Learning Bayesian Networks from Data Nir Friedman Hebrew

Learning Bayesian Networks from Data Nir Friedman Hebrew U. . Daphne Koller Stanford

Overview u Introduction u Parameter Estimation u Model Selection u Structure Discovery u Incomplete Data u Learning from Structured Data 2
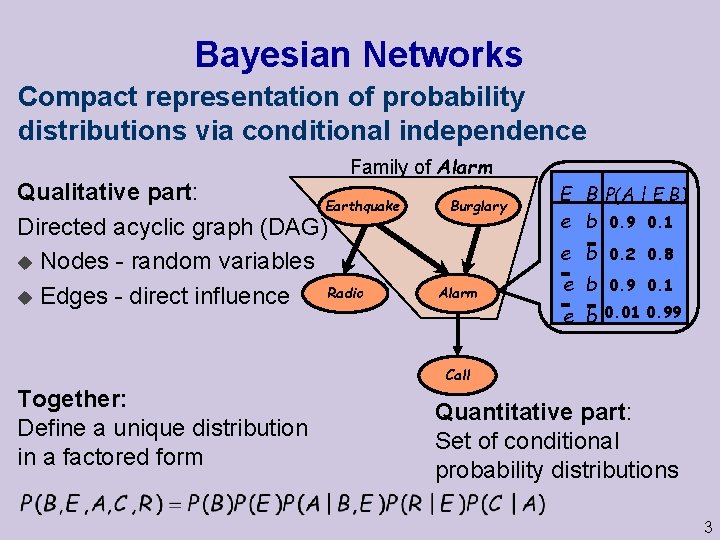
Bayesian Networks Compact representation of probability distributions via conditional independence Family of Alarm Qualitative part: Earthquake Directed acyclic graph (DAG) u Nodes - random variables Radio u Edges - direct influence Burglary Alarm E B P(A | E, B) e b 0. 9 0. 1 e b 0. 2 0. 8 e b 0. 9 0. 1 e b 0. 01 0. 99 Call Together: Define a unique distribution in a factored form Quantitative part: Set of conditional probability distributions 3
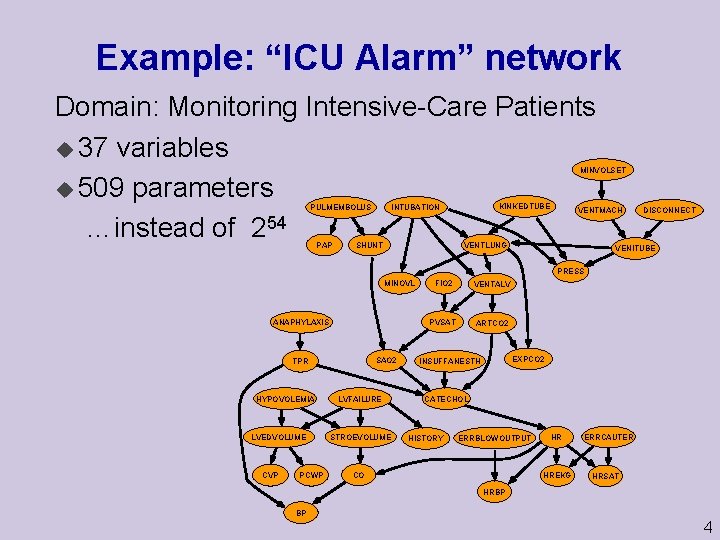
Example: “ICU Alarm” network Domain: Monitoring Intensive-Care Patients u 37 variables u 509 parameters …instead of 254 MINVOLSET PULMEMBOLUS PAP KINKEDTUBE INTUBATION SHUNT VENTMACH VENTLUNG DISCONNECT VENITUBE PRESS MINOVL ANAPHYLAXIS SAO 2 TPR HYPOVOLEMIA LVEDVOLUME CVP PCWP LVFAILURE STROEVOLUME FIO 2 VENTALV PVSAT ARTCO 2 EXPCO 2 INSUFFANESTH CATECHOL HISTORY ERRBLOWOUTPUT CO HR HREKG ERRCAUTER HRSAT HRBP BP 4
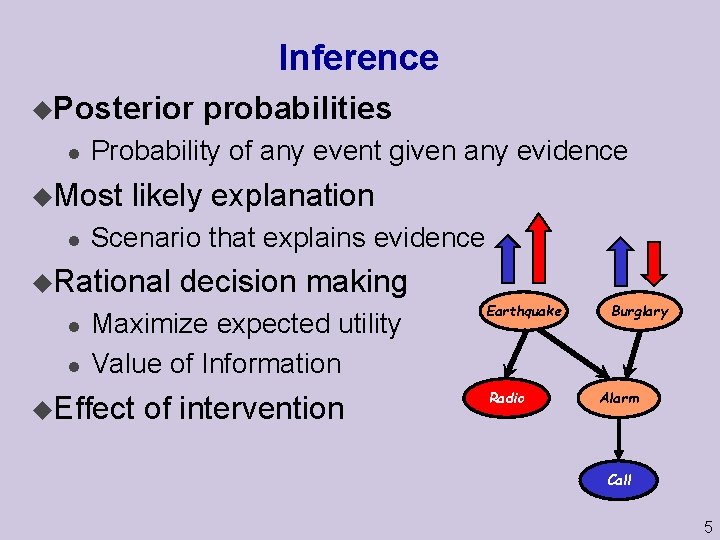
Inference u. Posterior l Probability of any event given any evidence u. Most l likely explanation Scenario that explains evidence u. Rational l l probabilities decision making Maximize expected utility Value of Information u. Effect of intervention Earthquake Radio Burglary Alarm Call 5

Why learning? Knowledge acquisition bottleneck u Knowledge acquisition is an expensive process u Often we don’t have an expert Data is cheap u Amount of available information growing rapidly u Learning allows us to construct models from raw data 6

Why Learn Bayesian Networks? u Conditional independencies & graphical language capture structure of many real-world distributions u Graph l structure provides much insight into domain Allows “knowledge discovery” u Learned model can be used for many tasks u Supports l l all the features of probabilistic learning Model selection criteria Dealing with missing data & hidden variables 7
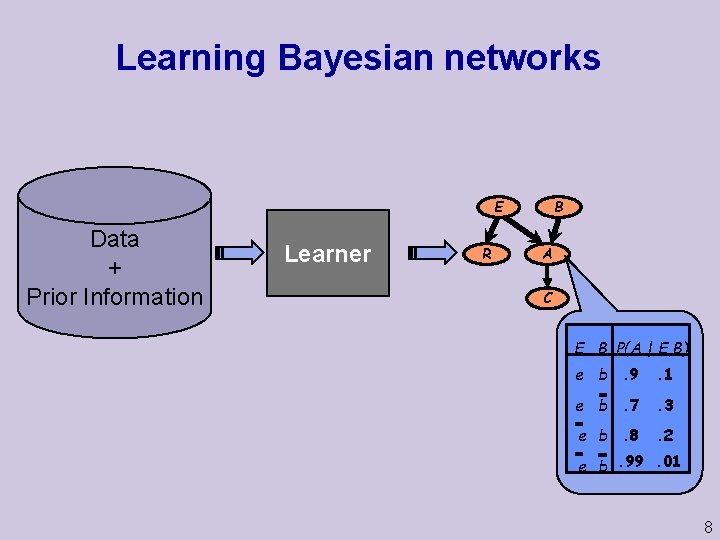
Learning Bayesian networks B E Data + Prior Information Learner R A C E B P(A | E, B) e b. 9 . 1 e b. 7 . 3 e b. 8 . 2 e b. 99. 01 8
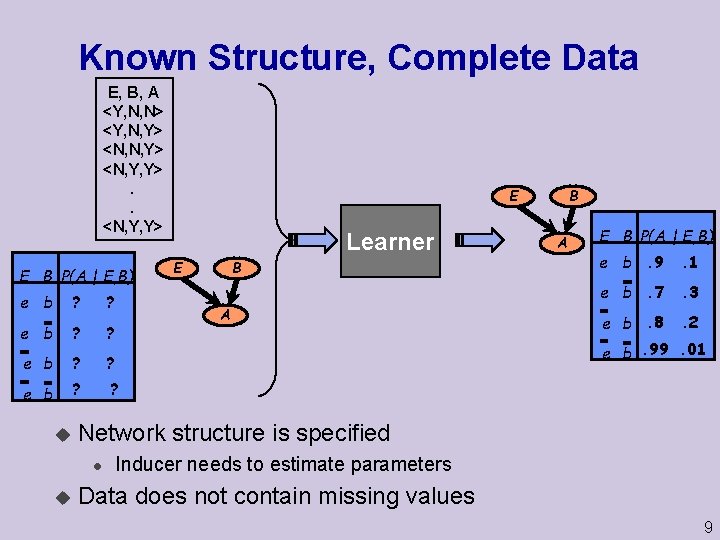
Known Structure, Complete Data E, B, A <Y, N, N> <Y, N, Y> <N, Y, Y>. . <N, Y, Y> E B P(A | E, B) e b ? ? u Learner B E A A E B P(A | E, B) e b. 9 . 1 e b. 7 . 3 e b. 8 . 2 e b. 99. 01 Network structure is specified l u B E Inducer needs to estimate parameters Data does not contain missing values 9
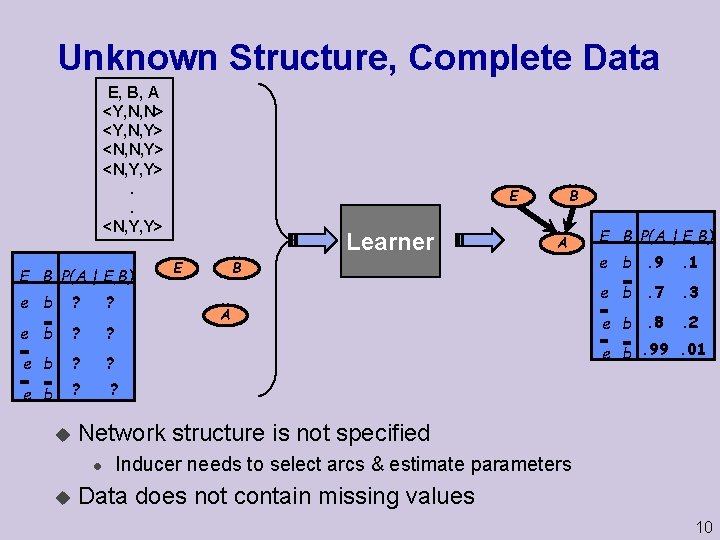
Unknown Structure, Complete Data E, B, A <Y, N, N> <Y, N, Y> <N, Y, Y>. . <N, Y, Y> E B P(A | E, B) e b ? ? u Learner A B E A E B P(A | E, B) e b. 9 . 1 e b. 7 . 3 e b. 8 . 2 e b. 99. 01 Network structure is not specified l u B E Inducer needs to select arcs & estimate parameters Data does not contain missing values 10
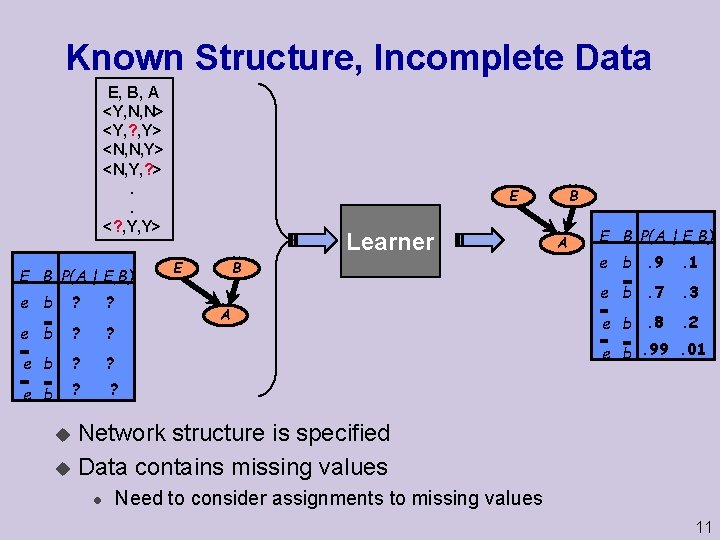
Known Structure, Incomplete Data E, B, A <Y, N, N> <Y, ? , Y> <N, N, Y> <N, Y, ? >. . <? , Y, Y> E B P(A | E, B) e b ? ? B E Learner B E A A E B P(A | E, B) e b. 9 . 1 e b. 7 . 3 e b. 8 . 2 e b. 99. 01 Network structure is specified u Data contains missing values u l Need to consider assignments to missing values 11
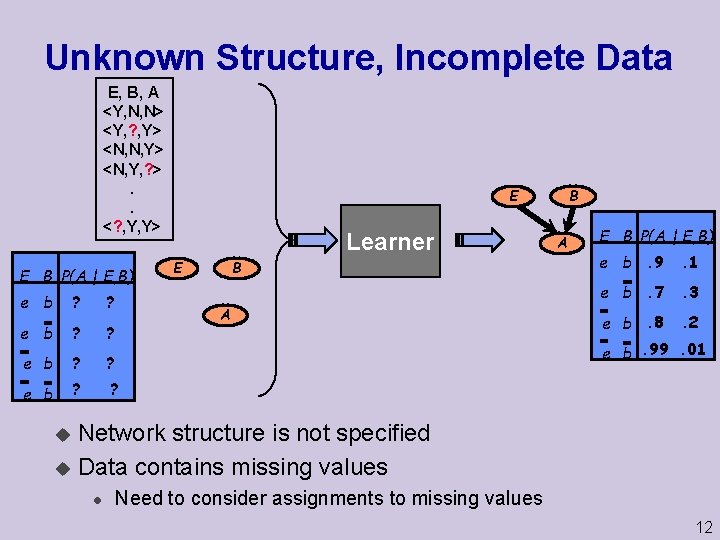
Unknown Structure, Incomplete Data E, B, A <Y, N, N> <Y, ? , Y> <N, N, Y> <N, Y, ? >. . <? , Y, Y> E B P(A | E, B) e b ? ? B E Learner B E A A E B P(A | E, B) e b. 9 . 1 e b. 7 . 3 e b. 8 . 2 e b. 99. 01 Network structure is not specified u Data contains missing values u l Need to consider assignments to missing values 12

Overview u Introduction u Parameter Estimation l Likelihood function l Bayesian estimation u Model Selection u Structure Discovery u Incomplete Data u Learning from Structured Data 13
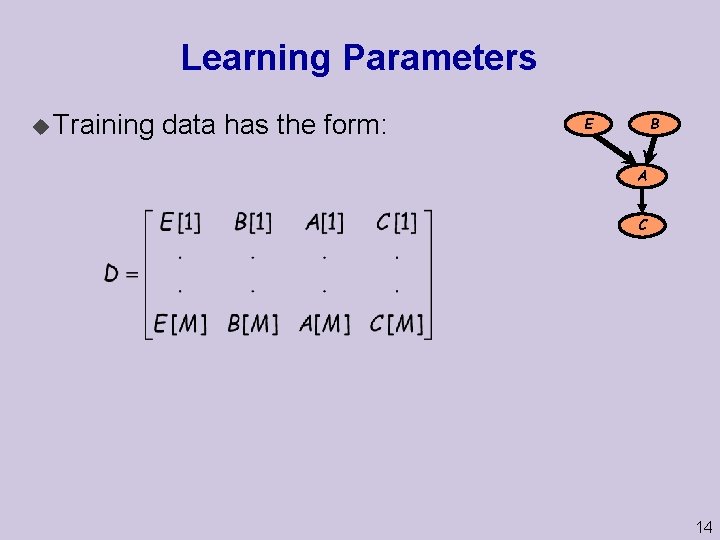
Learning Parameters u Training data has the form: B E A C 14

Likelihood Function u Assume i. i. d. samples u Likelihood function is B E A C 15
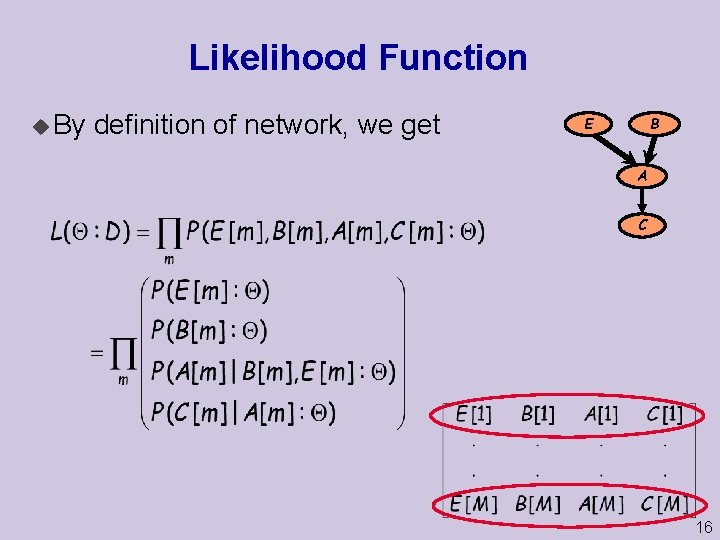
Likelihood Function u By definition of network, we get B E A C 16

Likelihood Function u Rewriting terms, we get B E A C = 17

General Bayesian Networks Generalizing for any Bayesian network: Decomposition Independent estimation problems 18

Likelihood Function: Multinomials likelihood for the sequence H, T, T, H, H is L( : D) u The 0 0. 2 0. 4 General case: 0. 6 0. 8 1 Count of kth outcome in D Probability of kth outcome 19
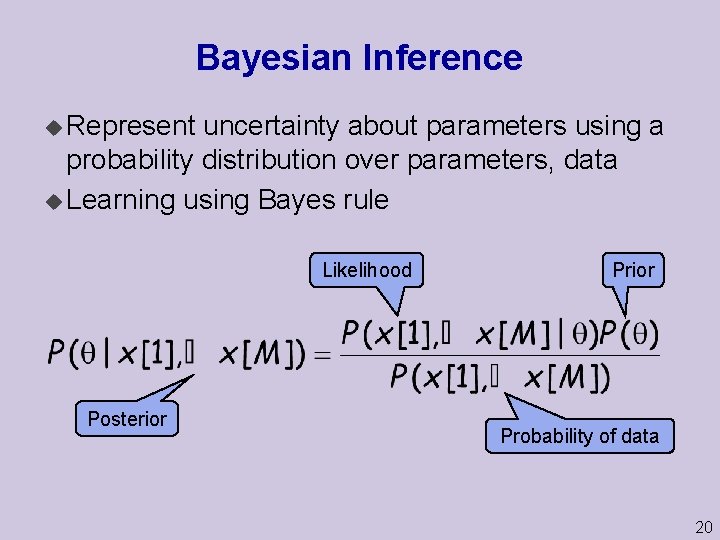
Bayesian Inference u Represent uncertainty about parameters using a probability distribution over parameters, data u Learning using Bayes rule Likelihood Posterior Probability of data 20
![Bayesian Inference u Represent Bayesian distribution as Bayes net X[1] X[2] X[m] Observed data Bayesian Inference u Represent Bayesian distribution as Bayes net X[1] X[2] X[m] Observed data](http://slidetodoc.com/presentation_image_h2/09309c47a36aa199612bc0b9622ab1d9/image-21.jpg)
Bayesian Inference u Represent Bayesian distribution as Bayes net X[1] X[2] X[m] Observed data values of X are independent given u P(x[m] | ) = u Bayesian prediction is inference in this network u The 21
![Example: Binomial Data uniform for in [0, 1] P( |D) the likelihood L( : Example: Binomial Data uniform for in [0, 1] P( |D) the likelihood L( :](http://slidetodoc.com/presentation_image_h2/09309c47a36aa199612bc0b9622ab1d9/image-22.jpg)
Example: Binomial Data uniform for in [0, 1] P( |D) the likelihood L( : D) u Prior: (NH, NT ) = (4, 1) u MLE for P(X = H ) is 4/5 = 0. 8 u Bayesian prediction is 0 0. 2 0. 4 0. 6 0. 8 1 22

Dirichlet Priors u Recall that the likelihood function is u Dirichlet prior with hyperparameters 1, …, K the posterior has the same form, with hyperparameters 1+N 1, …, K +N K 23
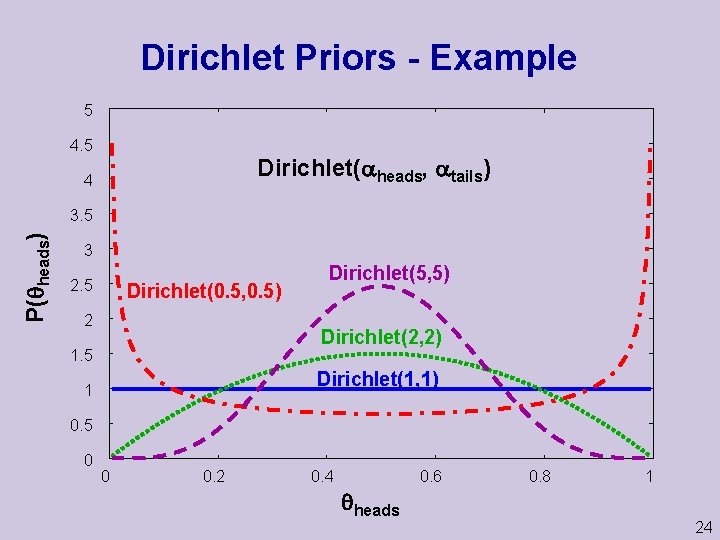
Dirichlet Priors Example 5 4. 5 Dirichlet( heads, tails) 4 P( heads) 3. 5 3 2. 5 Dirichlet(0. 5, 0. 5) 2 Dirichlet(5, 5) Dirichlet(2, 2) 1. 5 Dirichlet(1, 1) 1 0. 5 0 0 0. 2 0. 4 0. 6 heads 0. 8 1 24

Dirichlet Priors (cont. ) u If P( ) is Dirichlet with hyperparameters 1, …, K u Since the posterior is also Dirichlet, we get 25
![Bayesian Nets & Bayesian Prediction Y|X X X m X[1] X[2] X[M+1] Y[2] Y[M+1] Bayesian Nets & Bayesian Prediction Y|X X X m X[1] X[2] X[M+1] Y[2] Y[M+1]](http://slidetodoc.com/presentation_image_h2/09309c47a36aa199612bc0b9622ab1d9/image-26.jpg)
Bayesian Nets & Bayesian Prediction Y|X X X m X[1] X[2] X[M+1] Y[2] Y[M+1] Observed data Query X[m] Y|X Y[m] Plate notation u Priors for each parameter group are independent u Data instances are independent given the unknown parameters 26
![Bayesian Nets & Bayesian Prediction Y|X X X[1] X[2] X[M] Y[1] Y[2] Y[M] Observed Bayesian Nets & Bayesian Prediction Y|X X X[1] X[2] X[M] Y[1] Y[2] Y[M] Observed](http://slidetodoc.com/presentation_image_h2/09309c47a36aa199612bc0b9622ab1d9/image-27.jpg)
Bayesian Nets & Bayesian Prediction Y|X X X[1] X[2] X[M] Y[1] Y[2] Y[M] Observed data u We can also “read” from the network: Complete data posteriors on parameters are independent u Can compute posterior over parameters separately! 27

Learning Parameters: Summary u Estimation l l relies on sufficient statistics For multinomials: counts N(xi, pai) Parameter estimation MLE Bayesian (Dirichlet) u Both are asymptotically equivalent and consistent u Both can be implemented in an on-line manner by accumulating sufficient statistics 28
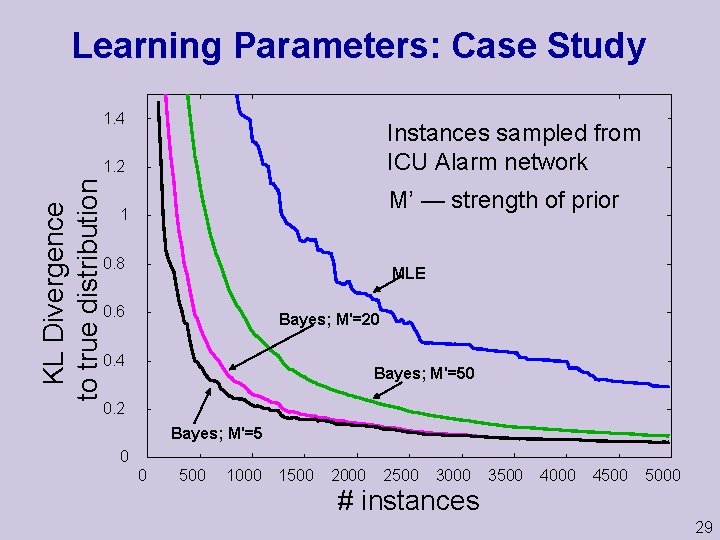
Learning Parameters: Case Study 1. 4 Instances sampled from ICU Alarm network KL Divergence to true distribution 1. 2 M’ — strength of prior 1 0. 8 MLE 0. 6 Bayes; M'=20 0. 4 Bayes; M'=50 0. 2 Bayes; M'=5 0 0 500 1000 1500 2000 2500 3000 3500 4000 4500 5000 # instances 29

Overview u Introduction u Parameter Learning u Model Selection l Scoring function l Structure search u Structure Discovery u Incomplete Data u Learning from Structured Data 30
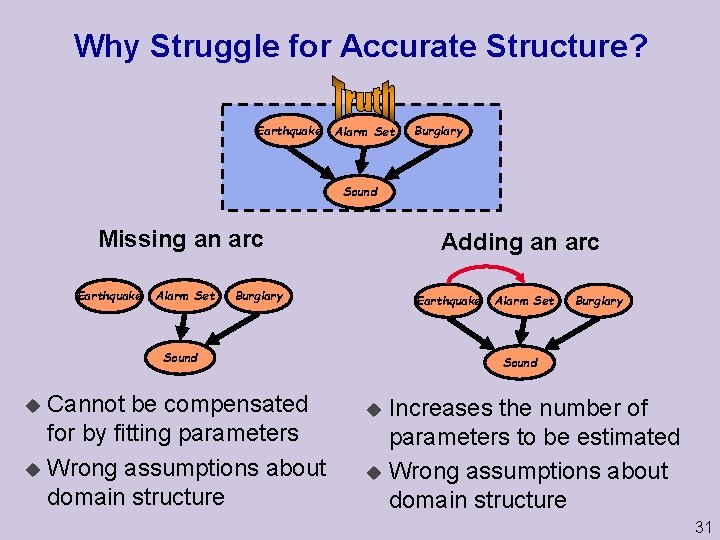
Why Struggle for Accurate Structure? Earthquake Alarm Set Burglary Sound Missing an arc Earthquake Alarm Set Adding an arc Burglary Earthquake Sound Cannot be compensated for by fitting parameters u Wrong assumptions about domain structure u Alarm Set Burglary Sound Increases the number of parameters to be estimated u Wrong assumptions about domain structure u 31
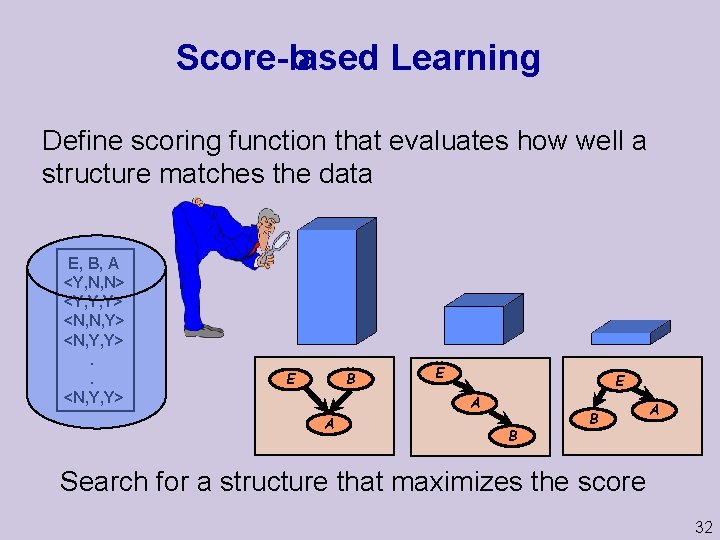
Score based Learning Define scoring function that evaluates how well a structure matches the data E, B, A <Y, N, N> <Y, Y, Y> <N, N, Y> <N, Y, Y>. . <N, Y, Y> B E E E A A B B A Search for a structure that maximizes the score 32

Likelihood Score for Structure Mutual information between Xi and its parents u Larger dependence of Xi on Pai higher score u Adding l l l arcs always helps I(X; Y) I(X; {Y, Z}) Max score attained by fully connected network Overfitting: A bad idea… 33
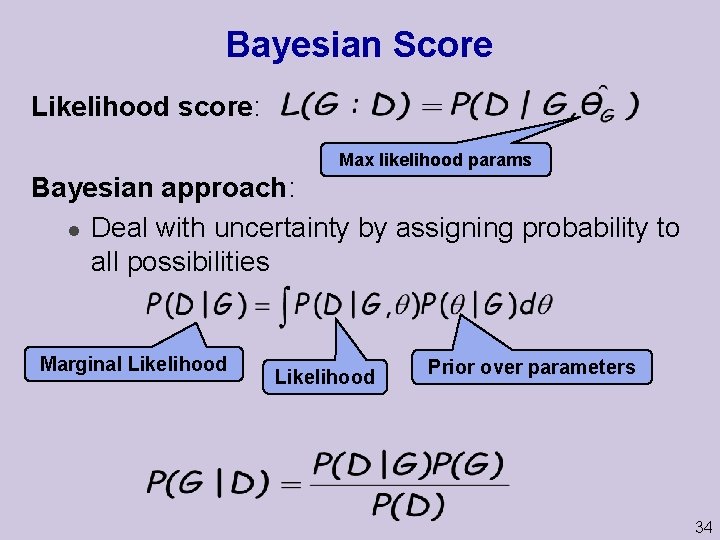
Bayesian Score Likelihood score: Max likelihood params Bayesian approach: l Deal with uncertainty by assigning probability to all possibilities Marginal Likelihood Prior over parameters 34

Marginal Likelihood: Multinomials Fortunately, in many cases integral has closed form u P( ) is Dirichlet with hyperparameters 1, …, K u D is a dataset with sufficient statistics N 1, …, NK Then 35
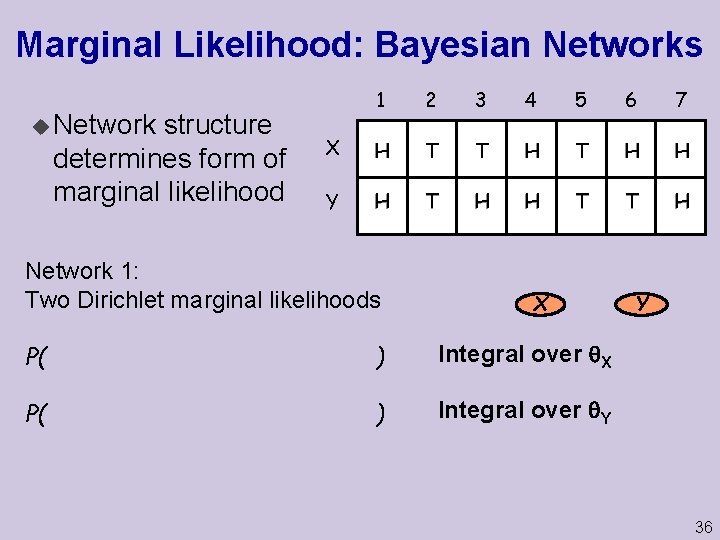
Marginal Likelihood: Bayesian Networks u Network structure determines form of marginal likelihood 1 2 3 4 5 6 7 X H T T H H Y H T H H T T H Network 1: Two Dirichlet marginal likelihoods X P( ) Integral over Y Y 36
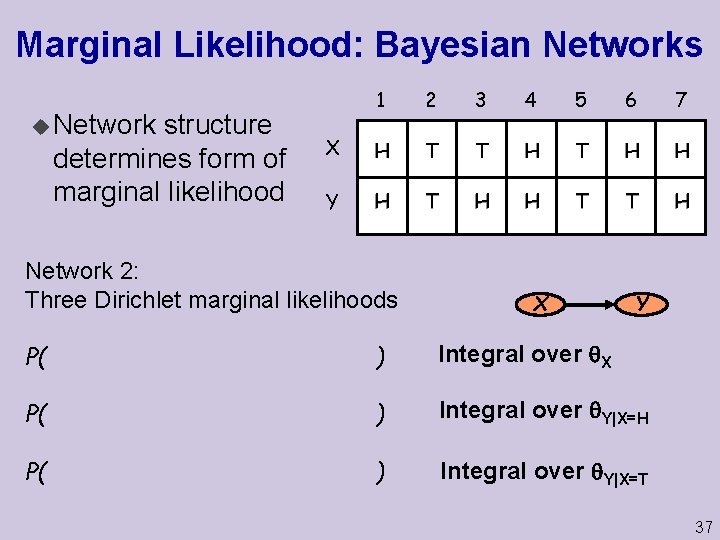
Marginal Likelihood: Bayesian Networks u Network structure determines form of marginal likelihood 1 2 3 4 5 6 7 X H T T H H Y H T H H T T H Network 2: Three Dirichlet marginal likelihoods X Y P( ) Integral over X P( ) Integral over Y|X=H P( ) Integral over Y|X=T 37

Marginal Likelihood for Networks The marginal likelihood has the form: Dirichlet marginal likelihood for multinomial P(Xi | pai) N(. . ) are counts from the data (. . ) are hyperparameters for each family given G 38

Bayesian Score: Asymptotic Behavior Fit dependencies in empirical distribution u As Complexity penalty M (amount of data) grows, Increasing pressure to fit dependencies in distribution l Complexity term avoids fitting noise l u Asymptotic equivalence to MDL score u Bayesian score is consistent l Observed data eventually overrides prior 39

Structure Search as Optimization Input: l Training data l Scoring function l Set of possible structures Output: l A network that maximizes the score Key Computational Property: Decomposability: score(G) = score ( family of X in G ) 40
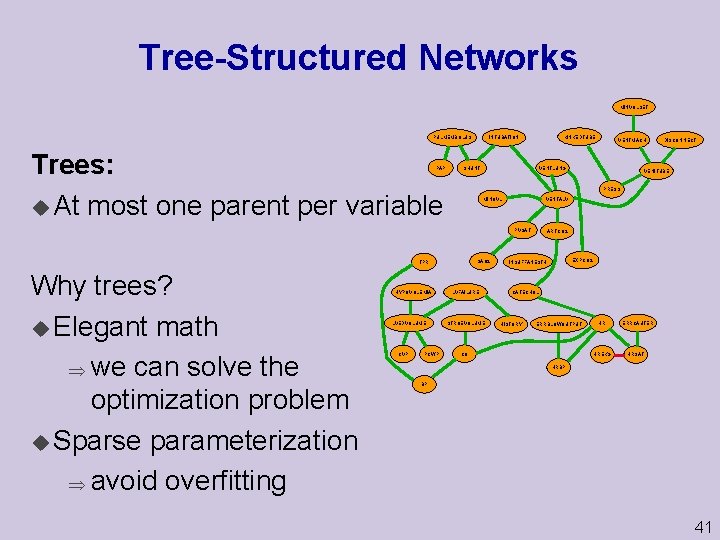
Tree Structured Networks MINVOLSET PULMEMBOLUS Trees: u At most one parent per variable PAP KINKEDTUBE INTUBATION SHUNT VENTLUNG SAO 2 TPR HYPOVOLEMIA LVEDVOLUME CVP PCWP DISCONNECT VENITUBE PRESS MINOVL VENTALV PVSAT Why trees? u Elegant math we can solve the optimization problem u Sparse parameterization avoid overfitting VENTMACH LVFAILURE STROEVOLUME ARTCO 2 EXPCO 2 INSUFFANESTH CATECHOL HISTORY ERRBLOWOUTPUT CO HR HREKG ERRCAUTER HRSAT HRBP BP 41
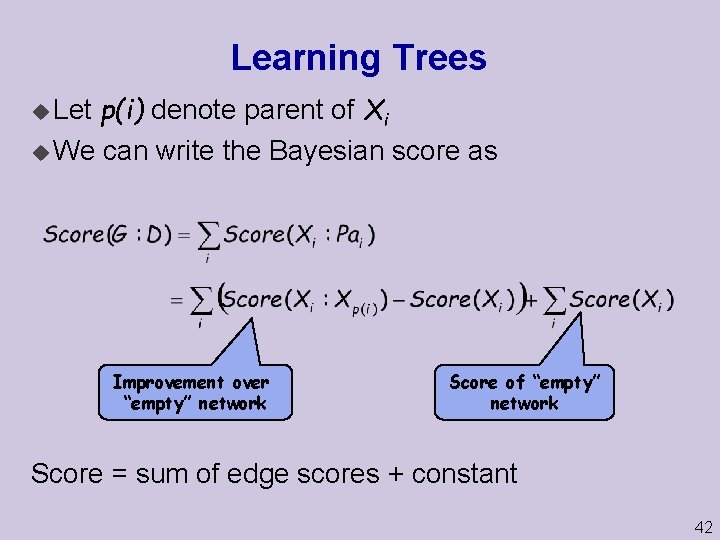
Learning Trees p(i) denote parent of Xi u We can write the Bayesian score as u Let Improvement over “empty” network Score of “empty” network Score = sum of edge scores + constant 42
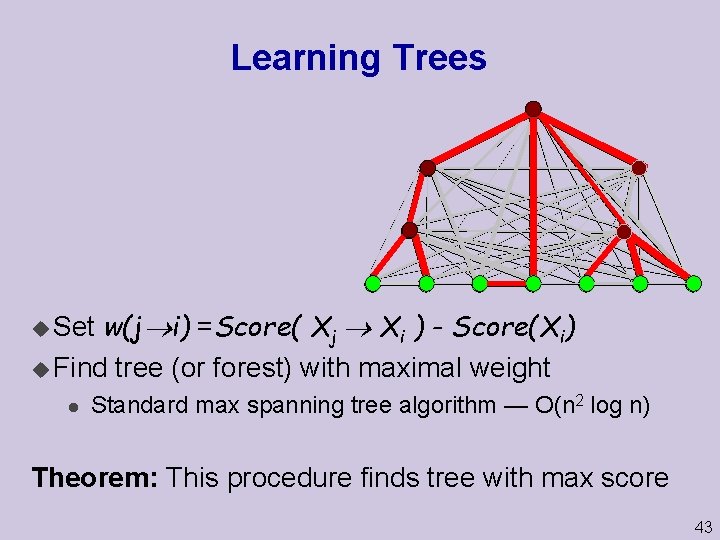
Learning Trees w(j i) =Score( Xj Xi ) - Score(Xi) u Find tree (or forest) with maximal weight u Set l Standard max spanning tree algorithm — O(n 2 log n) Theorem: This procedure finds tree with max score 43

Beyond Trees When we consider more complex network, the problem is not as easy u Suppose we allow at most two parents per node u A greedy algorithm is no longer guaranteed to find the optimal network u In fact, no efficient algorithm exists Theorem: Finding maximal scoring structure with at most k parents per node is NP-hard for k > 1 44

Heuristic Search u Define a search space: l search states are possible structures l operators make small changes to structure u Traverse space looking for high-scoring structures u Search techniques: l Greedy hill-climbing l Best first search l Simulated Annealing l. . . 45

Local Search u Start l l l with a given network empty network best tree a random network u At each iteration l Evaluate all possible changes l Apply change based on score u Stop when no modification improves score 46
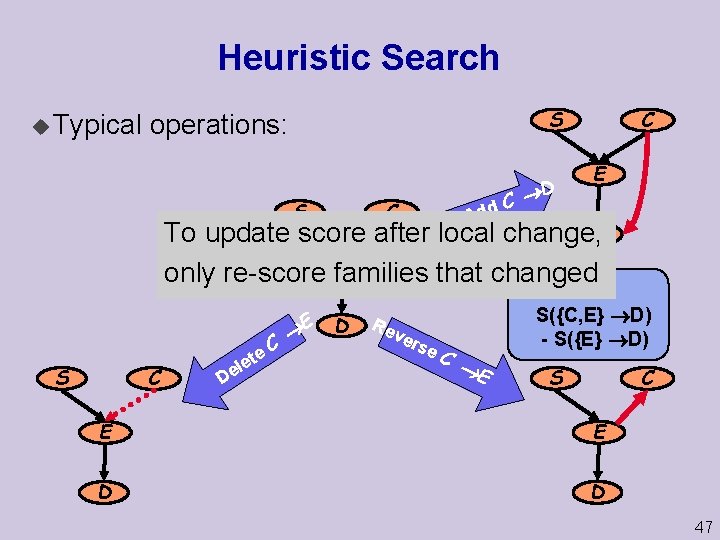
Heuristic Search u Typical S operations: S D C dd C C E A To update score after local change, D E only re-score families that changed S C D el C e t e E D Re ver se C E score = S({C, E} D) S({E} D) S C E E D D 47

Learning in Practice: Alarm domain 2 true distribution KL Divergence to 1. 5 Structure known, fit params 1 Learn both structure & params 0. 5 0 0 500 1000 1500 2000 2500 3000 #samples 3500 4000 4500 5000 48

Local Search: Possible Pitfalls u Local l search can get stuck in: Local Maxima: ØAll l one-edge changes reduce the score Plateaux: ØSome u Standard l l l one-edge changes leave the score unchanged heuristics can escape both Random restarts TABU search Simulated annealing 49
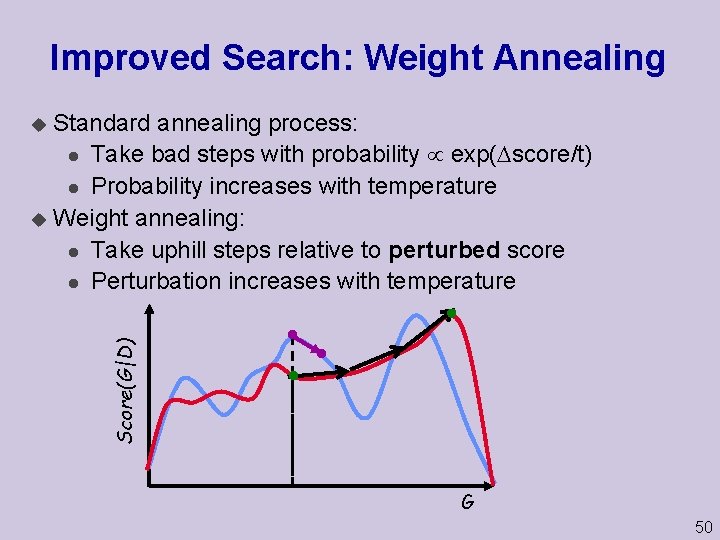
Improved Search: Weight Annealing Standard annealing process: l Take bad steps with probability exp( score/t) l Probability increases with temperature u Weight annealing: l Take uphill steps relative to perturbed score l Perturbation increases with temperature Score(G|D) u G 50

Perturbing the Score u Perturb the score by reweighting instances u Each weight sampled from distribution: l Mean = 1 l Variance temperature u Instances sampled from “original” distribution u … but perturbation changes emphasis Benefit: u Allows global moves in the search space 51
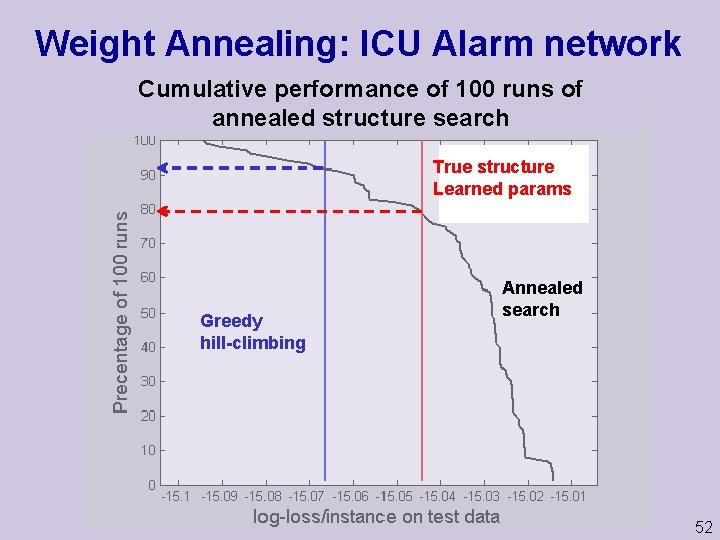
Weight Annealing: ICU Alarm network Cumulative performance of 100 runs of annealed structure search True structure Learned params Greedy hill climbing Annealed search 52

Structure Search: Summary u Discrete optimization problem u In some cases, optimization problem is easy l Example: learning trees u In general, NP-Hard l Need to resort to heuristic search l In practice, search is relatively fast (~100 vars in ~2 -5 min): ØDecomposability ØSufficient l statistics Adding randomness to search is critical 53

Overview u Introduction u Parameter Estimation u Model Selection u Structure Discovery u Incomplete Data u Learning from Structured Data 54

Structure Discovery Task: Discover structural properties l Is there a direct connection between X & Y l Does X separate between two “subsystems” l Does X causally effect Y Example: scientific data mining l Disease properties and symptoms l Interactions between the expression of genes 55
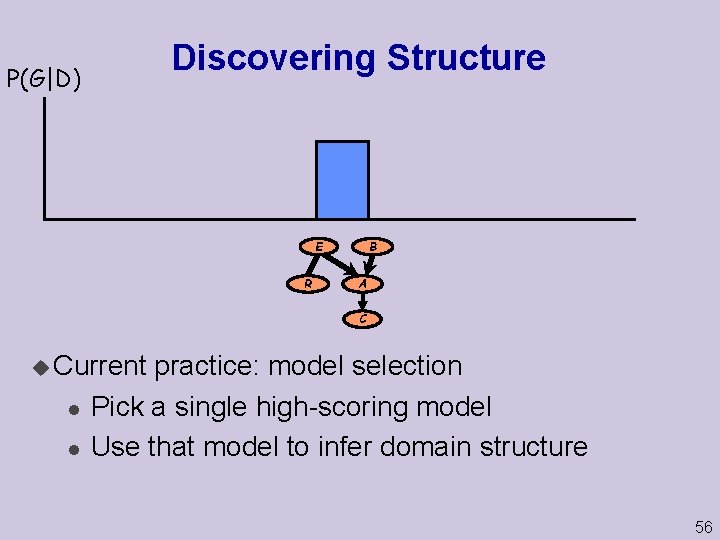
Discovering Structure P(G|D) E R B A C u Current l l practice: model selection Pick a single high-scoring model Use that model to infer domain structure 56
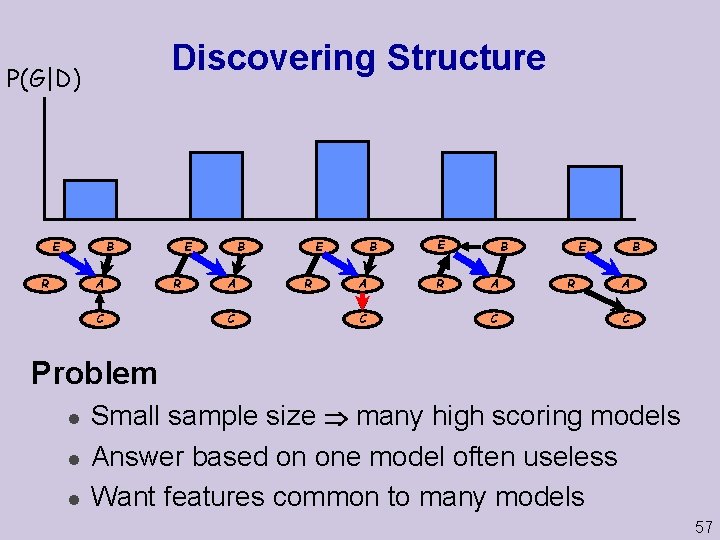
Discovering Structure P(G|D) E B R A C R E B E A C R B A C E R B A C Problem l l l Small sample size many high scoring models Answer based on one model often useless Want features common to many models 57
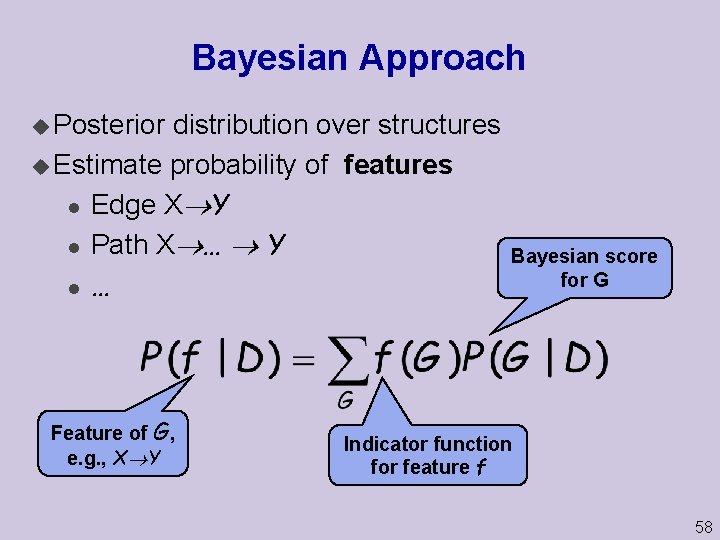
Bayesian Approach u Posterior distribution over structures u Estimate probability of features l Edge X Y l Path X … Y l … Feature of G, e. g. , X Y Bayesian score for G Indicator function for feature f 58

MCMC over Networks u Cannot enumerate structures, so sample structures u MCMC Sampling l Define Markov chain over BNs l Run chain to get samples from posterior P(G | D) Possible pitfalls: l Huge (superexponential) number of networks l Time for chain to converge to posterior is unknown l Islands of high posterior, connected by low bridges 59

ICU Alarm BN: No Mixing instances: Score of cuurent sample u 500 MCMC Iteration u The runs clearly do not mix 60
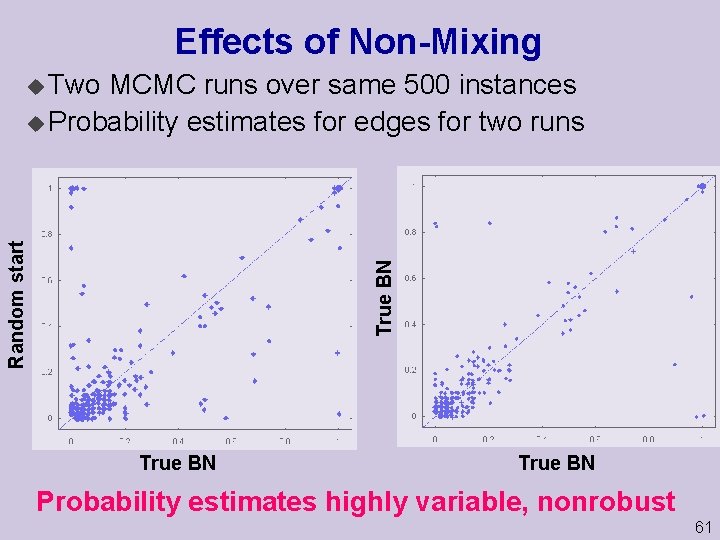
Effects of Non Mixing MCMC runs over same 500 instances u Probability estimates for edges for two runs True BN Random start u Two True BN Probability estimates highly variable, nonrobust 61
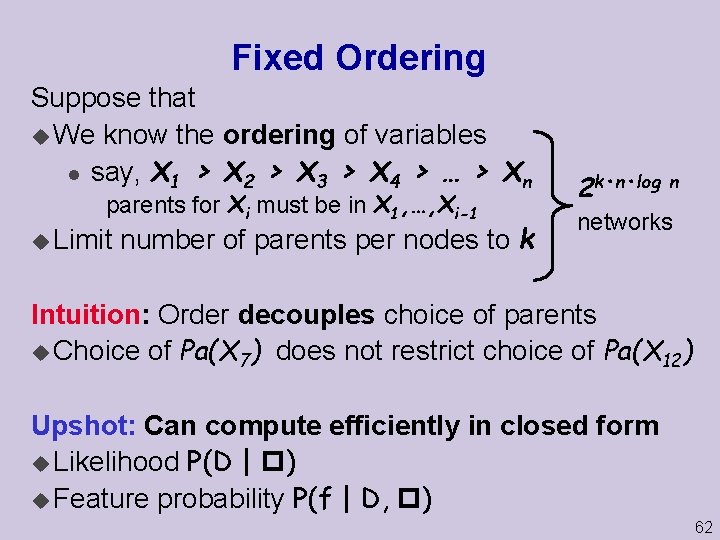
Fixed Ordering Suppose that u We know the ordering of variables l say, X 1 > X 2 > X 3 > X 4 > … > Xn parents for Xi must be in X 1, …, Xi-1 u Limit number of parents per nodes to k 2 k • n • log n networks Intuition: Order decouples choice of parents u Choice of Pa(X 7) does not restrict choice of Pa(X 12) Upshot: Can compute efficiently in closed form u Likelihood P(D | ) u Feature probability P(f | D, ) 62

Our Approach: Sample Orderings We can write Sample orderings and approximate u MCMC Sampling l Define Markov chain over orderings l Run chain to get samples from posterior P( | D) 63

Mixing with MCMC Orderings u 4 runs on ICU-Alarm with 500 instances l Score of cuurent sample l fewer iterations than MCMC-Nets approximately same amount of computation MCMC Iteration Process appears to be mixing! 64
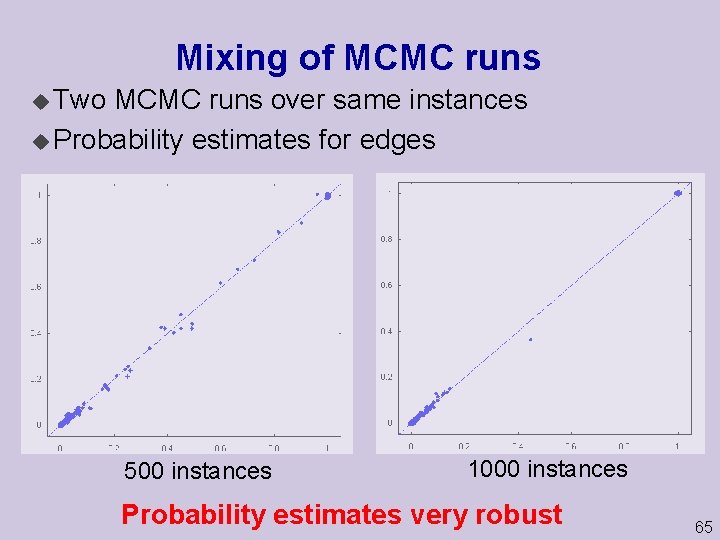
Mixing of MCMC runs u Two MCMC runs over same instances u Probability estimates for edges 500 instances 1000 instances Probability estimates very robust 65

Application: Gene expression Input: u. Measurement of gene expression under different conditions l Thousands of genes l Hundreds of experiments Output: u. Models of gene interaction l Uncover pathways 66
![Map of Feature Confidence Yeast data [Hughes et al 2000] u 600 genes u Map of Feature Confidence Yeast data [Hughes et al 2000] u 600 genes u](http://slidetodoc.com/presentation_image_h2/09309c47a36aa199612bc0b9622ab1d9/image-67.jpg)
Map of Feature Confidence Yeast data [Hughes et al 2000] u 600 genes u 300 experiments 67
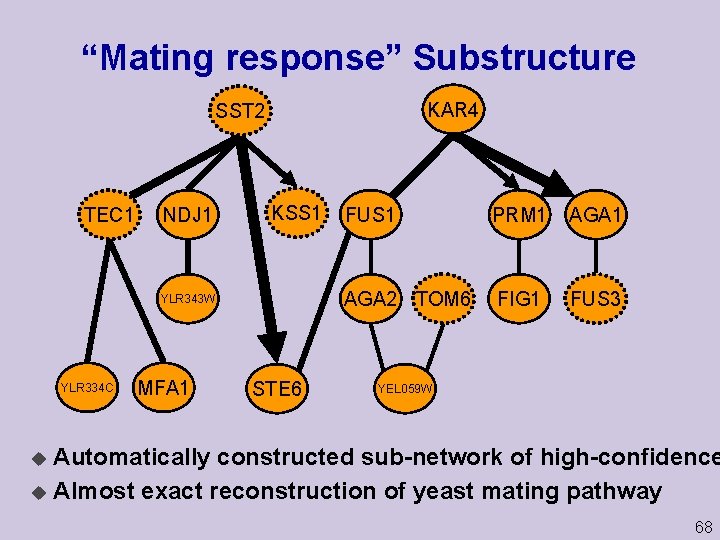
“Mating response” Substructure KAR 4 SST 2 TEC 1 NDJ 1 KSS 1 YLR 343 W YLR 334 C MFA 1 STE 6 FUS 1 PRM 1 AGA 2 TOM 6 FIG 1 FUS 3 YEL 059 W Automatically constructed sub network of high confidence u Almost exact reconstruction of yeast mating pathway u 68

Overview u Introduction u Parameter Estimation u Model Selection u Structure Discovery u Incomplete Data l Parameter estimation l Structure search u Learning from Structured Data 69

Incomplete Data is often incomplete u Some variables of interest are not assigned values This phenomenon happens when we have u Missing values: l Some variables unobserved in some instances u Hidden variables: l Some variables are never observed l We might not even know they exist 70
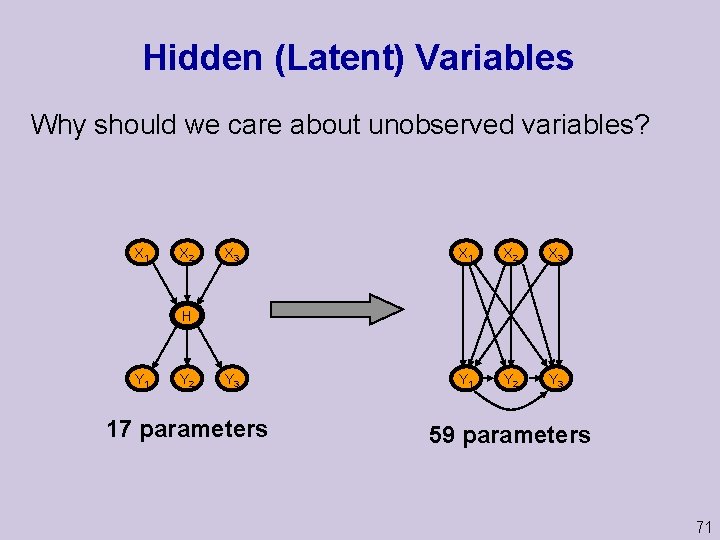
Hidden (Latent) Variables Why should we care about unobserved variables? X 1 X 2 X 3 Y 3 Y 1 Y 2 Y 3 H Y 1 Y 2 17 parameters 59 parameters 71
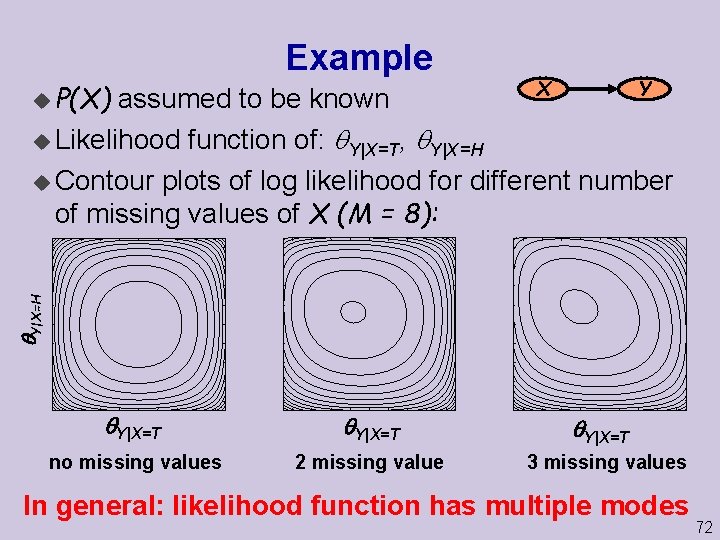
u P(X) Example X Y Y|X=H assumed to be known u Likelihood function of: Y|X=T, Y|X=H u Contour plots of log likelihood for different number of missing values of X (M = 8): Y|X=T no missing values 2 missing value Y|X=T 3 missing values In general: likelihood function has multiple modes 72

Incomplete Data u In the presence of incomplete data, the likelihood can have multiple maxima H Y Example: u We can rename the values of hidden variable H u If H has two values, likelihood has two maxima u In practice, many local maxima 73
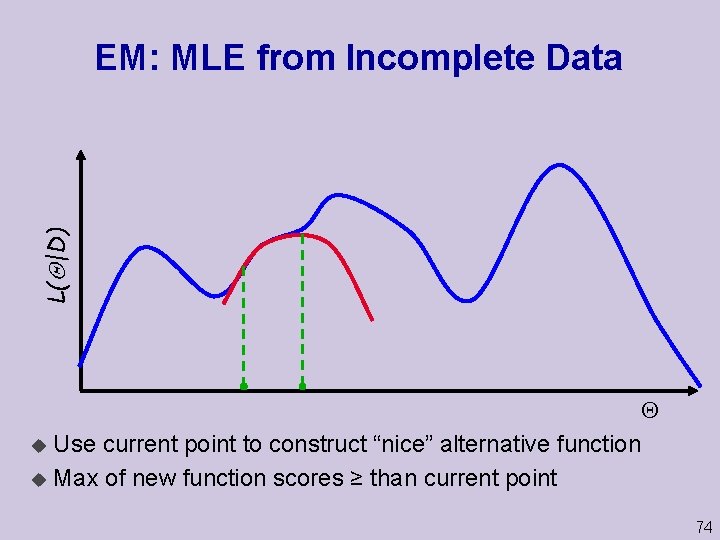
L( |D) EM: MLE from Incomplete Data u Use current point to construct “nice” alternative function u Max of new function scores ≥ than current point 74

Expectation Maximization (EM) u. A general purpose method for learning from incomplete data Intuition: u If we had true counts, we could estimate parameters u But with missing values, counts are unknown u We “complete” counts using probabilistic inference based on current parameter assignment u We use completed counts as if real to re-estimate parameters 75
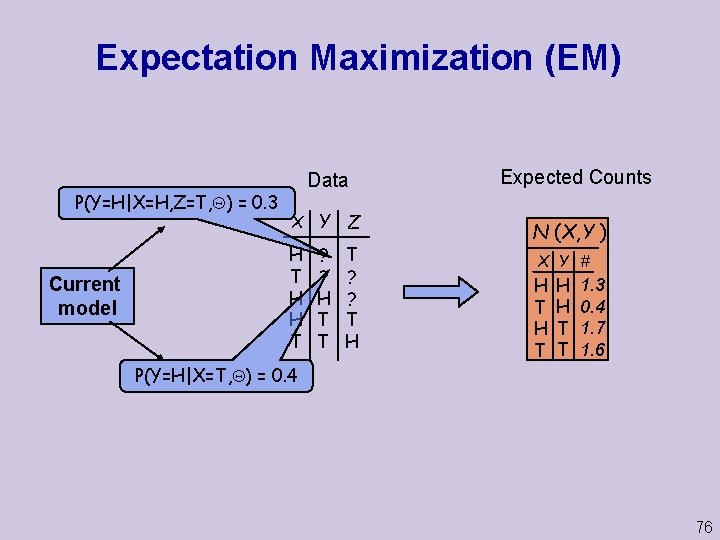
Expectation Maximization (EM) P(Y=H|X=H, Z=T, ) = 0. 3 Current model Data X Y Z H T H H T T ? ? T H ? ? H T T Expected Counts N (X, Y ) X Y # H T H H T T 1. 3 0. 4 1. 7 1. 6 P(Y=H|X=T, ) = 0. 4 76
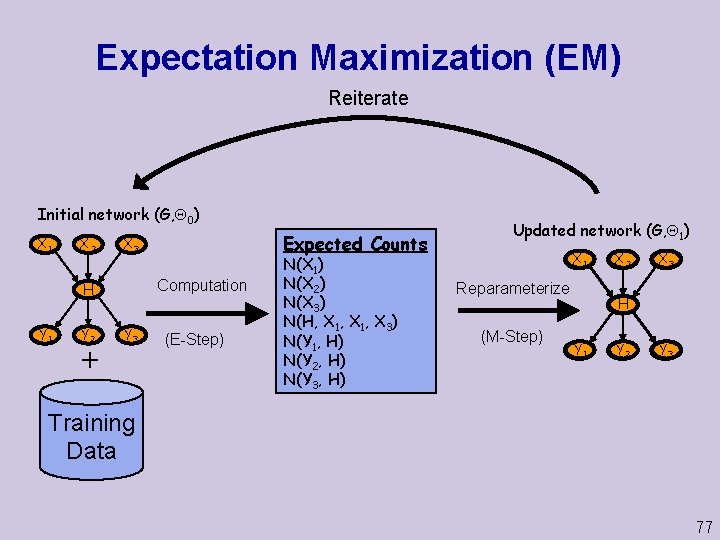
Expectation Maximization (EM) Reiterate Initial network (G, 0) X 1 X 2 Computation H Y 1 Y 2 Expected Counts X 3 Y 3 (E-Step) N(X 1) N(X 2) N(X 3) N(H, X 1, X 3) N(Y 1, H) N(Y 2, H) N(Y 3, H) Updated network (G, 1) X 1 Reparameterize (M-Step) X 2 X 3 H Y 1 Y 2 Y 3 Training Data 77

Expectation Maximization (EM) Formal Guarantees: u L( 1: D) L( 0: D) l Each iteration improves the likelihood u If 1 = 0 , then 0 is a stationary point of L( : D) l Usually, this means a local maximum 78

Expectation Maximization (EM) Computational bottleneck: u Computation of expected counts in E-Step l Need to compute posterior for each unobserved variable in each instance of training set l All posteriors for an instance can be derived from one pass of standard BN inference 79

Summary: Parameter Learning with Incomplete Data u Incomplete data makes parameter estimation hard u Likelihood function l Does not have closed form l Is multimodal u Finding max likelihood parameters: EM l Gradient ascent u Both exploit inference procedures for Bayesian networks to compute expected sufficient statistics l 80

Incomplete Data: Structure Scores Recall, Bayesian score: With incomplete data: u Cannot evaluate marginal likelihood in closed form u We have to resort to approximations: l l Evaluate score around MAP parameters Need to find MAP parameters (e. g. , EM) 81
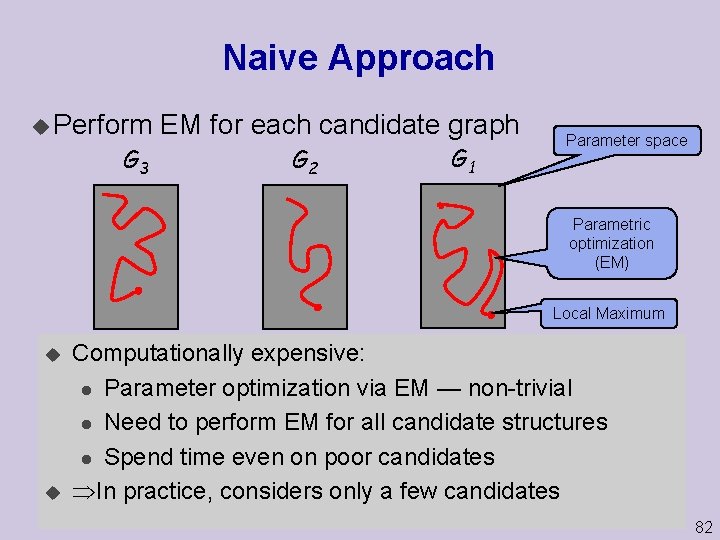
Naive Approach u Perform G 3 EM for each candidate graph G 2 G 1 Parameter space Parametric optimization (EM) Local Maximum u u Computationally expensive: Gn G 4 l Parameter optimization via EM — non-trivial l Need to perform EM for all candidate structures l Spend time even on poor candidates In practice, considers only a few candidates 82

Structural EM Recall, in complete data we had l Decomposition efficient search Idea: u Instead of optimizing the real score… u Find decomposable alternative score u Such that maximizing new score improvement in real score 83

Structural EM Idea: u Use current model to help evaluate new structures Outline: u Perform search in (Structure, Parameters) space u At each iteration, use current model for finding either: l Better scoring parameters: “parametric” EM step or l Better scoring structure: “structural” EM step 84
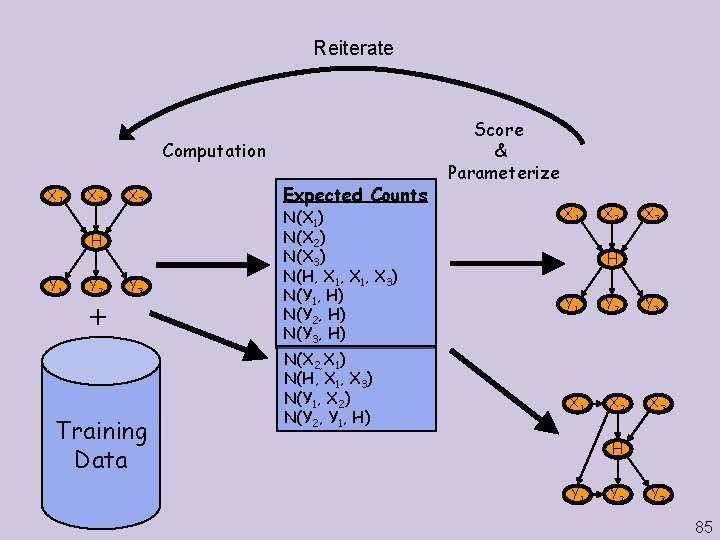
Reiterate Computation X 1 X 2 X 3 H Y 1 Y 2 Y 3 Training Data Expected Counts N(X 1) N(X 2) N(X 3) N(H, X 1, X 3) N(Y 1, H) N(Y 2, H) N(Y 3, H) N(X 2, X 1) N(H, X 1, X 3) N(Y 1, X 2) N(Y 2, Y 1, H) Score & Parameterize X 1 X 2 X 3 H Y 1 X 1 Y 2 X 2 Y 3 X 3 H Y 1 Y 2 Y 3 85
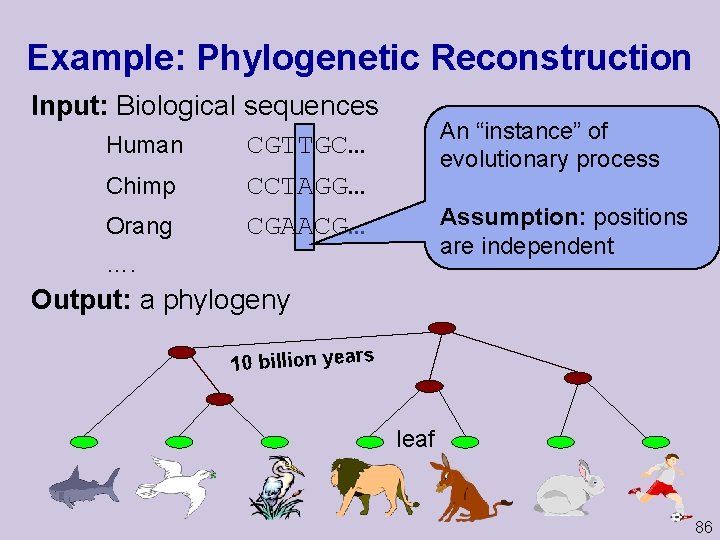
Example: Phylogenetic Reconstruction Input: Biological sequences Human CGTTGC… Chimp CCTAGG… Orang CGAACG… An “instance” of evolutionary process Assumption: positions are independent …. Output: a phylogeny 10 billion years leaf 86
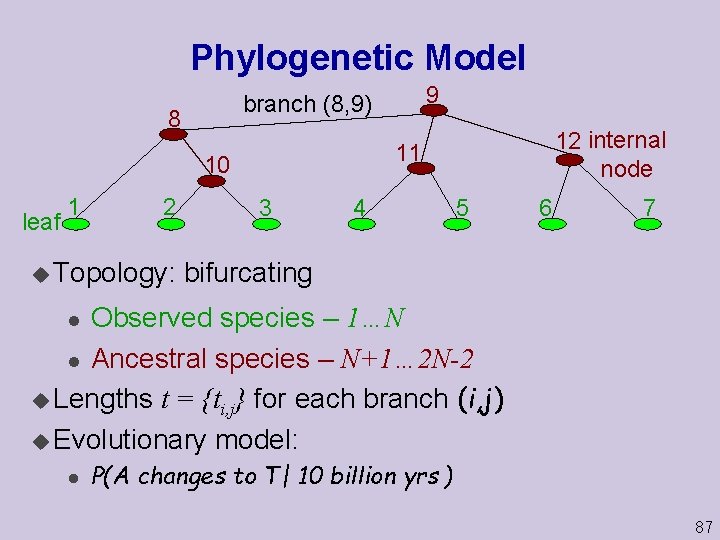
Phylogenetic Model 8 leaf 2 u Topology: 12 internal node 11 10 1 9 branch (8, 9) 3 4 5 6 7 bifurcating Observed species – 1…N l Ancestral species – N+1… 2 N-2 u Lengths t = {ti, j} for each branch (i, j) u Evolutionary model: l l P(A changes to T| 10 billion yrs ) 87

Phylogenetic Tree as a Bayes Net u Variables: Letter at each position for each species l Current day species – observed l Ancestral species - hidden u BN Structure: Tree topology u BN Parameters: Branch lengths (time spans) Main problem: Learn topology If ancestral were observed easy learning problem (learning trees) 88
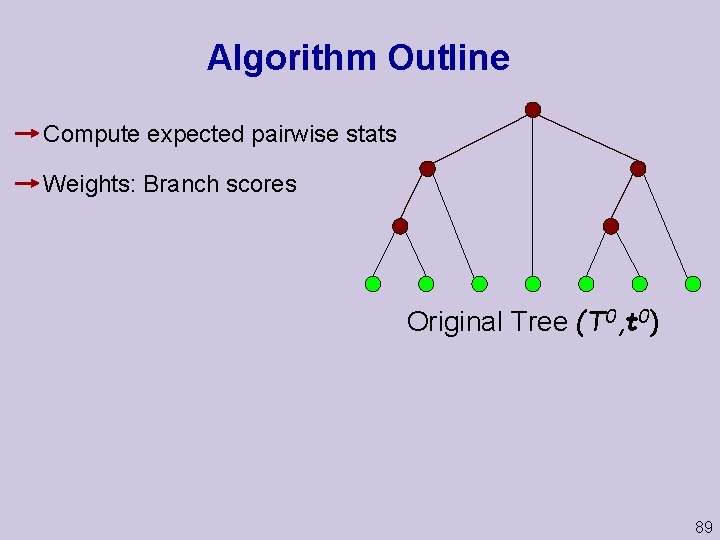
Algorithm Outline Compute expected pairwise stats Weights: Branch scores Original Tree (T 0, t 0) 89
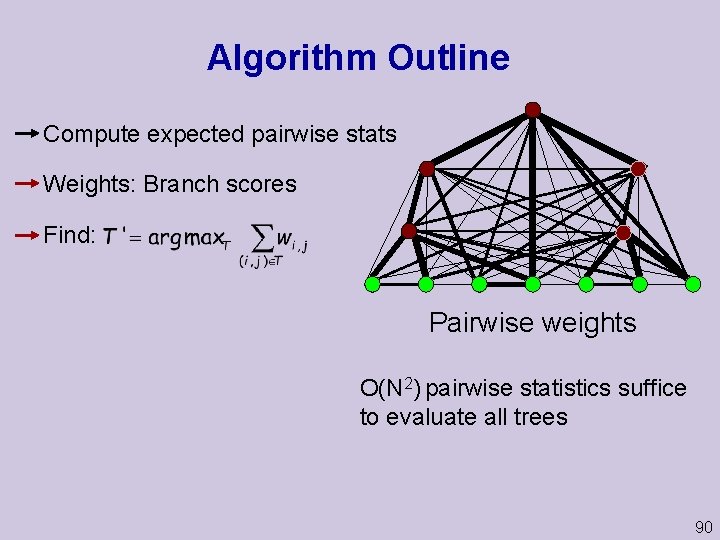
Algorithm Outline Compute expected pairwise stats Weights: Branch scores Find: Pairwise weights O(N 2) pairwise statistics suffice to evaluate all trees 90
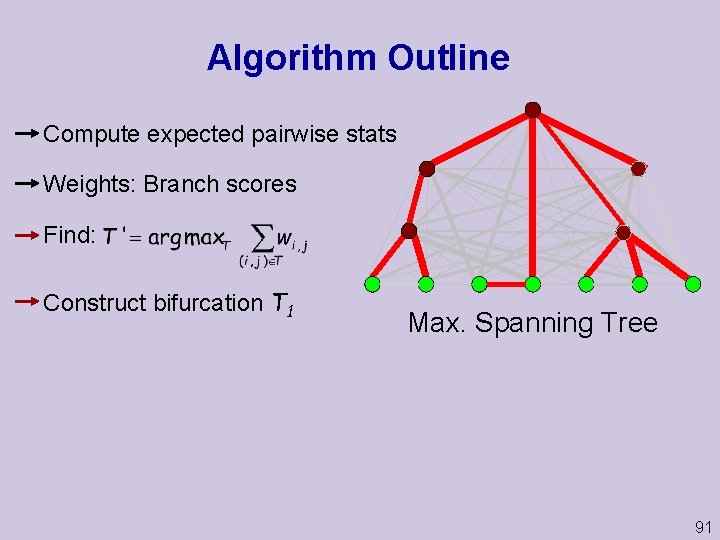
Algorithm Outline Compute expected pairwise stats Weights: Branch scores Find: Construct bifurcation T 1 Max. Spanning Tree 91
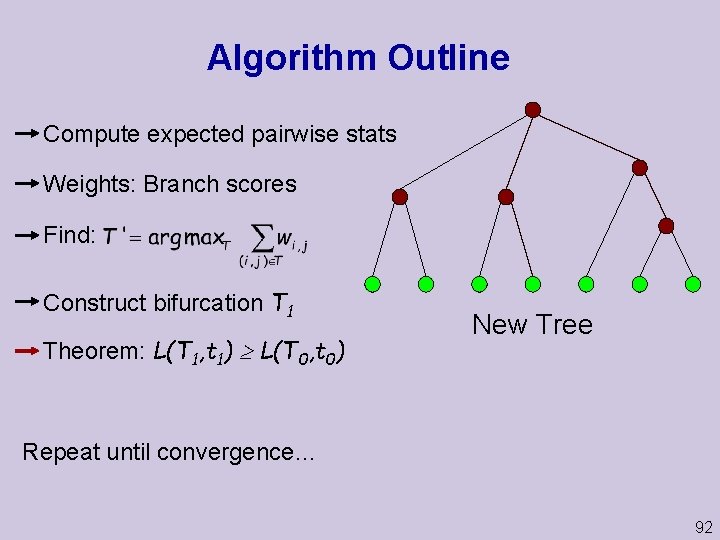
Algorithm Outline Compute expected pairwise stats Weights: Branch scores Find: Construct bifurcation T 1 Theorem: L(T 1, t 1) L(T 0, t 0) New Tree Repeat until convergence… 92

Real Life Data Lysozyme c Mitochondrial genomes 34 # sequences 43 # pos 122 3, 578 -2, 916. 2 -74, 227. 9 -2, 892. 1 -70, 533. 5 0. 19 1. 03 Traditional approach Log. Structural EM likelihood Approach Difference per position Each position twice as likely 93

Overview u Introduction u Parameter Estimation u Model Selection u Structure Discovery u Incomplete Data u Learning from Structured Data 94

Bayesian Networks: Problem Bayesian nets use propositional representation u Real world has objects, related to each other u Intelligence Difficulty Grade 95
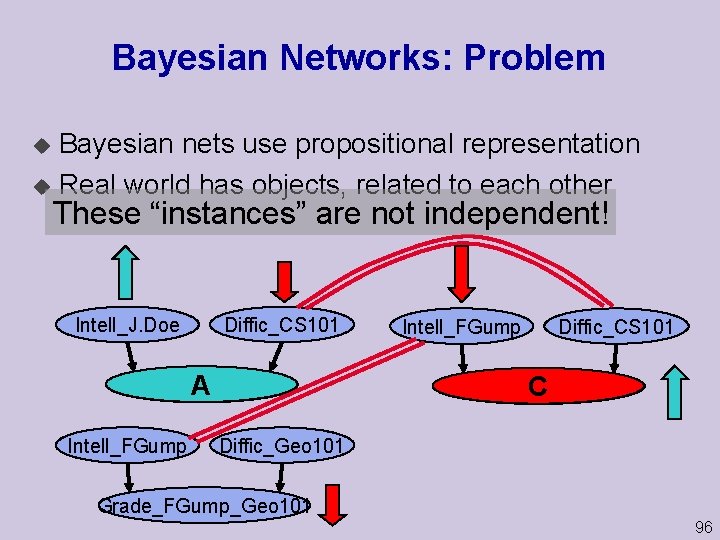
Bayesian Networks: Problem Bayesian nets use propositional representation u Real world has objects, related to each other u These “instances” are not independent! Intell_J. Doe Diffic_CS 101 Grade_JDoe_CS 101 A Intell_FGump Diffic_CS 101 Grade_FGump_CS 101 C Diffic_Geo 101 Grade_FGump_Geo 101 96
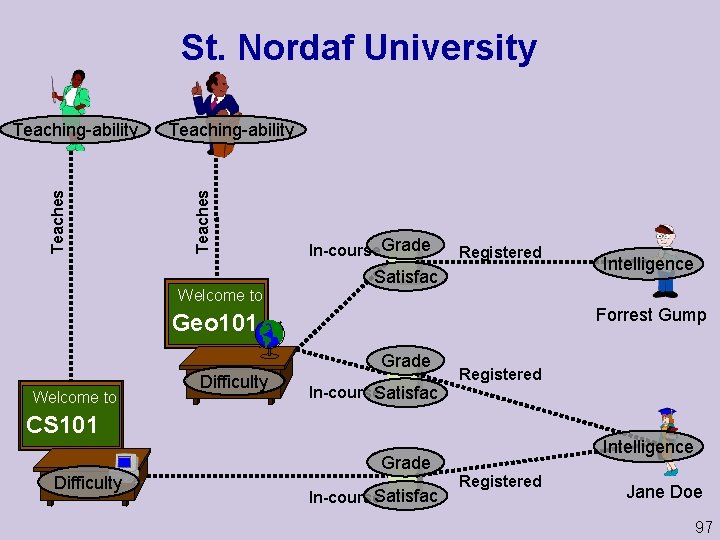
St. Nordaf University Teaching-ability Teaches Teaching-ability Welcome to In course. Grade Registered Satisfac Forrest Gump Geo 101 Grade Welcome to Difficulty Registered In course. Satisfac CS 101 Difficulty Intelligence Grade In course. Satisfac Intelligence Registered Jane Doe 97
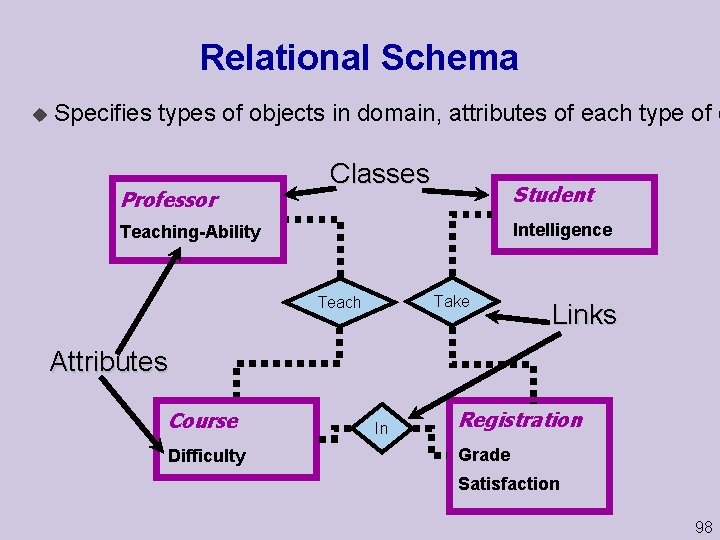
Relational Schema u Specifies types of objects in domain, attributes of each type of o Professor Classes Student Intelligence Teaching Ability Take Teach Links Attributes Course Difficulty In Registration Grade Satisfaction 98

Representing the Distribution u Many l possible worlds for a given university All possible assignments of all attributes of all objects u Infinitely l many potential universities Each associated with a very different set of worlds Need to represent infinite set of complex distributions 99
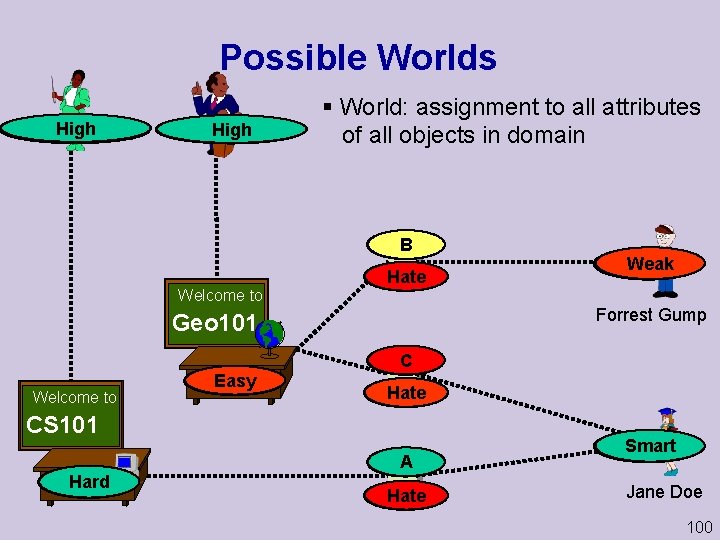
Possible Worlds Prof. Jones High Low Teaching-ability Prof. Smith High Teaching-ability § World: assignment to all attributes of all objects in domain B Grade C Welcome to Hate Satisfac Like Forrest Gump Geo 101 Welcome to Easy Difficulty C B Grade Hate Satisfac CS 101 Hard Easy Difficulty Weak Intelligence A Grade Hate Like Satisfac Smart Intelligence Jane Doe 100
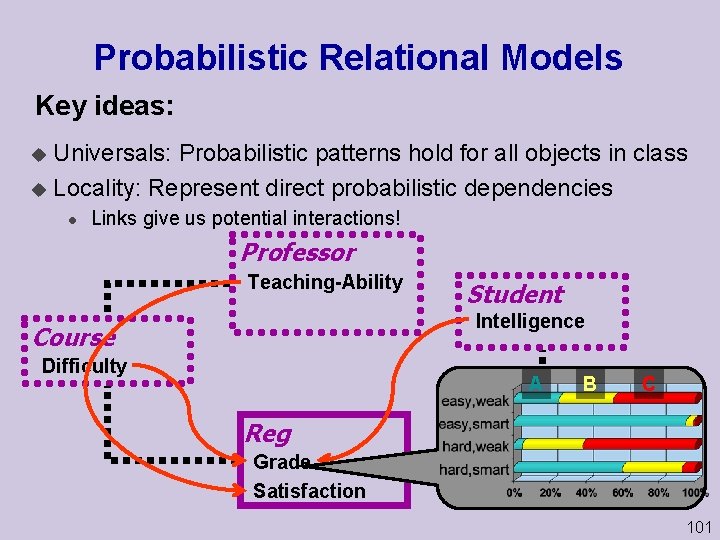
Probabilistic Relational Models Key ideas: Universals: Probabilistic patterns hold for all objects in class u Locality: Represent direct probabilistic dependencies u l Links give us potential interactions! Professor Teaching Ability Student Intelligence Course Difficulty A B C Reg Grade Satisfaction 101
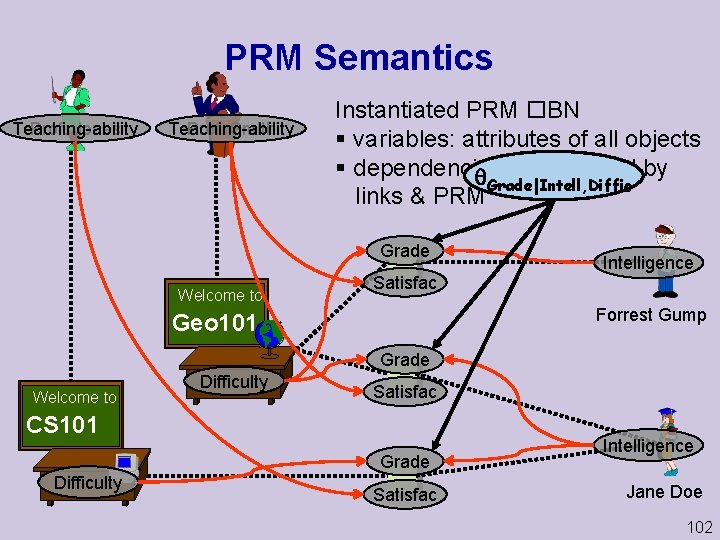
PRM Semantics Prof. Jones Teaching-ability Prof. Smith Teaching-ability Instantiated PRM BN § variables: attributes of all objects § dependencies: determined by Grade|Intell, Diffic links & PRM Grade Welcome to Satisfac Intelligence Forrest Gump Geo 101 Grade Welcome to Difficulty Satisfac CS 101 Grade Difficulty Satisfac Intelligence Jane Doe 102
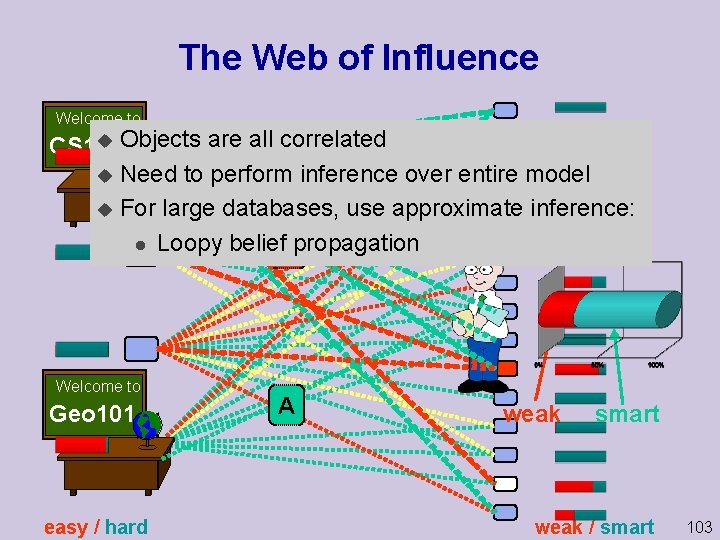
The Web of Influence Welcome to u Objects are all correlated CS 101 u Need to perform inference over entire model u For large databases, use approximate inference: l Loopy belief. Cpropagation Welcome to Geo 101 easy / hard A weak smart weak / smart 103
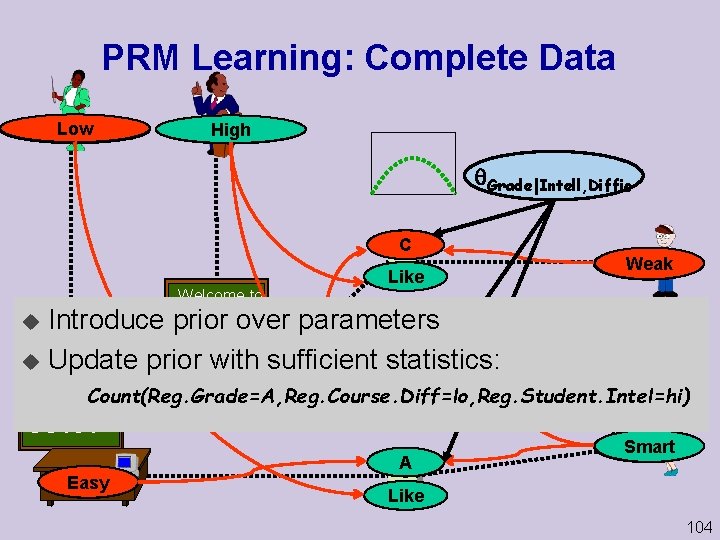
PRM Learning: Complete Data Prof. Jones Low Teaching-ability Prof. Smith High Teaching-ability Grade|Intell, Diffic Grade C Welcome to Satisfac Like Weak Intelligence Introduce Geo 101 prior over parameters u Entire database is single “instance” B u Update prior with sufficient Grade statistics: u Parameters Easyused many times in instance Difficulty u Hate Satisfac Count(Reg. Grade=A, Reg. Course. Diff=lo, Reg. Student. Intel=hi) Welcome to CS 101 Easy Difficulty A Grade Smart Intelligence Like Satisfac 104
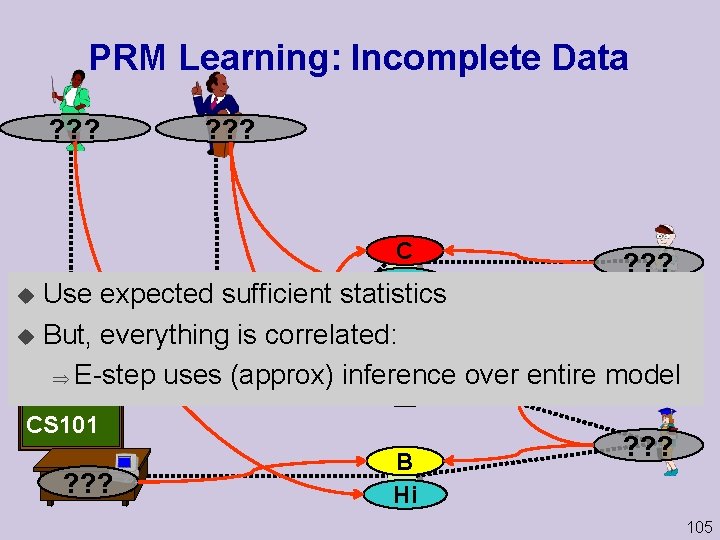
PRM Learning: Incomplete Data ? ? ? C Hi ? ? ? Welcome to Use expected sufficient statistics Geo 101 u But, everything is correlated: A E-step uses (approx) inference over entire model ? ? ? u Welcome to Low CS 101 ? ? ? B Hi ? ? ? 105
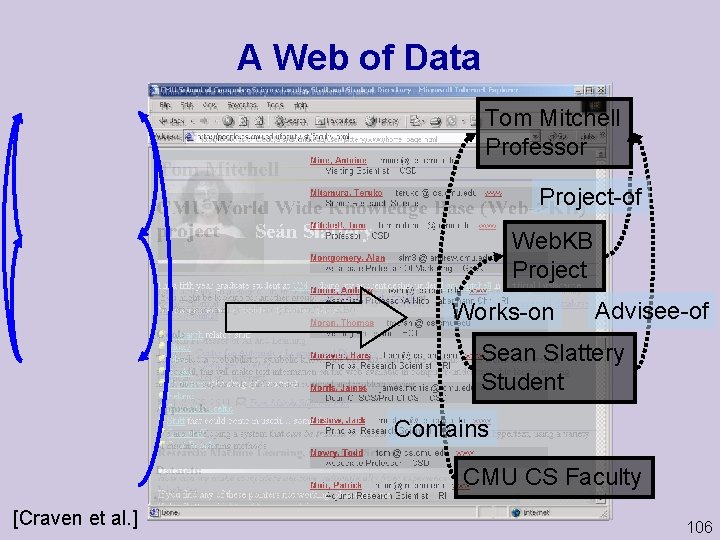
A Web of Data Tom Mitchell Professor Project-of Web. KB Project Works-on Advisee-of Sean Slattery Student Contains CMU CS Faculty [Craven et al. ] 106
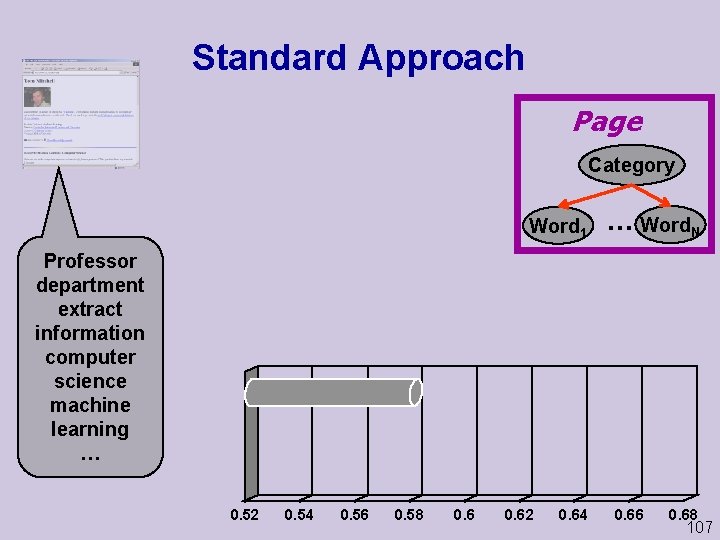
Standard Approach Page Category Word 1 . . . Word. N Professor department extract information computer science machine learning … 0. 52 0. 54 0. 56 0. 58 0. 62 0. 64 0. 66 0. 68 107
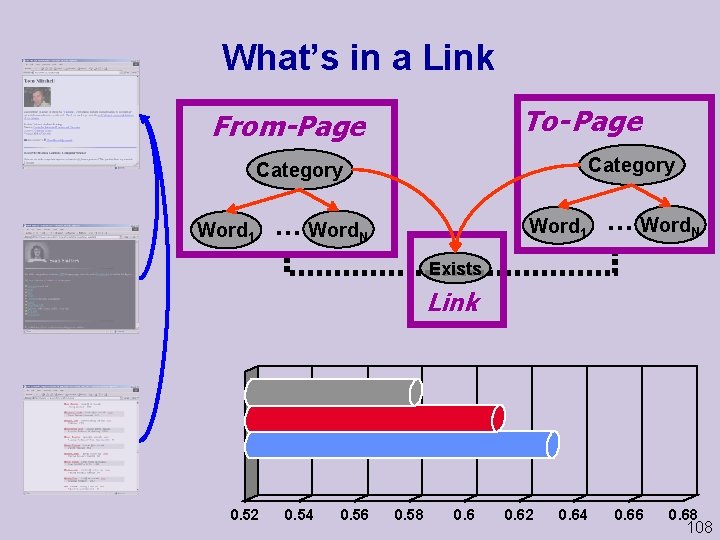
What’s in a Link To- Page From-Page Category Word 1 . . . Word. N Exists Link 0. 52 0. 54 0. 56 0. 58 0. 62 0. 64 0. 66 0. 68 108
Discovering Hidden Concepts Internet Movie Database http: //www. imdb. com 109
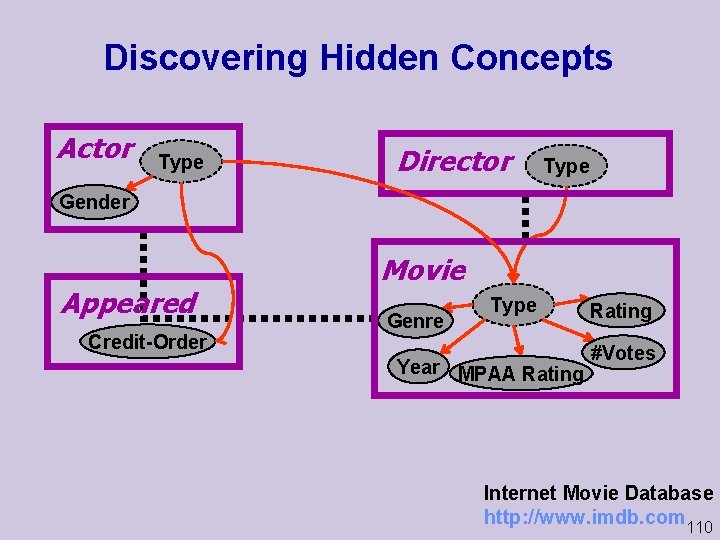
Discovering Hidden Concepts Actor Type Director Type Gender Appeared Credit Order Movie Genre Type Year MPAA Rating #Votes Internet Movie Database http: //www. imdb. com 110

Web of Influence, Yet Again Movies Actors Directors Wizard of Oz Cinderella Sound of Music The Love Bug Pollyanna The Parent Trap Mary Poppins Swiss Family Robinson Sylvester Stallone Bruce Willis Harrison Ford Steven Seagal Kurt Russell Kevin Costner Jean Claude Van Damme Arnold Schwarzenegger Alfred Hitchcock Stanley Kubrick David Lean Milos Forman Terry Gilliam Francis Coppola Terminator 2 Batman Forever Mission: Impossible Golden. Eye Starship Troopers Hunt for Red October Anthony Hopkins Robert De Niro Tommy Lee Jones Harvey Keitel Morgan Freeman Gary Oldman Steven Spielberg Tim Burton Tony Scott James Cameron John Mc. Tiernan Joel Schumacher … … … 111

Conclusion u Many distributions have combinatorial dependency structure u Utilizing this structure is good u Discovering this structure has implications: l To density estimation l To knowledge discovery u Many applications l Medicine l Biology l Web 112

The END Thanks to Gal Elidan u Lise Getoor u Moises Goldszmidt u Matan Ninio u Dana Pe’er u Eran Segal u Ben Taskar u Slides will be available from: http: //www. cs. huji. ac. il/~nir/ http: //robotics. stanford. edu/~koller/ 113
- Slides: 113