Learning recursive Bayesian multinets for data clustering by

Learning recursive Bayesian multinets for data clustering by means of constructive induction Pegna, J. M. , Lozano, J. A. , and Larragnaga, P. Machine Learning, 47(1), pp. 63 -89, 2002. Summarized by Kyu-Baek Hwang (c) 2003 SNU CSE Biointelligence Laboratory

data clustering problem l data partitioning ¨ k-means l representing the joint probability distribution of a database ¨ mixture density models (e. g. , Gaussian mixtures) ¨ Bayesian networks C Y 1 Y 2 Y 3 Y 4 Y 5 (c) 2003 SNU CSE Biointelligence Laboratory 2
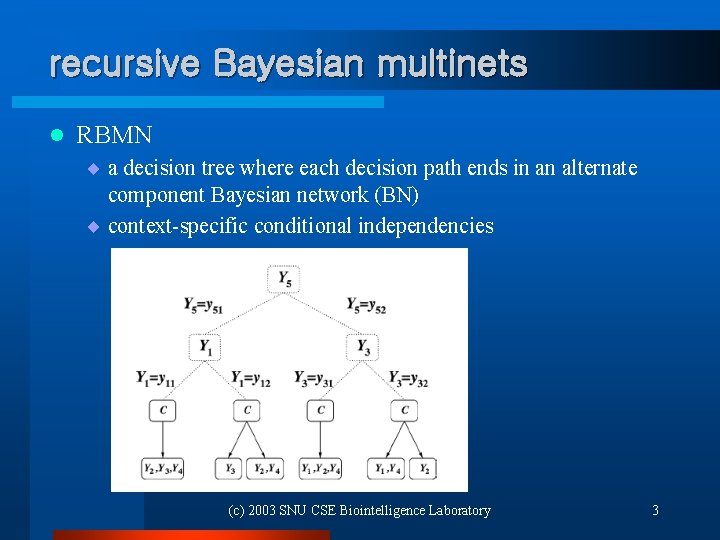
recursive Bayesian multinets l RBMN ¨ a decision tree where each decision path ends in an alternate component Bayesian network (BN) ¨ context-specific conditional independencies (c) 2003 SNU CSE Biointelligence Laboratory 3

BNs for data clustering l the joint probability distribution by a BN (c) 2003 SNU CSE Biointelligence Laboratory 4
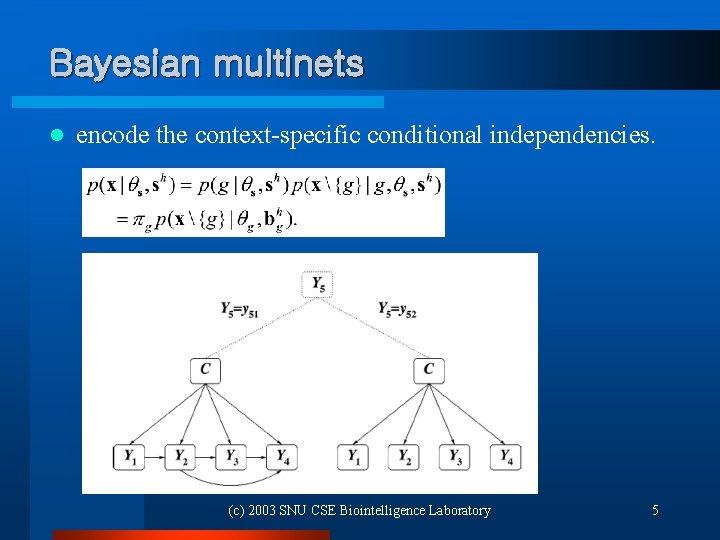
Bayesian multinets l encode the context-specific conditional independencies. (c) 2003 SNU CSE Biointelligence Laboratory 5
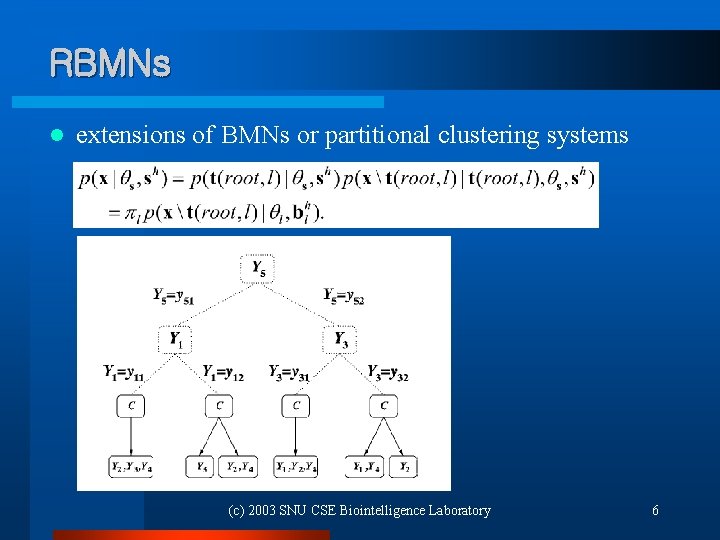
RBMNs l extensions of BMNs or partitional clustering systems (c) 2003 SNU CSE Biointelligence Laboratory 6
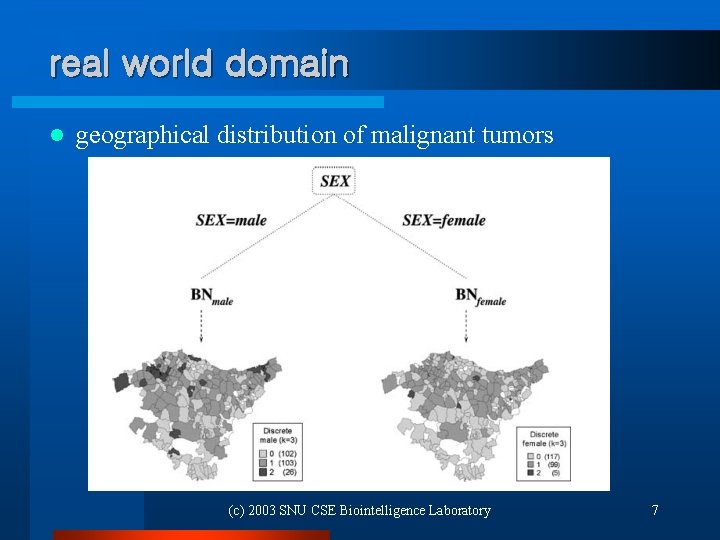
real world domain l geographical distribution of malignant tumors (c) 2003 SNU CSE Biointelligence Laboratory 7
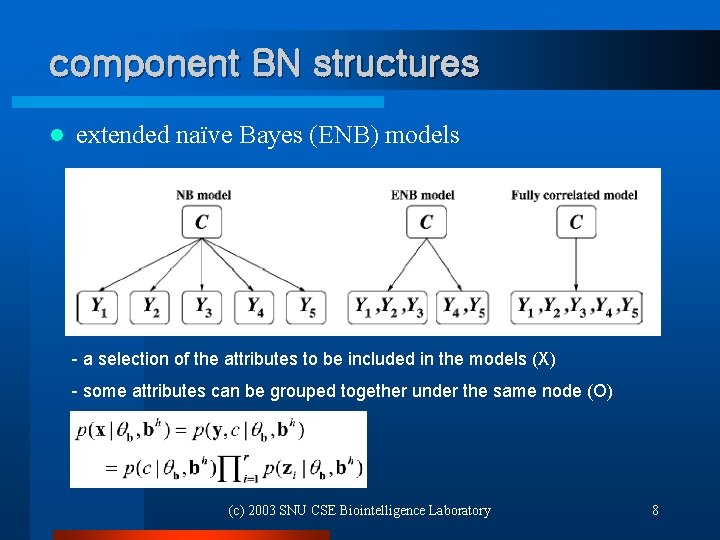
component BN structures l extended naïve Bayes (ENB) models - a selection of the attributes to be included in the models (X) - some attributes can be grouped together under the same node (O) (c) 2003 SNU CSE Biointelligence Laboratory 8

learning algorithm for ENB models (c) 2003 SNU CSE Biointelligence Laboratory 9

parameter search - EM (expectation maximization) algorithm - BC (bound and collapse) + EM algorithm (O) (c) 2003 SNU CSE Biointelligence Laboratory 10

structure search l constructive induction ¨ the process of changing the representation of the cases in the database by creating new attributes from existing attributes. ¨ forward algorithm ¨ backward algorithm (c) 2003 SNU CSE Biointelligence Laboratory 11

marginal likelihood criterion for RBMNs l with uninformative Dirichlet prior, l for BMNs, with some reasonable assumptions including parameter independence, l for RBMNs, (c) 2003 SNU CSE Biointelligence Laboratory 12

learning algorithm for RBMNs (c) 2003 SNU CSE Biointelligence Laboratory 13

experimental setup both synthetic data and real data l discrete variables with (unrestricted) multinomial distributions l convergence criterion for BC + EM algorithm l ¨ change in the log marginal likelihood value is less than 10 -6 ¨ or 150 iterations fixing_probability_threshold: 0. 51 l initial structure: naïve Bayes model l 5 independent runs at each experiment l (c) 2003 SNU CSE Biointelligence Laboratory 14

1 -level RBMNs for the experiments (c) 2003 SNU CSE Biointelligence Laboratory 15

2 -level RBMNs for the experiments (c) 2003 SNU CSE Biointelligence Laboratory 16
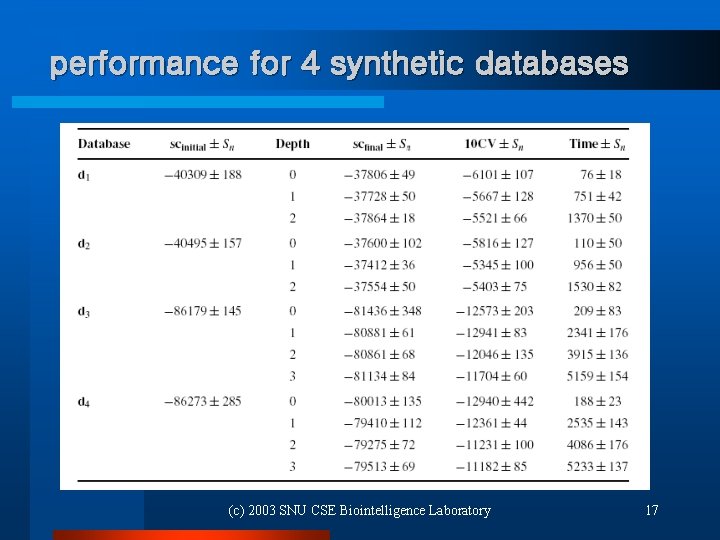
performance for 4 synthetic databases (c) 2003 SNU CSE Biointelligence Laboratory 17
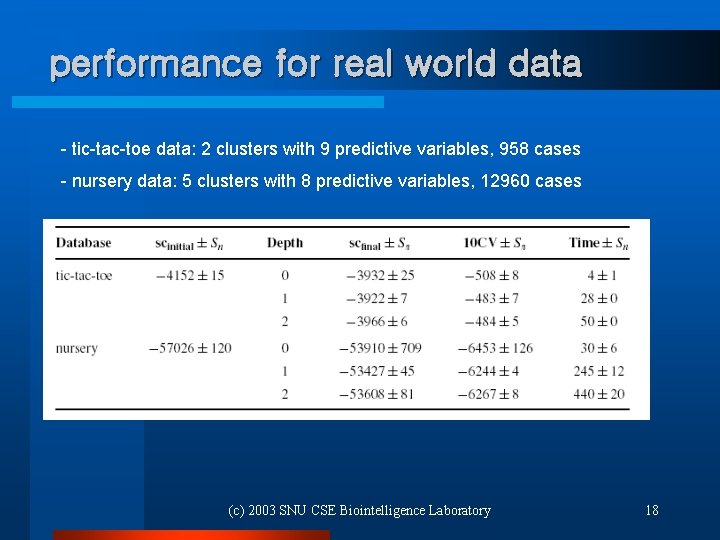
performance for real world data - tic-tac-toe data: 2 clusters with 9 predictive variables, 958 cases - nursery data: 5 clusters with 8 predictive variables, 12960 cases (c) 2003 SNU CSE Biointelligence Laboratory 18

conclusions and future research l context-specific conditional independencies ¨ data partitioning ¨ efficient representation, Bayesian committees, mixture of experts l learning speed problem ¨ trade-off with the efficient representation l monothetic decision tree ¨ polythetic paths enrich the modeling power l extensions to the continuous domain (c) 2003 SNU CSE Biointelligence Laboratory 19
- Slides: 19