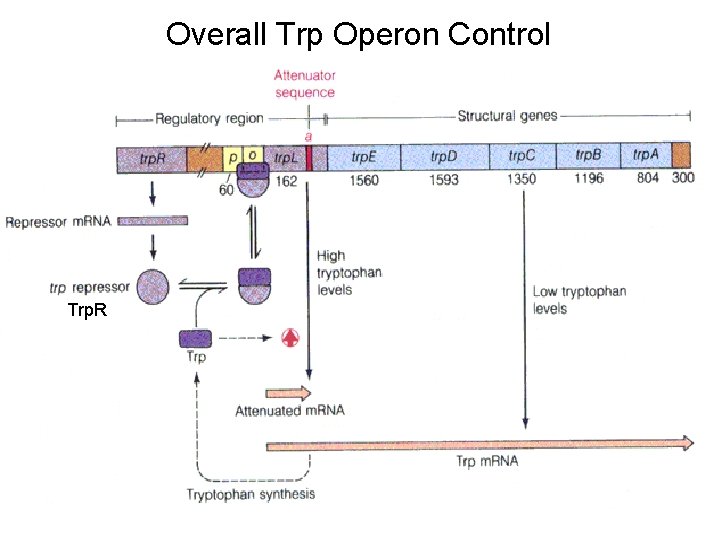
Overall Trp Operon Control Trp. R

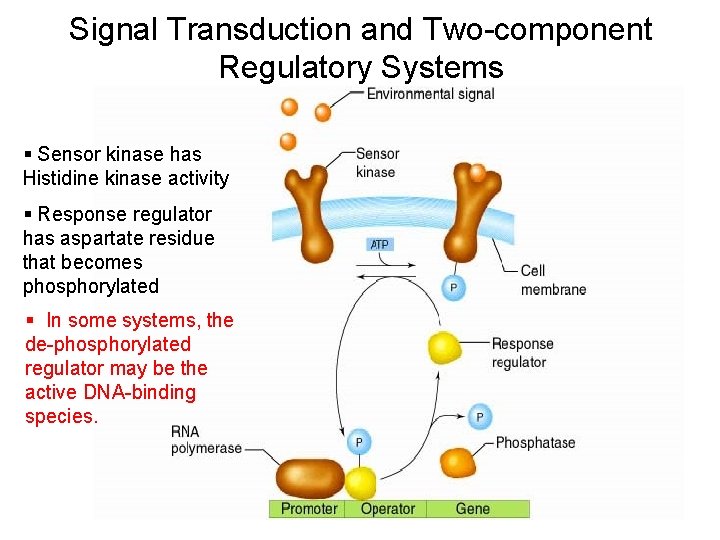
Signal Transduction and Two-component Regulatory Systems § Sensor kinase has Histidine kinase activity § Response regulator has aspartate residue that becomes phosphorylated § In some systems, the de-phosphorylated regulator may be the active DNA-binding species.

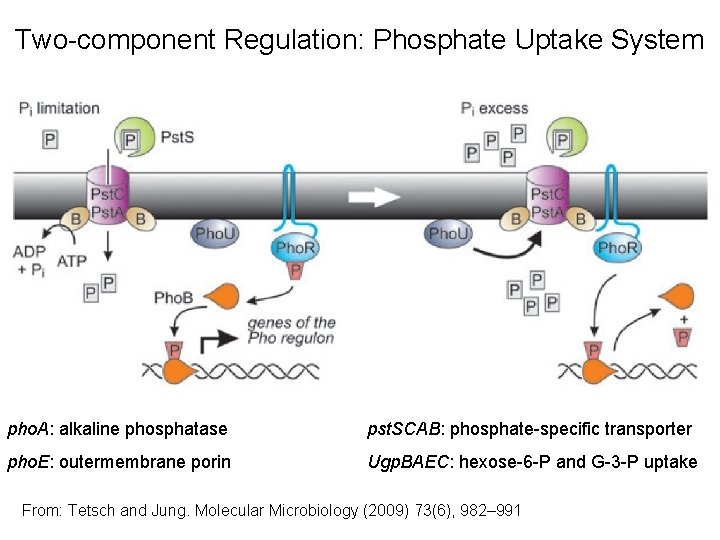
Two-component Regulation: Phosphate Uptake System pho. A: alkaline phosphatase pst. SCAB: phosphate-specific transporter pho. E: outermembrane porin Ugp. BAEC: hexose-6 -P and G-3 -P uptake From: Tetsch and Jung. Molecular Microbiology (2009) 73(6), 982– 991

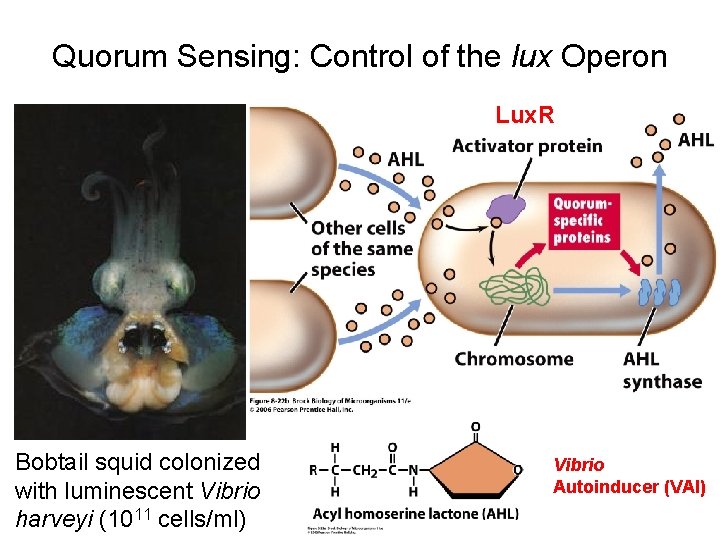
Quorum Sensing: Control of the lux Operon Lux. R Bobtail squid colonized with luminescent Vibrio harveyi (1011 cells/ml) Vibrio Autoinducer (VAI)

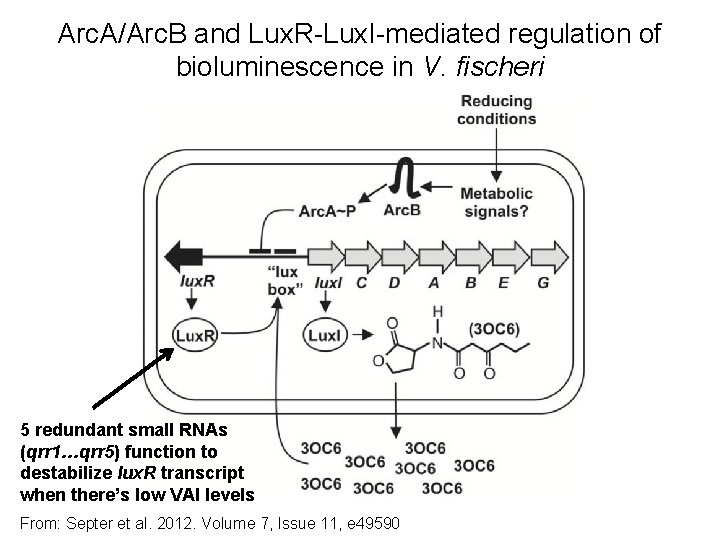
Arc. A/Arc. B and Lux. R-Lux. I-mediated regulation of bioluminescence in V. fischeri 5 redundant small RNAs (qrr 1…qrr 5) function to destabilize lux. R transcript when there’s low VAI levels From: Septer et al. 2012. Volume 7, Issue 11, e 49590

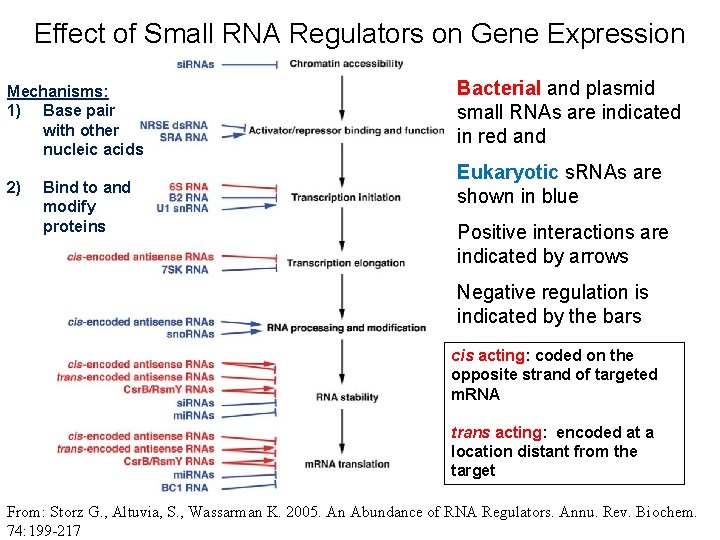
Effect of Small RNA Regulators on Gene Expression Mechanisms: 1) Base pair with other nucleic acids 2) Bind to and modify proteins Bacterial and plasmid small RNAs are indicated in red and Eukaryotic s. RNAs are shown in blue Positive interactions are indicated by arrows Negative regulation is indicated by the bars cis acting: coded on the opposite strand of targeted m. RNA trans acting: encoded at a location distant from the target From: Storz G. , Altuvia, S. , Wassarman K. 2005. An Abundance of RNA Regulators. Annu. Rev. Biochem. 74: 199 -217

Advantages of small RNAs over protein Regulators 1) Small RNAs require fewer resources to make than a (Cheaper) regulatory protein. Many small RNAs are expressed when carbon and energy are limiting. 2) Small RNAs act at the posttranscriptional level allowing a fast regulatory response. (Faster) 3) cis-encoded RNA regulators evolve with the target (Adapting)

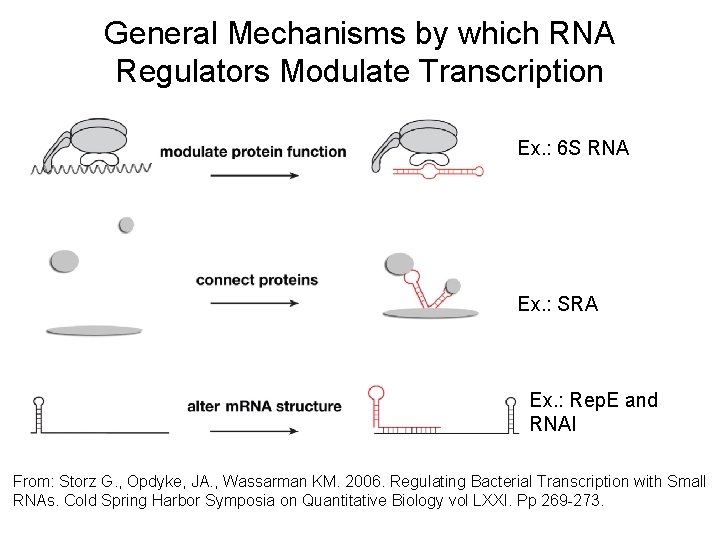
General Mechanisms by which RNA Regulators Modulate Transcription Ex. : 6 S RNA Ex. : SRA Ex. : Rep. E and RNAI From: Storz G. , Opdyke, JA. , Wassarman KM. 2006. Regulating Bacterial Transcription with Small RNAs. Cold Spring Harbor Symposia on Quantitative Biology vol LXXI. Pp 269 -273.

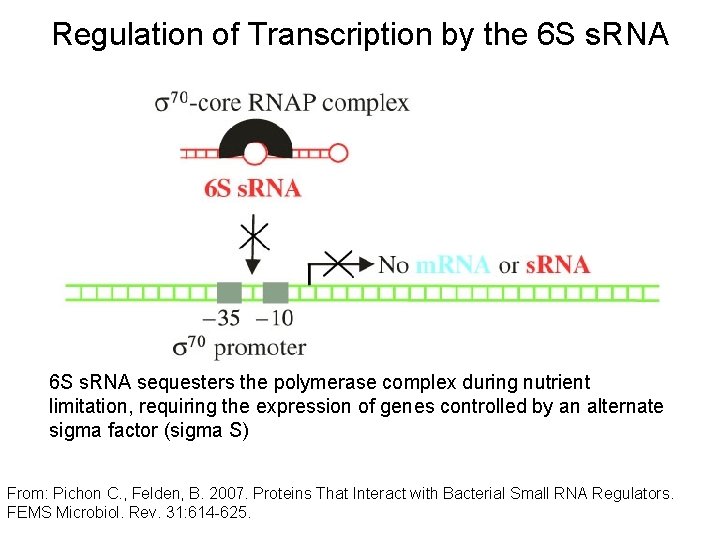
Regulation of Transcription by the 6 S s. RNA sequesters the polymerase complex during nutrient limitation, requiring the expression of genes controlled by an alternate sigma factor (sigma S) From: Pichon C. , Felden, B. 2007. Proteins That Interact with Bacterial Small RNA Regulators. FEMS Microbiol. Rev. 31: 614 -625.

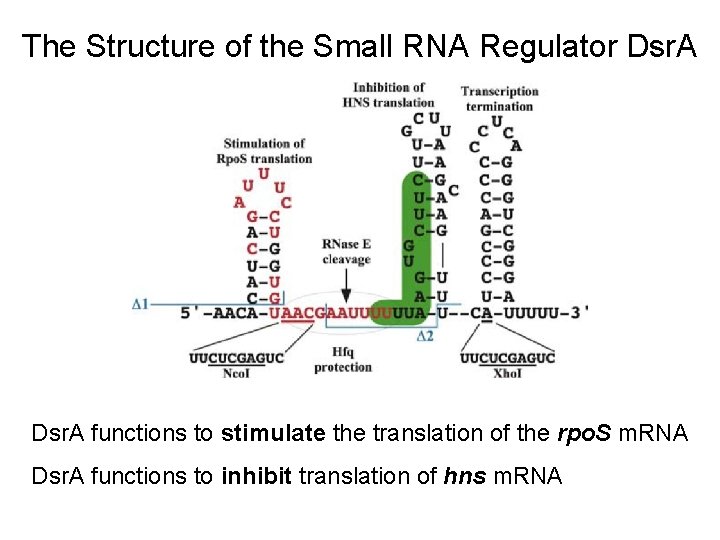
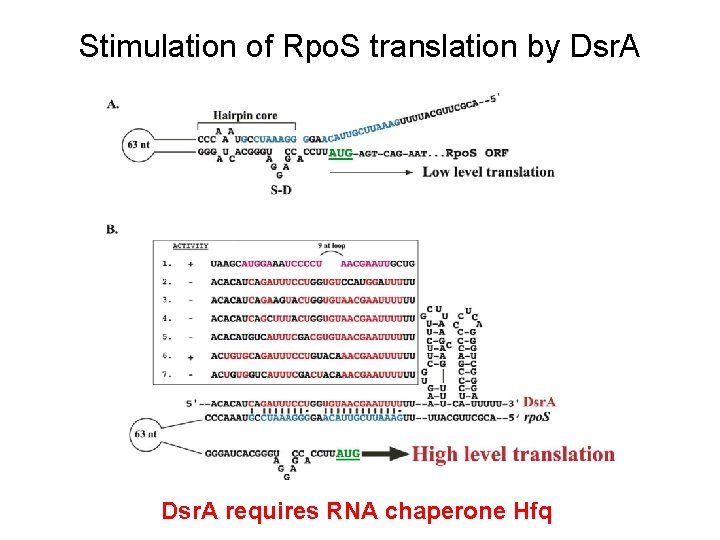
The Structure of the Small RNA Regulator Dsr. A functions to stimulate the translation of the rpo. S m. RNA Dsr. A functions to inhibit translation of hns m. RNA

Stimulation of Rpo. S translation by Dsr. A requires RNA chaperone Hfq

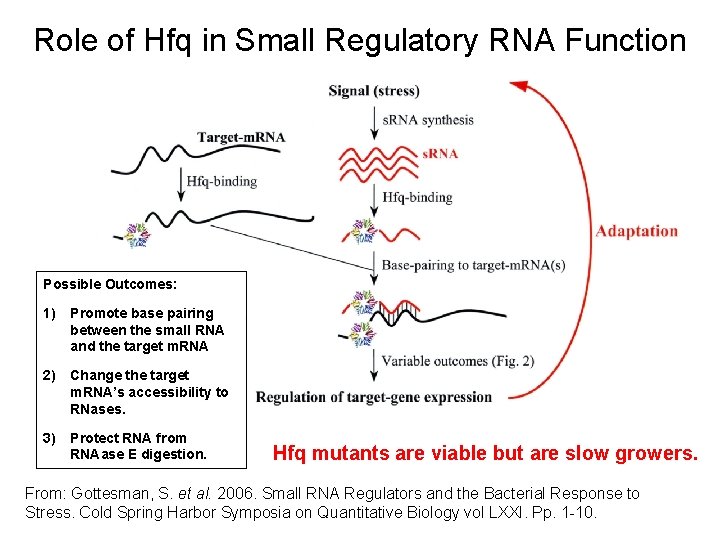
Role of Hfq in Small Regulatory RNA Function Possible Outcomes: 1) Promote base pairing between the small RNA and the target m. RNA 2) Change the target m. RNA’s accessibility to RNases. 3) Protect RNA from RNAase E digestion. Hfq mutants are viable but are slow growers. From: Gottesman, S. et al. 2006. Small RNA Regulators and the Bacterial Response to Stress. Cold Spring Harbor Symposia on Quantitative Biology vol LXXI. Pp. 1 -10.

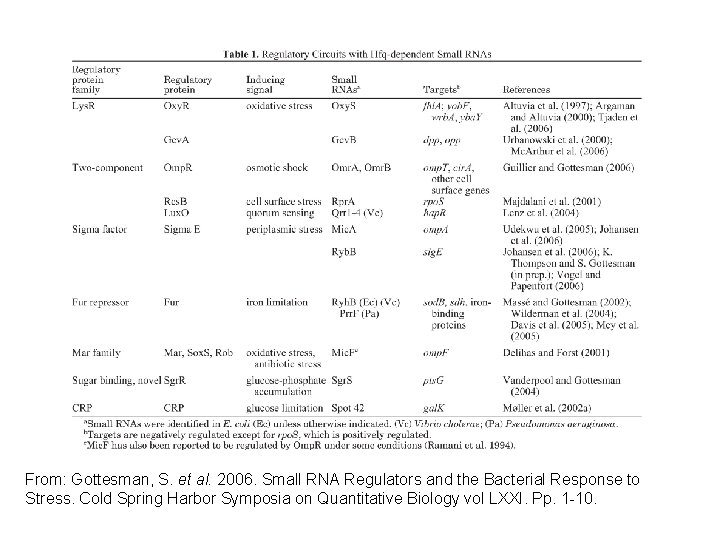
From: Gottesman, S. et al. 2006. Small RNA Regulators and the Bacterial Response to Stress. Cold Spring Harbor Symposia on Quantitative Biology vol LXXI. Pp. 1 -10.

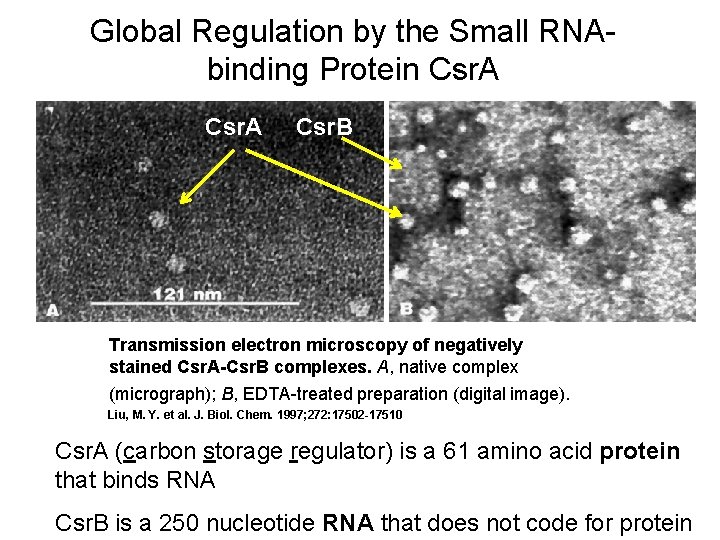
Global Regulation by the Small RNAbinding Protein Csr. A Csr. B Transmission electron microscopy of negatively stained Csr. A-Csr. B complexes. A, native complex (micrograph); B, EDTA-treated preparation (digital image). Liu, M. Y. et al. J. Biol. Chem. 1997; 272: 17502 -17510 Csr. A (carbon storage regulator) is a 61 amino acid protein that binds RNA Csr. B is a 250 nucleotide RNA that does not code for protein

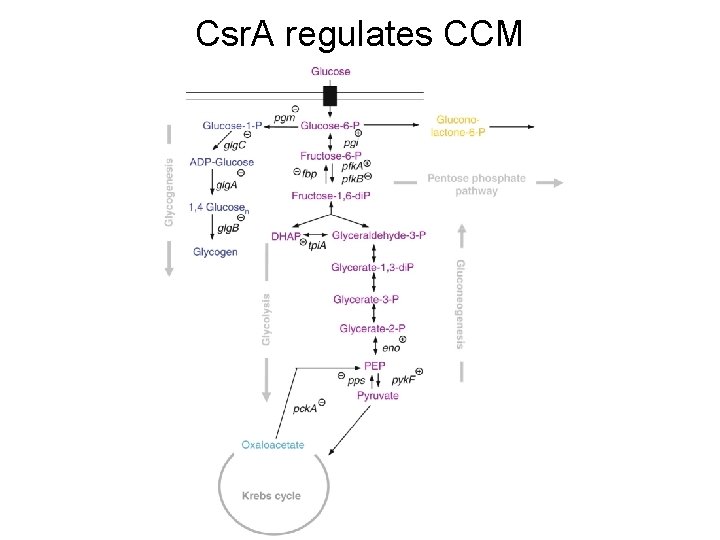
Expression of genes affected by Csr. A (E. coli) Pathways Neg. Regulated by Csr. A: § Glycogen biosynthesis § Glycogen catabolism § Gluconeogenesis Pathways Pos. Regulated by Csr. A: § Glycolysis § Glyoxylate shunt § Acetate metabolism § Flagella biosynthesis

Csr. A regulates CCM

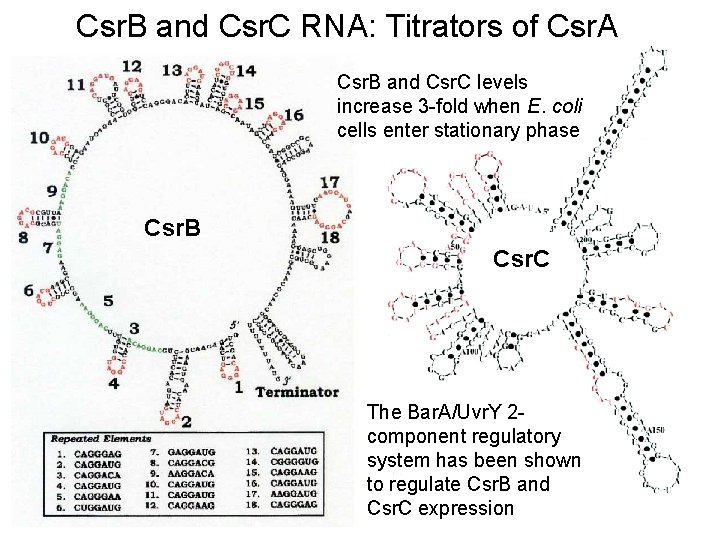
Csr. B and Csr. C RNA: Titrators of Csr. A Csr. B and Csr. C levels increase 3 -fold when E. coli cells enter stationary phase Csr. B Csr. C The Bar. A/Uvr. Y 2 component regulatory system has been shown to regulate Csr. B and Csr. C expression

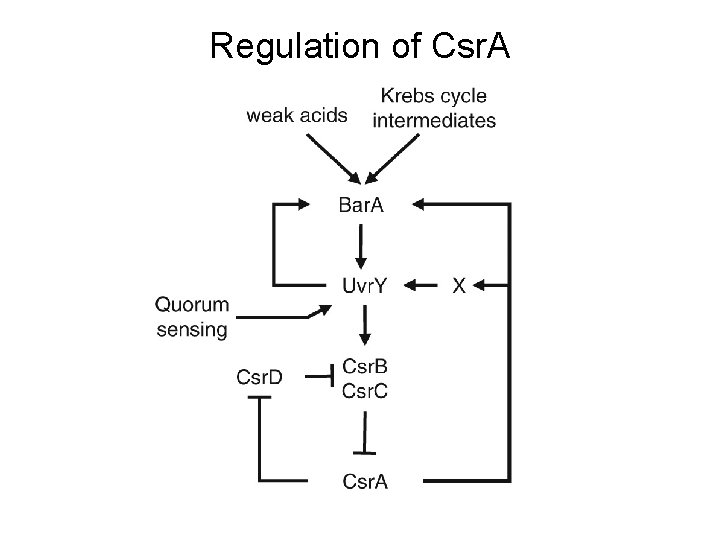
Regulation of Csr. A

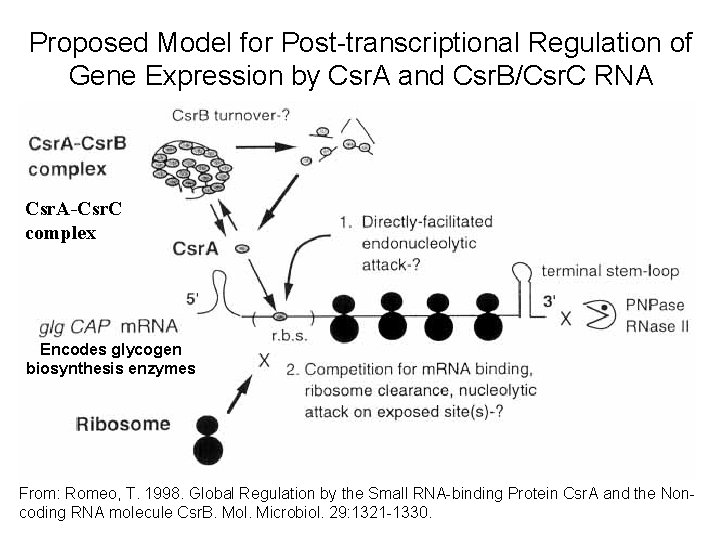
Proposed Model for Post-transcriptional Regulation of Gene Expression by Csr. A and Csr. B/Csr. C RNA Csr. A-Csr. C complex Encodes glycogen biosynthesis enzymes From: Romeo, T. 1998. Global Regulation by the Small RNA-binding Protein Csr. A and the Noncoding RNA molecule Csr. B. Mol. Microbiol. 29: 1321 -1330.

Proposed Model for Post-transcriptional Regulation of Gene Expression by Csr. A and Csr. B/Csr. C RNA Benefits of this Regulatory Strategy (1) There is a low energy investment and short synthesis time to produce an RNA regulator. (2) A single RNA can bind 18 regulator proteins, thereby amplifying its control (3) Post-transcriptional control results in almost immediate changes in gene expression.

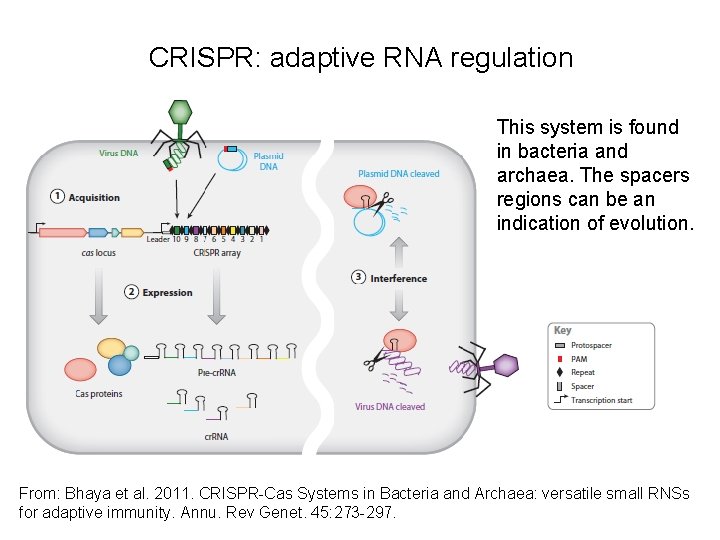
CRISPR: adaptive RNA regulation This system is found in bacteria and archaea. The spacers regions can be an indication of evolution. From: Bhaya et al. 2011. CRISPR-Cas Systems in Bacteria and Archaea: versatile small RNSs for adaptive immunity. Annu. Rev Genet. 45: 273 -297.

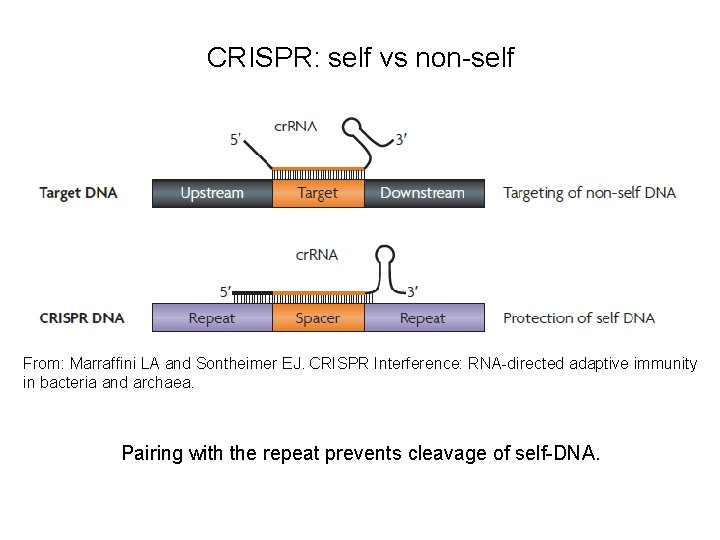
CRISPR: self vs non-self From: Marraffini LA and Sontheimer EJ. CRISPR Interference: RNA-directed adaptive immunity in bacteria and archaea. Pairing with the repeat prevents cleavage of self-DNA.

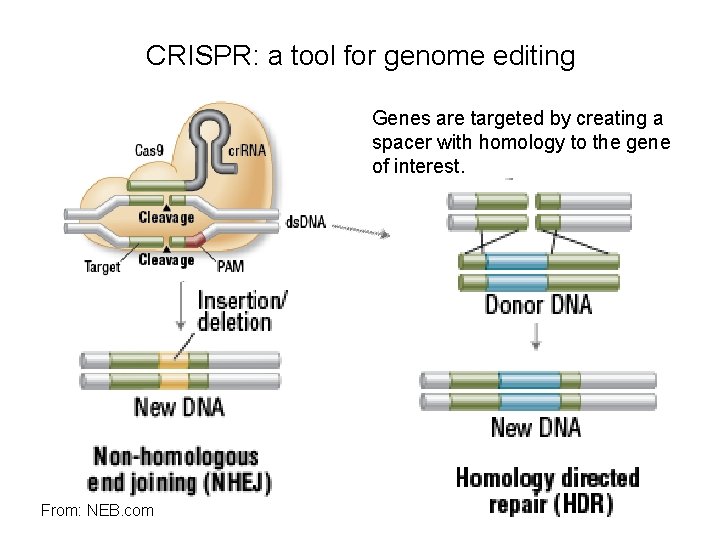
CRISPR: a tool for genome editing Genes are targeted by creating a spacer with homology to the gene of interest. From: NEB. com