Lecture 1 DNA damage Damage Reversal Base excision

Lecture 1 DNA damage. Damage Reversal. Base excision repair. Mismatch repair Lecture 2 Nucleotide excision repair: cellular and clinical aspects Nucleotide excision repair: genes and proteins Lecture 3 Replication of damaged DNA. Mutagenesis and carcinogenesis

Course Learning objectives • To gain an understanding of the molecular mechanisms that maintain genome stability • To appreciate the importance of this topic for human health. Learning outcomes (Lecture 1 a) Understanding: • Different types of DNA damage • Three examples of ways in which cells can reverse damage in situ • Basic mechanism of Base Excision Repair
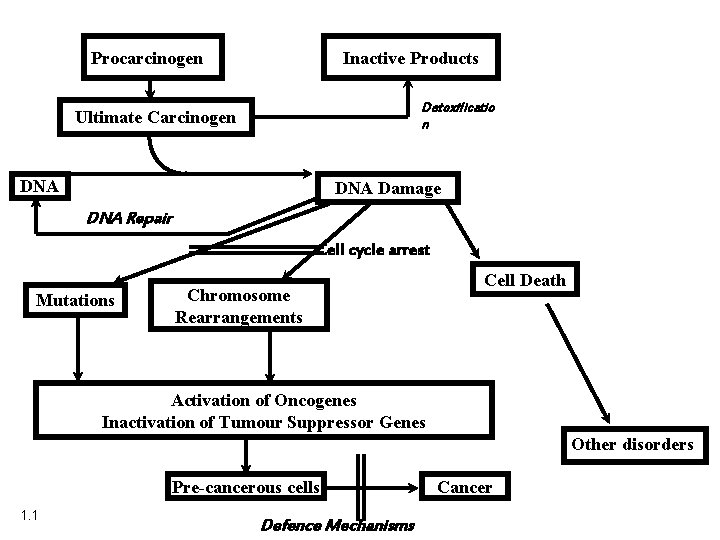
Procarcinogen Inactive Products Detoxificatio n Ultimate Carcinogen DNA Damage DNA Repair Cell cycle arrest Mutations Chromosome Rearrangements Cell Death Activation of Oncogenes Inactivation of Tumour Suppressor Genes Other disorders Pre-cancerous cells 1. 1 Defence Mechanisms Cancer
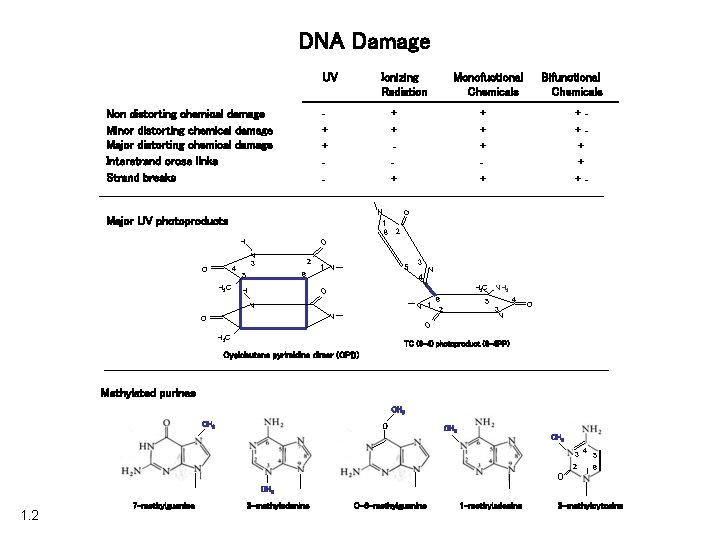
DNA Damage UV Ionizing Radiation + + - Non distorting chemical damage Minor distorting chemical damage Major distorting chemical damage Interstrand cross links Strand breaks + +++ + +- 2 N 3 2 6 5 1 N 5 3 N 4 H 3 C NH 2 O H 3 C + + O H 4 Bifunctional Chemicals O N 1 6 Major UV photoproducts O Monofuctional Chemicals H 6 N 1 2 N N 4 3 N O O O 5 H 3 C TC (6 -4) photoproduct (6 -4 PP) Cyclobutane pyrimidine dimer (CPD) Methylated purines CH 3 O 3 4 5 2 1 6 CH 3 1. 2 7 -methylguanine 3 -methyladenine O-6 -methylguanine 1 -methyladenine 3 -methylcytosine

Aspects of DNA repair 1. Initial damage 2. Repair of damage 3. Genes involved 4. Mechanism of action of gene products 5. Replication of unremoved damage. Cell cycle progression. 6. Biological consequences of damage, repair and failure to repair. 1. 3
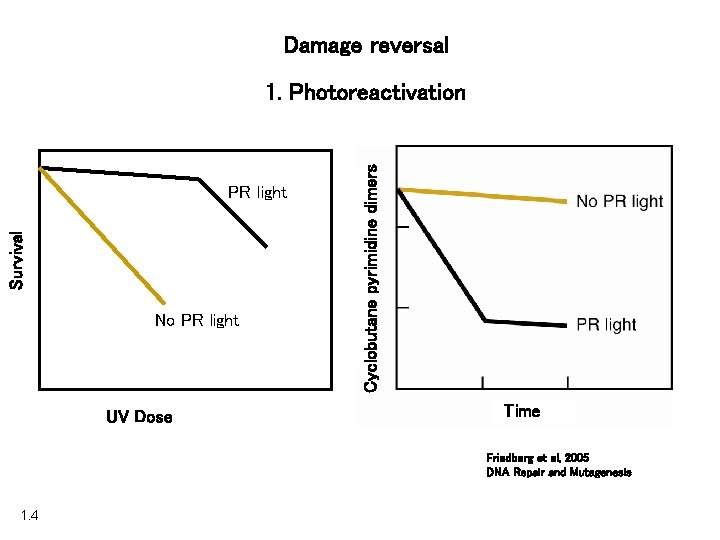
Damage reversal Survival PR light No PR light UV Dose Cyclobutane pyrimidine dimers 1. Photoreactivation Time Friedberg et al, 2005 DNA Repair and Mutagenesis 1. 4
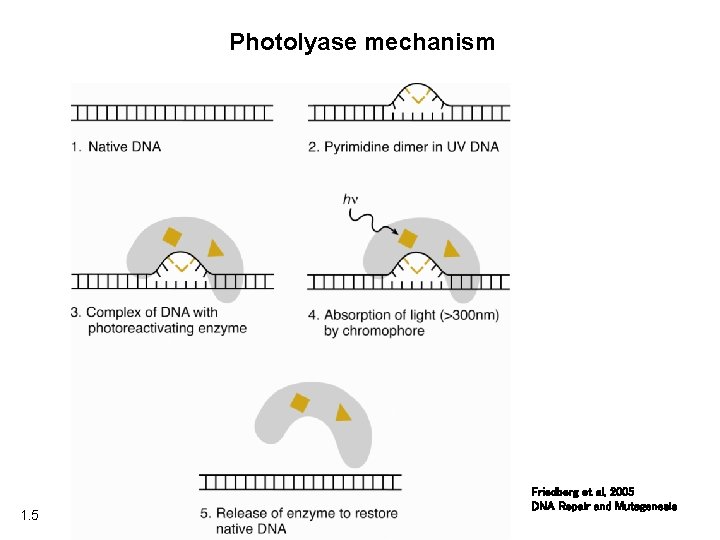
Photolyase mechanism 1. 5 Friedberg et al, 2005 DNA Repair and Mutagenesis
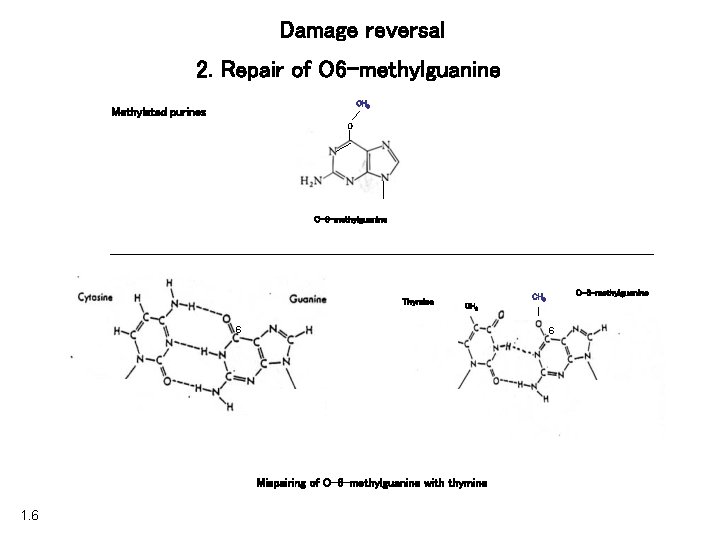
Damage reversal 2. Repair of O 6 -methylguanine CH 3 Methylated purines O O-6 -methylguanine Thymine CH 3 6 6 Mispairing of O-6 -methylguanine with thymine 1. 6 O-6 -methylguanine CH 3

Damage reversal 2. Repair of O 6 -methylguanine Adaptation Treat E. coli with indicated dose of MNNG, then expose to high dose Control Adapted Friedberg et al, 2005 DNA Repair and Mutagenesis 1. 7

Dual activities of Ada methyltransferase Friedberg et al, 2005 DNA Repair and Mutagenesis 1. 8
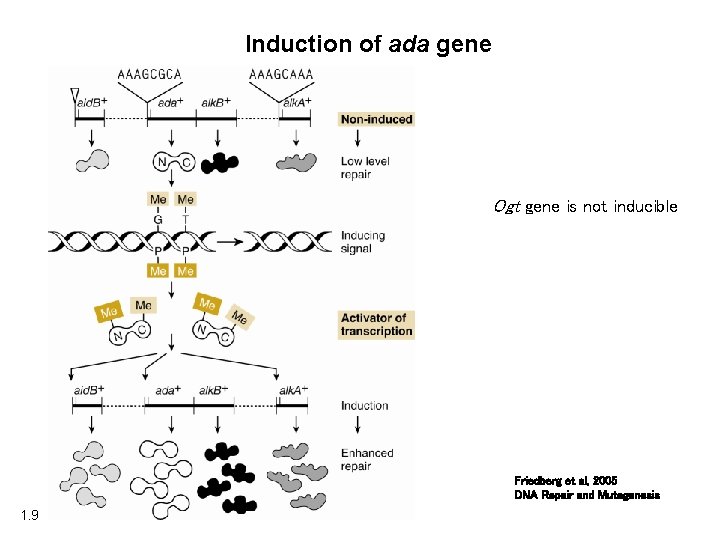
Induction of ada gene Ogt gene is not inducible Friedberg et al, 2005 DNA Repair and Mutagenesis 1. 9

Alkyltransferases in mammalian cells • Similar mechanism to E. coli, but for O-6 -me. G alone, like Ogt, not inducible. • K/o mouse constructed, very sensitive to carcinogenesis by methylating agents. • Conversely transgenic mice bearing MGMT gene are more resistant. • Many cancer cell lines are Mex-. MGMT silenced by methylation in about 50% of tumours. • Mex- cells are sensitive to killing and mutagenesis by alkylating agents. • Many cancer therapy drugs are alkylating agents, eg temozolomide. • Patrin 2 binds MGMT and depletes it. Currently in clinical trials together with temozolomide. 1. 10
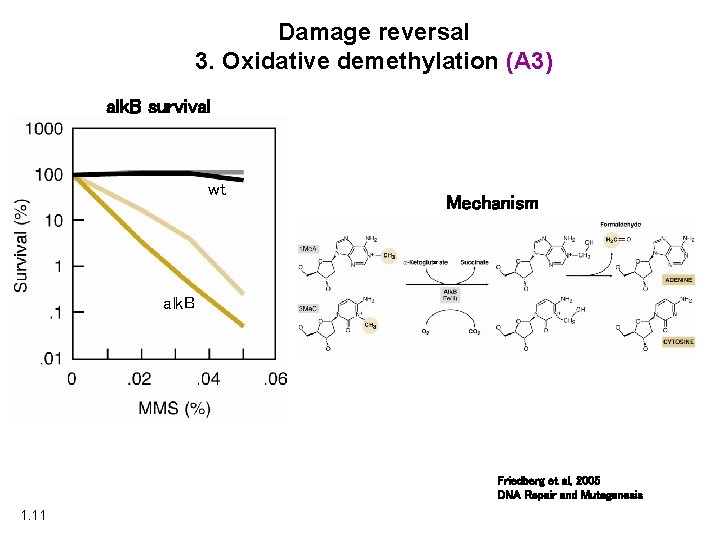
Damage reversal 3. Oxidative demethylation (A 3) alk. B survival wt Mechanism ar alk. B Friedberg et al, 2005 DNA Repair and Mutagenesis 1. 11

Base Excision Repair

Deamination of bases 8 -hydroxyguanine
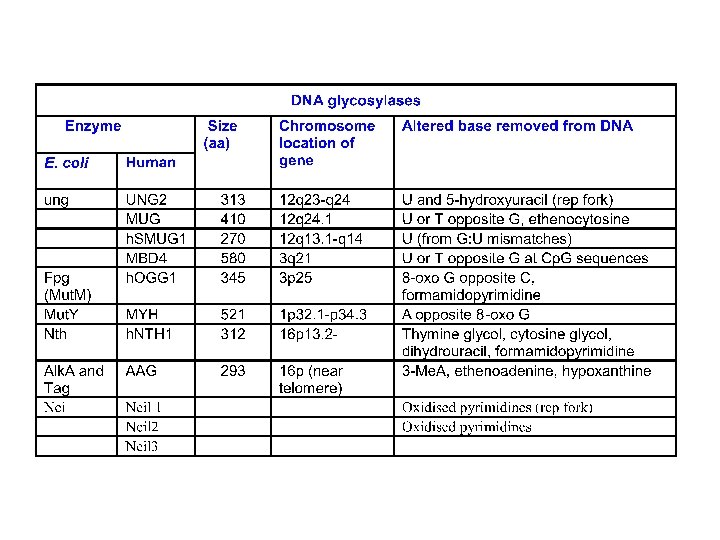
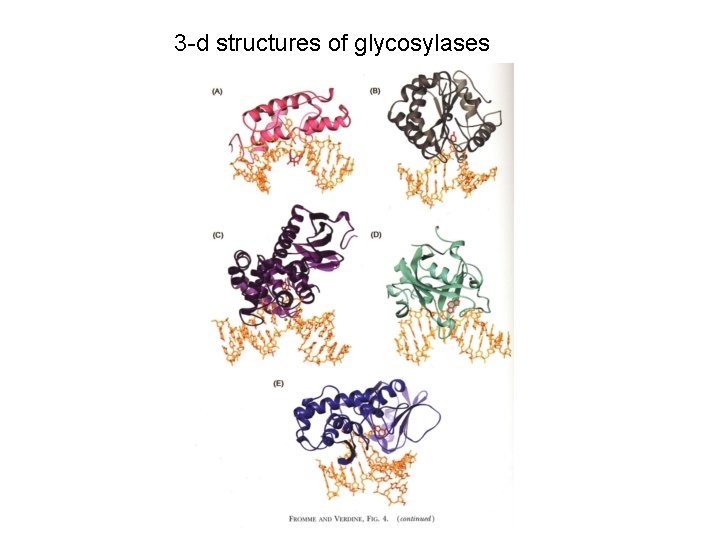
3 -d structures of glycosylases
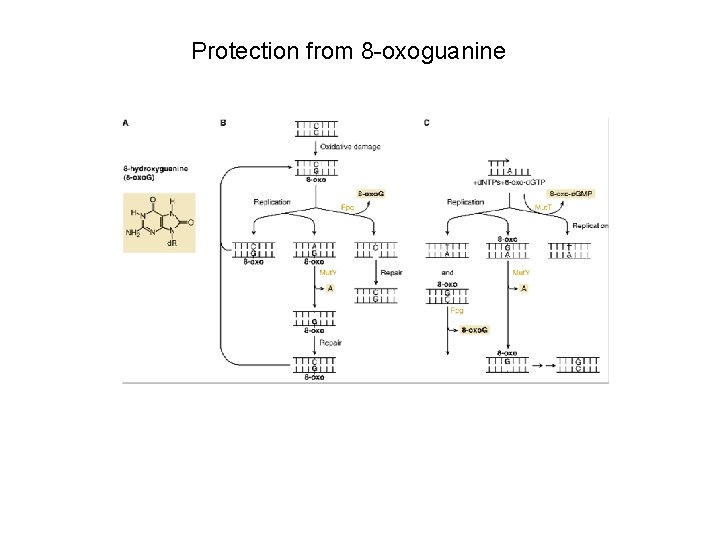
Protection from 8 -oxoguanine
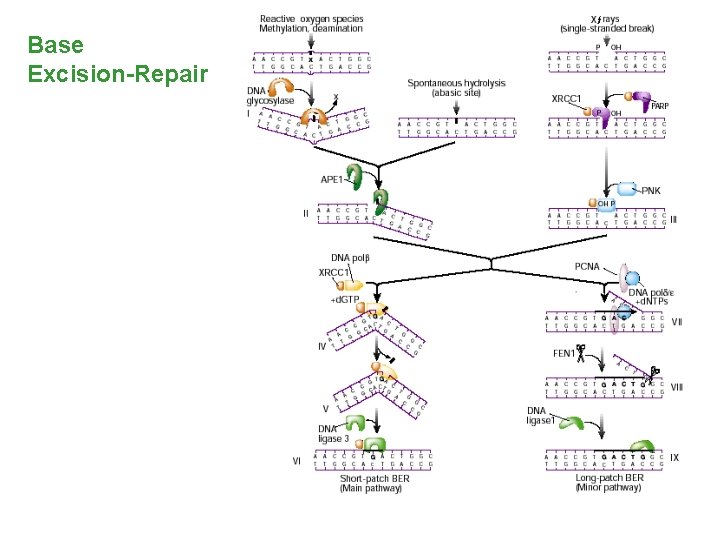
Base Excision-Repair
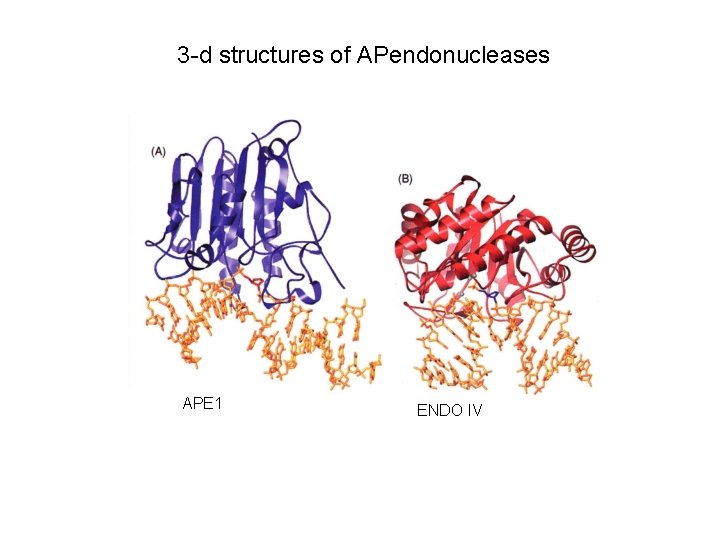
3 -d structures of APendonucleases APE 1 ENDO IV

Summary (Lecture 1 a) • DNA damage can cause distortions of different severity • UV damage is repaired by photoreversal (not in placental mammals) • O 6 -methylguanine is repaired by a specific methyltransferase • 1 -methyladenine and 3 -methylcytosine are repaired by oxidative demethylation • Spontaneous lesions are removed by Base Excision Repair

Learning outcomes (Lecture 1 b) Understanding: • Detailed mechanism of mismatch repair in E. coli and eukaryotes • How mismatch repair is important both for cancer protection and cancer therapy

Mismatch Repair (A 5, A 6) • DNA polymerases replicate DNA very faithfully Accurate insertion Associated 3’-5’ exonuclease for proof-reading Error rates c. 10 -6 or less • But genomes are big: E. coli 3 x 106 bp, mammals 3 x 109 • Errors can be single base mismatches or small insertions or deletions caused by base slippage • Mismatches are repaired by the MMR system which recognises the mismatched bases • But there’s a problem 1. 12
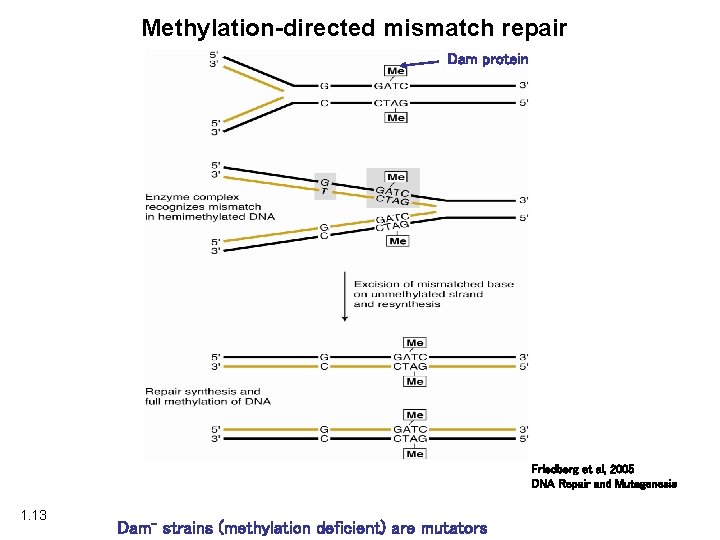
Methylation-directed mismatch repair Dam protein Friedberg et al, 2005 DNA Repair and Mutagenesis 1. 13 Dam- strains (methylation deficient) are mutators
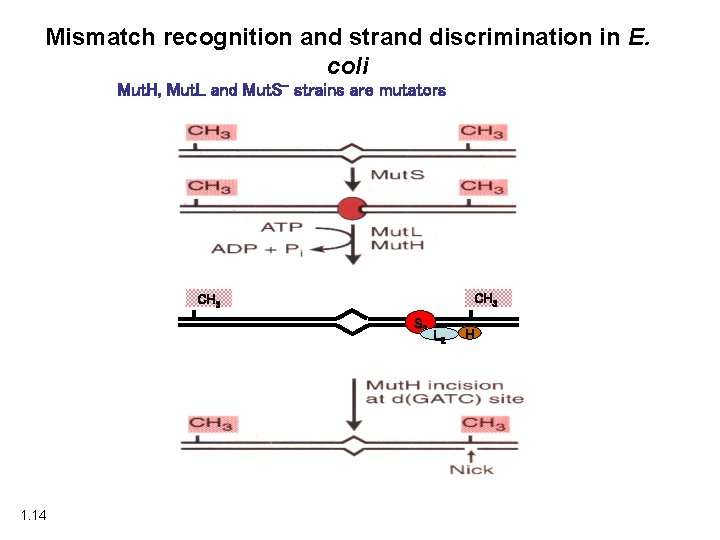
Mismatch recognition and strand discrimination in E. coli Mut. H, Mut. L and Mut. S- strains are mutators CH 3 S 2 1. 14 L 2 H
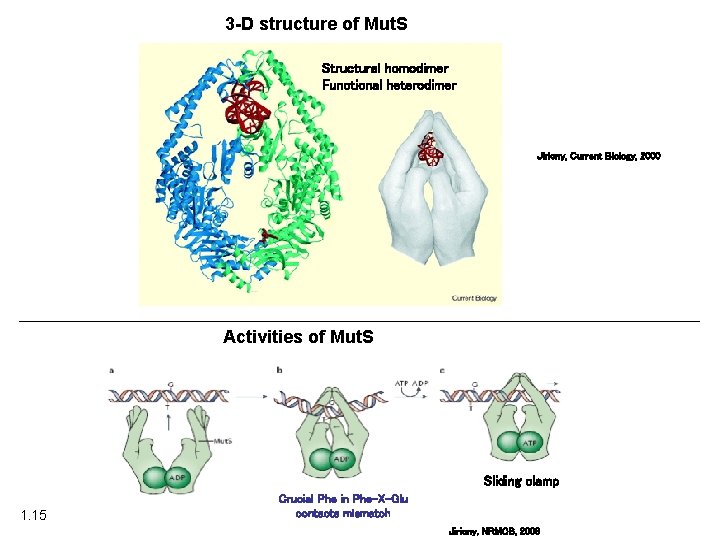
3 -D structure of Mut. S Structural homodimer Functional heterodimer Jiricny, Current Biology, 2000 Activities of Mut. S Sliding clamp 1. 15 Crucial Phe in Phe-X-Glu contacts mismatch Jiricny, NRMCB, 2006
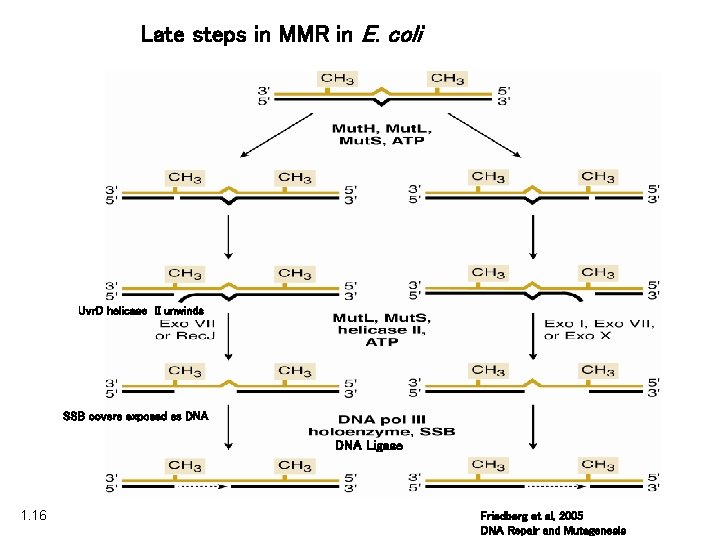
Late steps in MMR in E. coli Uvr. D helicase II unwinds SSB covers exposed ss DNA Ligase 1. 16 Friedberg et al, 2005 DNA Repair and Mutagenesis

Eukaryotic homologues of Mut. H, L, S Mut. L: Mut. S: Msh 2 Msh 3 Msh 4 Msh 5 Msh 6 MMR Meiosis MMR Mlh 1 Mlh 2 Mlh 3 Pms 1 Pms 2 MMR ? MMR (= Pms 1 in yeast) Mut. H: No homologues Neither yeast nor Drosophila has methylated DNA Strand discrimination based on nicks/ends in daughter DNA MMR proteins interact with PCNA at replication fork 1. 17
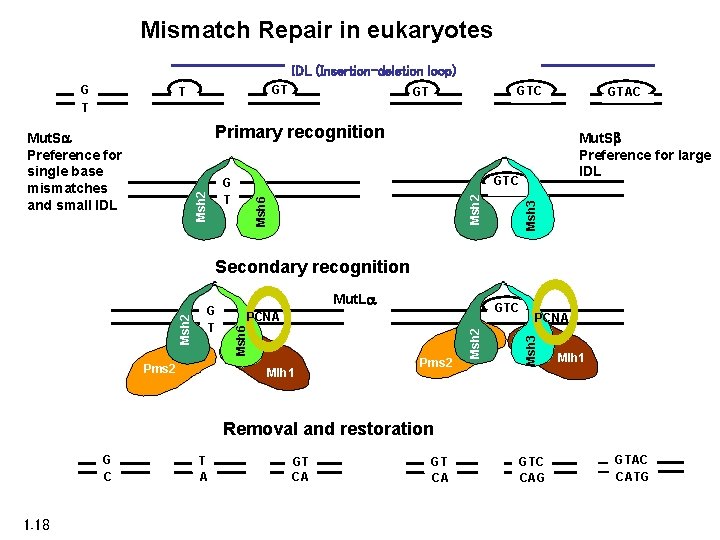
Mismatch Repair in eukaryotes IDL (Insertion-deletion loop) G T GTC GT GTAC Primary recognition Mut. Sb Preference for large IDL Msh 3 Msh 6 Msh 2 GTC G T Msh 2 Mut. Sa Preference for single base mismatches and small IDL GTC PCNA Mlh 1 Pms 2 PCNA Msh 3 Pms 2 Mut. La Msh 2 G T Msh 6 Msh 2 Secondary recognition Mlh 1 Removal and restoration G C 1. 18 T A GT CA GTC CAG GTAC CATG

Mismatch Repair in eukaryotes PCNA 3’ 5’ 5’ Mut. Sa or b binds mismatch Mut. Sa/Mut. La translocates to end 3’ (Exo. I) Leading strand: Mut. La (Pms 2 subunit) cleaves on 5’ side of mismatch. Exo 1 degrades 5’ to 3’ Lagging strand: Exo 1 degrades 5’ to 3’ 1. 19 Modified from Jiricny, Nature Rev Mol Cell Biol 2006 (A 5)

Microsatellite instability in tumour tissue from HNPCC (Hereditary non-polyposis colon carcinoma) Lynch Syndrome Aaltonen, et al. Science 260, 812 (1993) Primer 1 7 microsatellite repeats CA CA GT GT Replication Slippage 1. 20 CA CA GT GT GT Primer 2 6 repeats
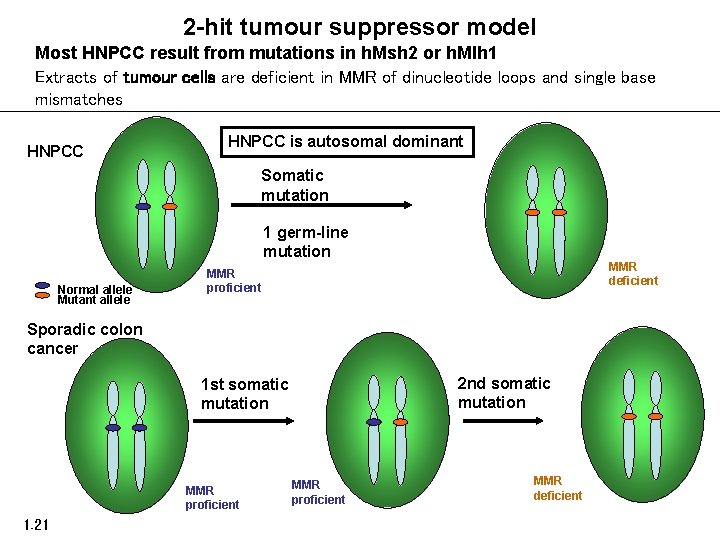
2 -hit tumour suppressor model Most HNPCC result from mutations in h. Msh 2 or h. Mlh 1 Extracts of tumour cells are deficient in MMR of dinucleotide loops and single base mismatches HNPCC is autosomal dominant Somatic mutation 1 germ-line mutation Normal allele Mutant allele MMR deficient MMR proficient Sporadic colon cancer 2 nd somatic mutation 1 st somatic mutation MMR proficient 1. 21 MMR proficient MMR deficient

Microsatellite instability results from loss of Mismatch Repair 5' T-G-T-G-T- G 3' A-C-A-C-A-C-5' T-G misalignment | | 5' T-G T- G-T- G 3' A-C-A-C-A-C extension T-G | | 5' T- G T-G-T- G 3' A-C-A-C-A-C-A-C reversal (proofreading) 5' T-G-T- G 3' A-C-A-C-A-C-A-C mismatch repair 5' T-G-T-G-T- G 3' A-C-A-C-A-C • • 1. 22 Microsatellite instability is a useful diagnostic tool. It’s not the cause of the cancers Cancers arise from high rate of single-base mismatches during replication These lead to high frequency of somatic mutations Why only in colon? Not known
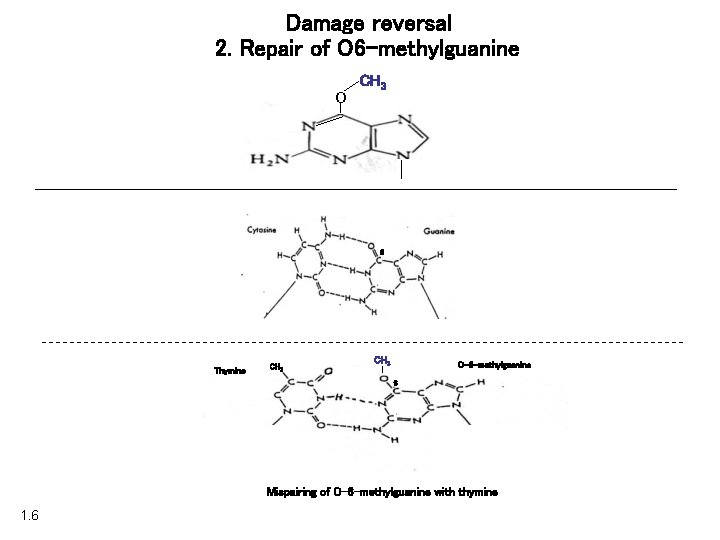
Damage reversal 2. Repair of O 6 -methylguanine O CH 3 6 Thymine CH 3 O-6 -methylguanine 6 Mispairing of O-6 -methylguanine with thymine 1. 6

MMR and resistant tumours • In many tumour cells MGMT is silenced. So alkylating agents are good for therapy. • But often develop resistance. • Select for alkylation-resistance in cells. • MGMT not restored. O 6 -Me. G remains in DNA. Instead cells have lost one of the MMR genes. • Implies MMR somehow sensitises cells to alkylation damage. • Result of futile cycles. O 6 -Me. G: C and O 6 -Me. G: T both recognised as mismatches. • C or T opposite O 6 -Me. G removed by MMR and replaced with C or T. Futile cycles. • Results in cell cycle arrest or apoptosis 1. 23

Loss of MMR protects against MNNG apoptosis Friedberg et al, 2005 DNA Repair and Mutagenesis MMR and cancer MMR deficiency • Increases cancer susceptibility (HNPCC) • Results in resistance to cancer therapy 1. 24

Summary (Lecture 1 b) • Mismatches are repaired by the Mut(H), L, S system • Mismatches are recognised by Mut. S and its homologues • Strand discrimination is brought about by methylation in E. coli and nicks/ends in daughter strands in eukaryotes • MMR deficiency leads to HNPCC and is detected by microsatellite instability • Loss of MMR results in resistance to alkylating agents
- Slides: 37