Learning Ensembles of FirstOrder Clauses for RecallPrecision Curves

Learning Ensembles of First-Order Clauses for Recall-Precision Curves A Case Study in Biomedical Information Extraction Mark Goadrich, Louis Oliphant and Jude Shavlik Department of Computer Sciences University of Wisconsin – Madison USA 6 Sept 2004

Talk Outline Link Learning and ILP n Our Gleaner Approach n Aleph Ensembles n Biomedical Information Extraction n Evaluation and Results n Future Work n
ILP Domains n Object Learning n n Trains, Carcinogenesis Link Learning n Learning binary predicates

Link Learning n Large skew toward negatives n n n Difficult to measure success n n n 500 relational objects 5000 positive links means 245, 000 negative links Always negative classifier is 98% accurate ROC curves look overly optimistic Enormous quantity of data n n 4, 285, 199, 774 web pages indexed by Google Pub. Med includes over 15 million citations

Our Approach Develop fast ensemble algorithms focused on recall and precision evaluation n Key Ideas of Gleaner n n Keep wide range of clauses Create separate theories for different recall ranges Evaluation n n Area Under Recall Precision Curve (AURPC) Time = Number of clauses considered

Gleaner - Background n Focus evaluation on positive examples n n n Recall = TP TP + FN Precision = TP TP + FP Rapid Random Restart (Zelezny et al ILP 2002) n n n Stochastic selection of starting clause Time-limited local heuristic search We store variety of clauses (based on recall)
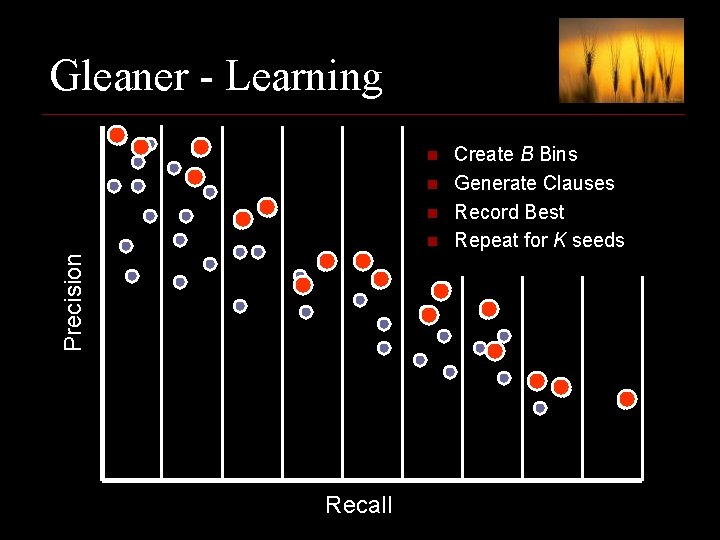
Gleaner - Learning n n n Precision n Recall Create B Bins Generate Clauses Record Best Repeat for K seeds

Gleaner - Combining n Combine K clauses per bin n n How to choose L ? n n n L=1 then high recall, low precision L=K then low recall, high precision Our method n n n If at least L of K clauses match, call example positive Choose L such that ensemble recall matches bin b Bin b’s precision should be higher than any clause in it We should now have set of high precision rule sets spanning space of recall levels
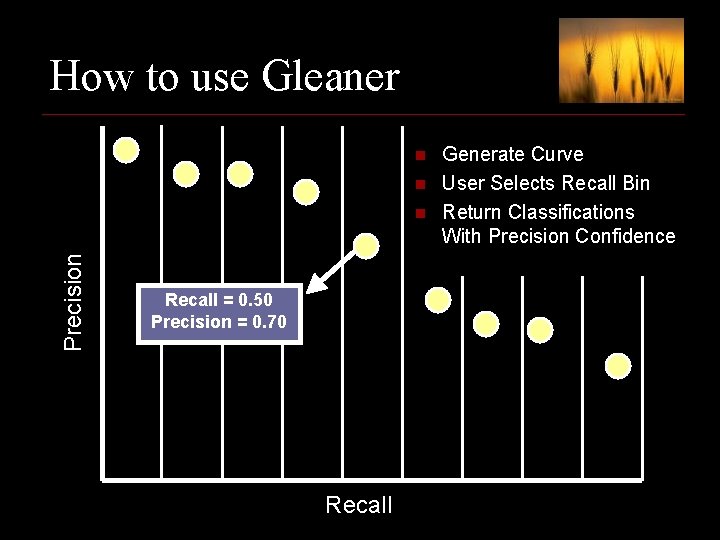
How to use Gleaner n n Precision n Recall = 0. 50 Precision = 0. 70 Recall Generate Curve User Selects Recall Bin Return Classifications With Precision Confidence

Aleph Ensembles We compare to ensembles of theories n Algorithm (Dutra et al ILP 2002) n n n Use K different initial seeds Learn K theories containing C clauses Rank examples by the number of theories Need to balance C for high performance n n Small C leads to low recall Large C leads to converging theories
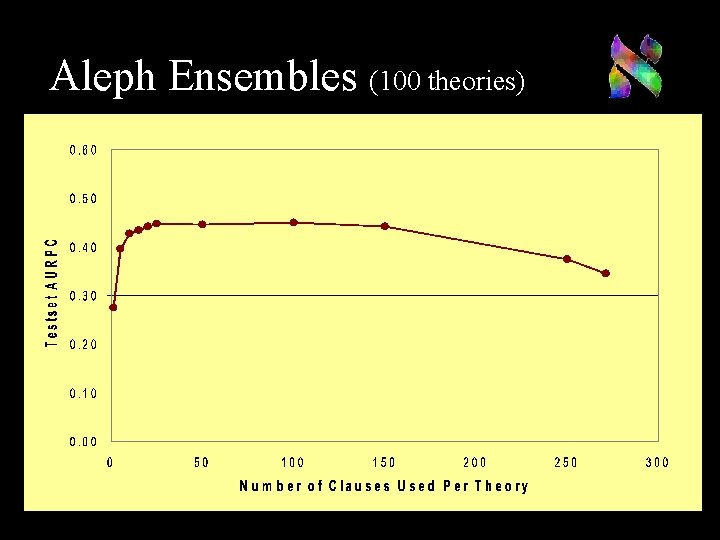
Aleph Ensembles (100 theories)

Biomedical Information Extraction Given: Medical Journal abstracts tagged with protein localization relations n Do: Construct system to extract protein localization phrases from unseen text n NPL 3 encodes a nuclear protein with an RNA recognition motif and similarities to a family of proteins involved in RNA metabolism.

Biomedical Information Extraction n Hand-labeled dataset (Ray & Craven ’ 01) n n 7, 245 sentences from 871 abstracts Examples are phrase-phrase combinations n n 1. 6 GB of background knowledge n n n 1, 810 positive & 279, 154 negative Structural, Statistical, Lexical and Ontological In total, 200+ distinct background predicates More info on dataset by Jude Shavlik later
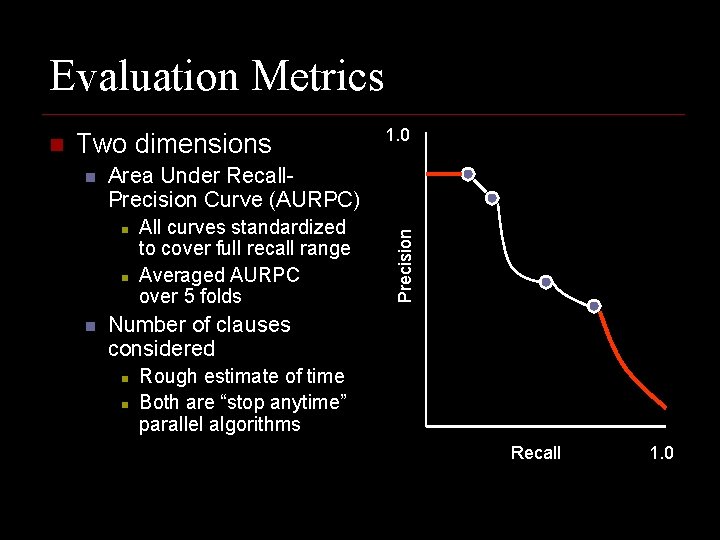
Evaluation Metrics Two dimensions n Area Under Recall. Precision Curve (AURPC) n n n 1. 0 All curves standardized to cover full recall range Averaged AURPC over 5 folds Precision n Number of clauses considered n n Rough estimate of time Both are “stop anytime” parallel algorithms Recall 1. 0
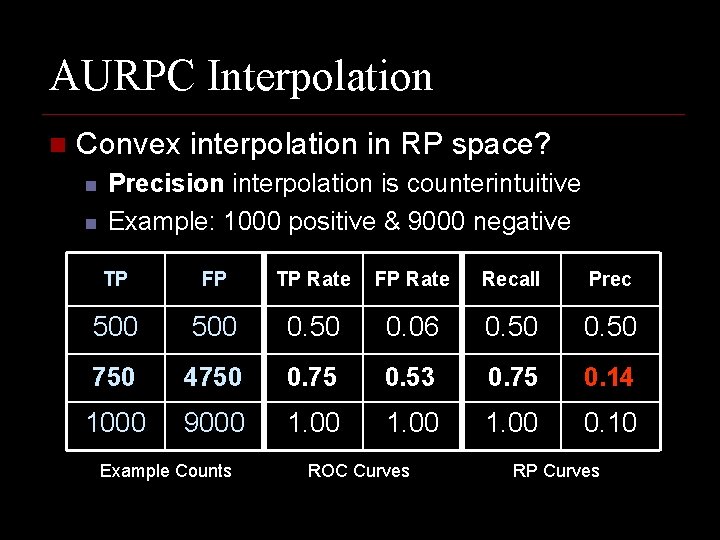
AURPC Interpolation n Convex interpolation in RP space? n n Precision interpolation is counterintuitive Example: 1000 positive & 9000 negative TP FP TP Rate FP Rate Recall Prec 500 0. 50 0. 06 0. 50 750 4750 0. 75 0. 53 0. 75 0. 14 1000 9000 1. 00 0. 10 Example Counts ROC Curves RP Curves
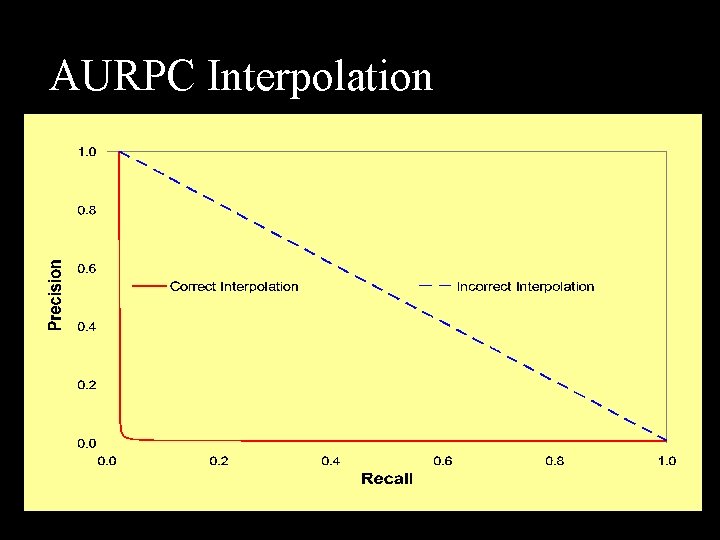
AURPC Interpolation

Experimental Methodology Performed five-fold cross-validation n Variation of parameters n n Gleaner (20 recall bins) n n n # seeds = {25, 50, 75, 100} # clauses = {1 K, 10 K, 25 K, 50 K, 100 K, 250 K, 500 K} Ensembles (0. 75 minacc, 35, 000 nodes) n n # theories = {10, 25, 50, 75, 100} # clauses per theory = {1, 5, 10, 15, 20, 25, 50}
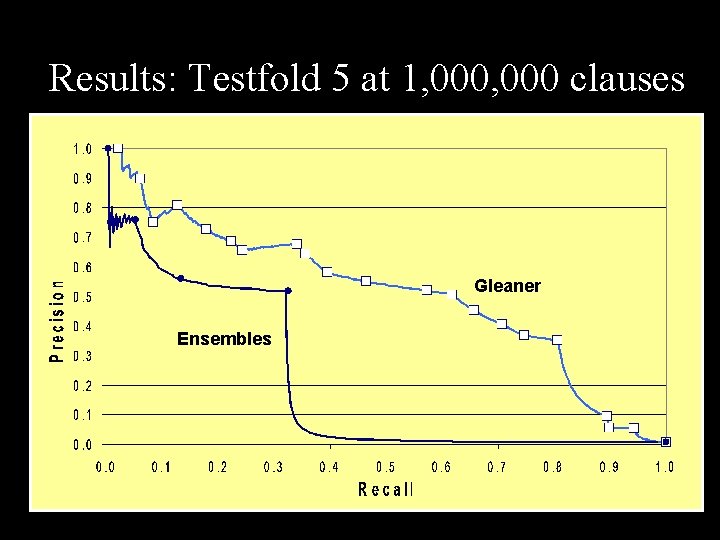
Results: Testfold 5 at 1, 000 clauses Gleaner Ensembles
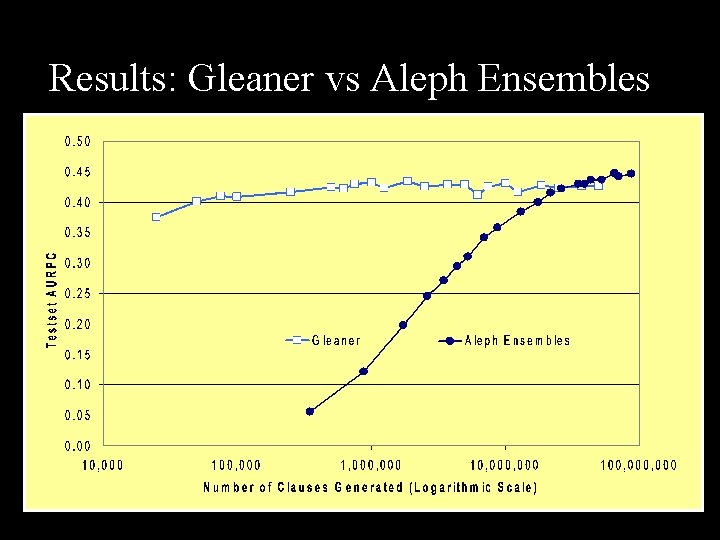
Results: Gleaner vs Aleph Ensembles

Conclusions n Gleaner n n Aleph ensembles n n n Focuses on recall and precision Keeps wide spectrum of clauses Good results in few cpu cycles ‘Early stopping’ helpful Require more cpu cycles AURPC n n Useful metric for comparison Interpolation unintuitive

Future Work Improve Gleaner performance over time n Explore alternate clause combinations n Better understanding of AURPC n Search for clauses that optimize AURPC n Examine more ILP link-learning datasets n Use Gleaner with other ML algorithms n

Take-Home Message n Definition of Gleaner n n Gleaner and ILP n n n One who gathers grain left behind by reapers Many clauses constructed and evaluated in ILP hypothesis search We need to make better use of those that aren’t the highest scoring ones Thanks, Questions?

Acknowledgements n n n n USA NLM Grant 5 T 15 LM 007359 -02 USA NLM Grant 1 R 01 LM 07050 -01 USA DARPA Grant F 30602 -01 -2 -0571 USA Air Force Grant F 30602 -01 -2 -0571 Condor Group David Page Vitor Santos Costa, Ines Dutra Soumya Ray, Marios Skounakis, Mark Craven Dataset available at (URL in proceedings) ftp: //ftp. cs. wisc. edu/machine-learning/shavlik-group/datasets/IE-protein-location

Deleted Scenes Aleph Learning n Clause Weighting n Sample Gleaner Recall-Precision Curve n Sample Extraction Clause n Gleaner Algorithm n Director Commentary on off

Aleph - Learning n Aleph learns theories of clauses (Srinivasan, v 4, 2003) n n Pick a positive seed example and saturate Use heuristic search to find best clause Pick new seed from uncovered positives and repeat until threshold of positives covered Theory produces one recall-precision point n n Learning complete theories is time-consuming Can produce ranking with theory ensembles

Clause Weighting n Single Theory Ensemble n n rank by how many clauses cover examples Weight clauses using tuneset statistics n n n CN 2 (average precision of matching clauses) Lowest False Positive Rate Score Cumulative n n F 1 score Recall Precision Diversity
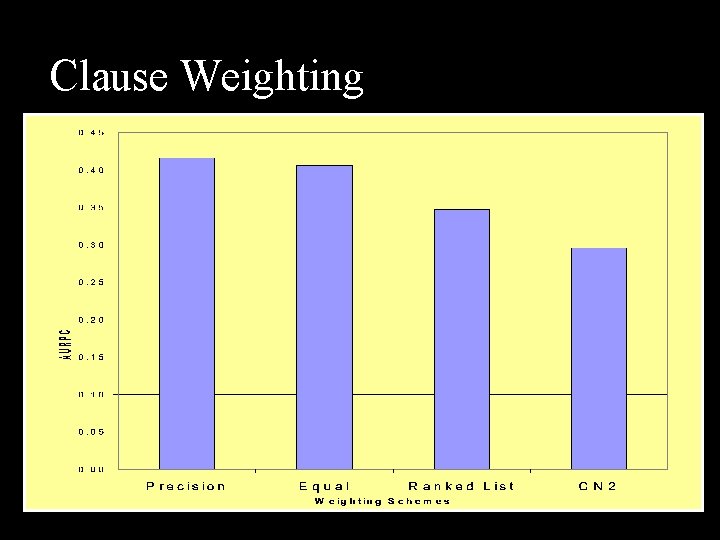
Clause Weighting
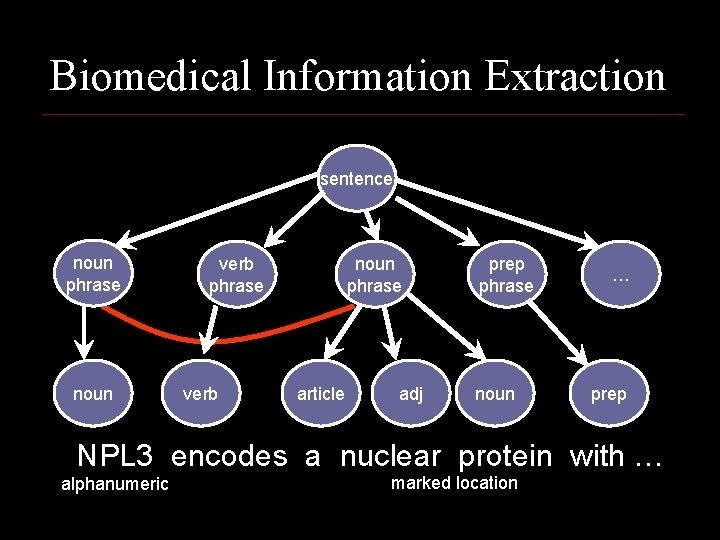
Biomedical Information Extraction sentence noun phrase noun verb phrase verb noun phrase article adj prep phrase noun … prep NPL 3 encodes a nuclear protein with … alphanumeric marked location
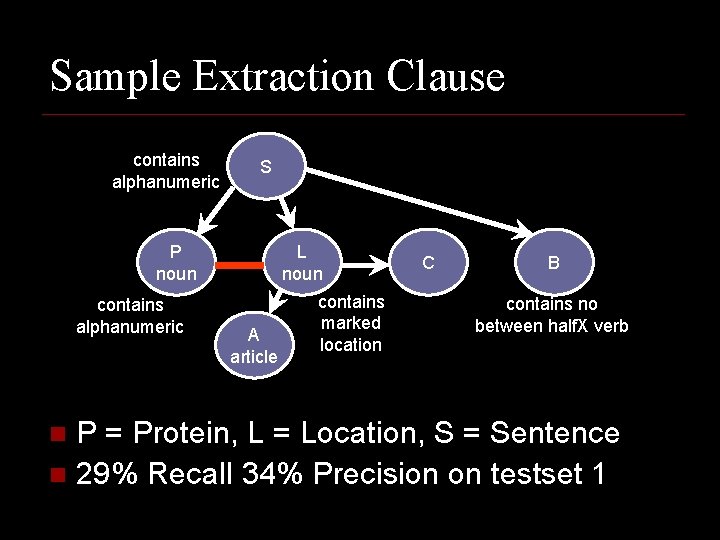
Sample Extraction Clause contains alphanumeric S P noun contains alphanumeric L noun A article contains marked location C B contains no between half. X verb P = Protein, L = Location, S = Sentence n 29% Recall 34% Precision on testset 1 n

Gleaner Algorithm n n Create B equal-sized recall bins For K different seeds n n n For each recall bin B n n Generate rules using Rapid Random Restart Record best rule (precision x recall) found for each bin Find threshold L of K clauses such that recall of “at least L of K clauses match examples” = recall for this bin Find recall and precision on testset using each bin’s “at least L of K” decision process
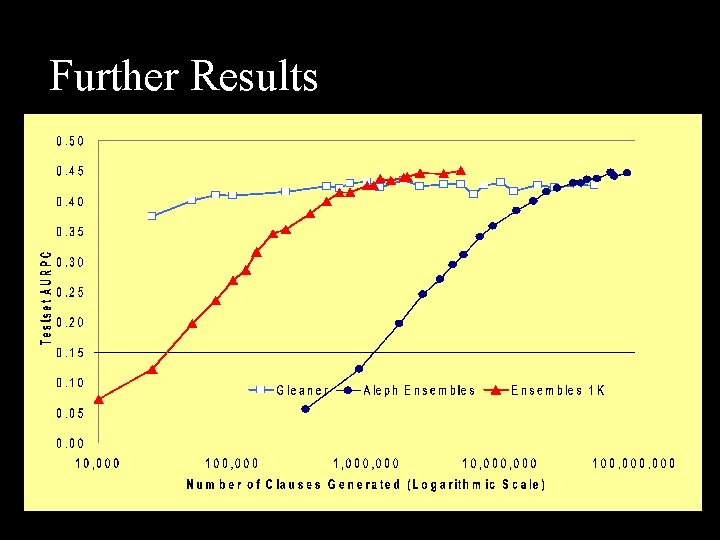
Further Results
- Slides: 31