Leaf area vs massproportionality of leaf traits within











- Slides: 17

Leaf area- vs. mass-proportionality of leaf traits within canopies and across species: patterns and analytical consequences Jeanne L. D. Osnas, Jeremy W. Lichstein, Stephen W. Pacala, Peter B. Reich June 2013

• 300, 000 vascular plant species • global vegetation models: 5 -10 plant functional types Foley et al. 1996 Barthlott et al. 1999

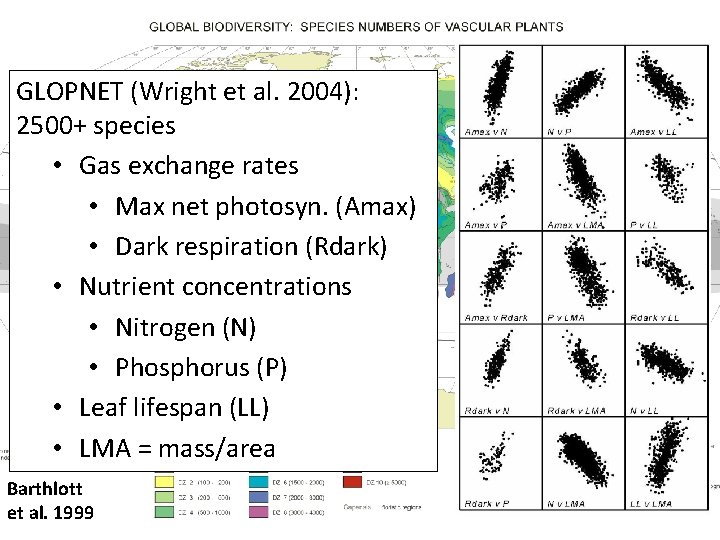
GLOPNET (Wright et al. 2004): 2500+ species • Gas exchange rates • Max net photosyn. (Amax) • Dark respiration (Rdark) • Nutrient concentrations • Nitrogen (N) • Phosphorus (P) • Leaf lifespan (LL) • LMA = mass/area Barthlott et al. 1999

Area-normalized Mass-normalized Xmass = Xarea/LMA X = Amax, Rdark, N, P GLOPNET

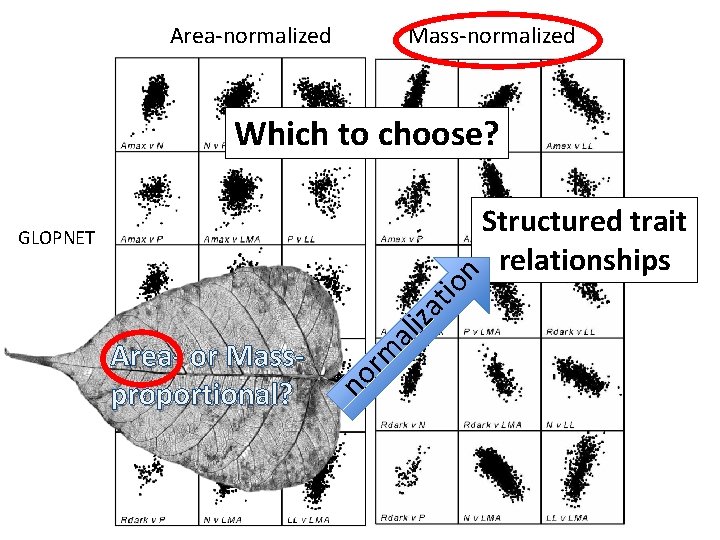
Area-normalized Mass-normalized Which to choose? no Area- or Massproportional? rm al iza tio n GLOPNET Structured trait relationships

Trait area- and mass-proportionality across species Total leaf trait i: Xik = (Massk μMi + Areak μAi)εik Mass-normalized: XMik = (μMi + LMAk-1 μAi)εik Area-normalized: XAik = (LMAkμMi + μAi)εik μMi, μAi constant across species εik = random variable (interspecific variation) Osnas et al. (2013) Science

Quantify trait area- and massproportionality across species Total leaf trait i: Xik = (Massk μMi + Areak μAi)εik Mass-normalized: XMik = (μMi + LMAk-1 μAi)εik Area-normalized: XAik = (LMAkμMi + μAi)εik μMi, μAi constant across species εik = random variable (interspecific variation) Osnas et al. (2013) Science

Quantify trait area- and massproportionality across species Total leaf trait i: Xik = (Massk μMi + Areak μAi)εik Mass-normalized: XMik = (μMi + LMAk-1 μAi)εik Area-normalized: XAik = (LMAkμMi + μAi)εik μMi, μAi constant across species εik = random variable (interspecific variation) Osnas et al. (2013) Science

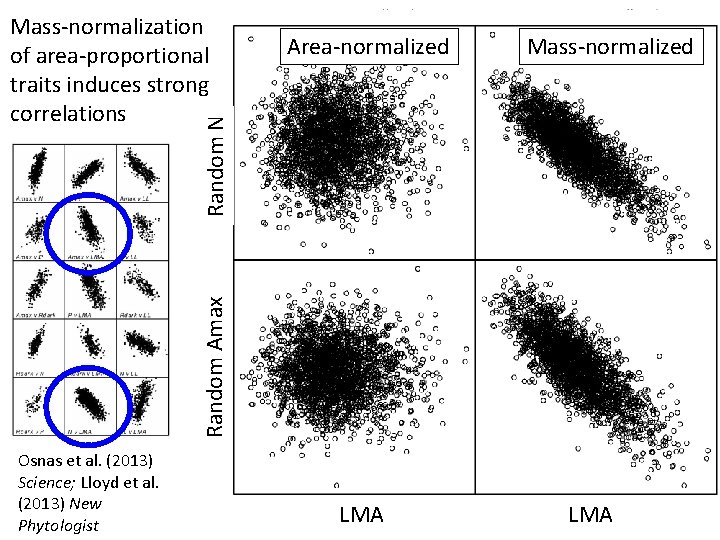
Mass-normalization of area-proportional traits induces strong correlations Mass-normalized Random N Area-normalized Random Amax LMA Random N = random draws from lognormal distribution parameterized with GLOPNET Narea GLOPNET LMA Osnas et al. (2013) Science; Lloyd et al. (2013) New Phytologist “area-proportional” LMA

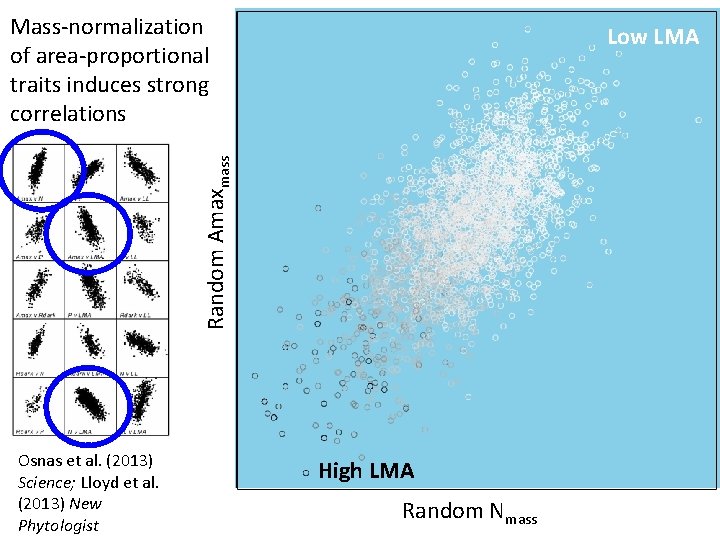
Mass-normalization of area-proportional traits induces strong correlations Random Amaxmass Low LMA Osnas et al. (2013) Science; Lloyd et al. (2013) New Phytologist High LMA Random Nmass

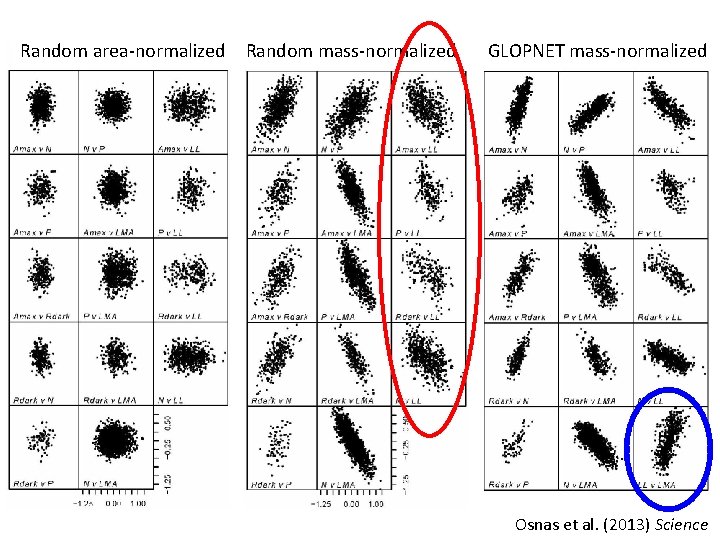
Random area-normalized Random mass-normalized GLOPNET mass-normalized Osnas et al. (2013) Science

How do we know if traits area-proportional, mass-proportional, or something in between? • Quantify trait mass-proportionality • Across species in the global flora • Normalization-independent trait relationships • Discuss consequences

Quantify trait area- and mass-proportionality across species Total leaf: Area-normalized: Si = massproportionality across species Mass-normalized: Area-normalized: log(XAik) = Ii + Si log(LMAk) + nik Mass-normalized: log(XMik) = Ii + (Si − 1) log(LMAk) + nik Ci, Si constant across species εik = distribution of interspecific variation nik is trait variation conditional on LMA (normalizationindependent) Osnas et al. (2013) Science

Quantify trait area- and mass-proportionality across species Total leaf: Area-normalized: Si = massproportionality across species Mass-normalized: Purely area-proportional: Si = 0 Area-normalized: log(XAik) = Ii + nik Mass-normalized: log(XMik) = Ii − log(LMAk) + nik Ci, Si constant across species εik = distribution of interspecific variation Osnas et al. (2013) Science

Quantify trait area- and mass-proportionality across species Total leaf: Area-normalized: Si = massproportionality across species Mass-normalized: Area-normalized: log(XAik) = Ii + log(LMAk) + nik Mass-normalized: log(XMik) = Ii + nik Purely mass-proportional: Si = 1 Ci, Si constant across species εik = distribution of interspecific variation Osnas et al. (2013) Science

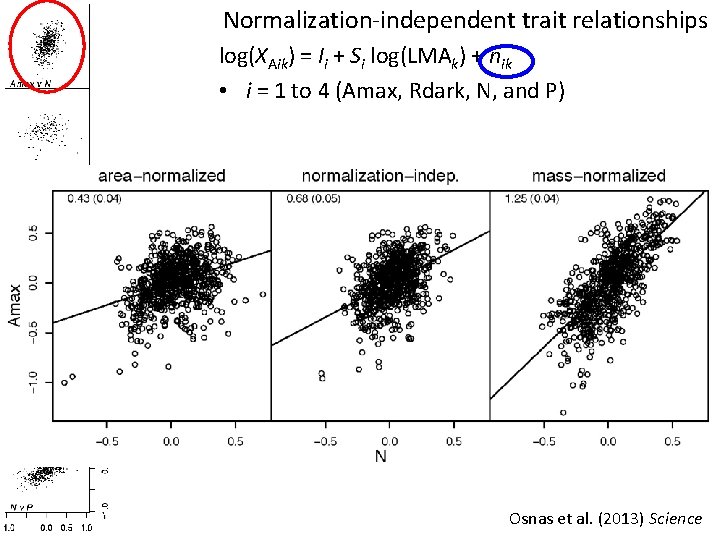
Normalization-independent trait relationships log(XAik) = Ii + Si log(LMAk) + nik • i = 1 to 4 (Amax, Rdark, N, and P) Osnas et al. (2013) Science

Traits are mostly area-proportional across species in the global flora, although N and Rdark have minor but significant massproportional components. Normalization by mass (substantially) or area (somewhat) can create potentially misleading structure in trait relationships – PC 1 of mass-normalized GLOPNET data ≈ LMA Using trait relationships – Functional diversity as a species continuum with at least 2 axes: • PC 1 of normalization-independent PCA • LMA • Maybe LL, other traits