LDBased Genotype and Haplotype Inference from LowCoverage Short

LD-Based Genotype and Haplotype Inference from Low-Coverage Short Sequencing Reads Ion Mandoiu Computer Science and Engineering Department University of Connecticut Joint work with S. Dinakar, J. Duitama, Y. Hernández, J. Kennedy, and Y. Wu

Outline n Introduction n Single SNP Genotype Calling n Multilocus Genotyping Problem n HMM-Posterior Algorithm n Experimental Results n Conclusion

Ultra-high throughput sequencing n Recent massively parallel sequencing technologies deliver orders of magnitude higher throughput compared to classic Sanger sequencing Roche/454 FLX Titanium 400 bp reads 400 Mb/10 h run ABI SOLi. D 2. 0 25 -35 bp reads 3 -4 Gb/6 day run Illumina Genome Analyzer II 35 -50 bp reads 1. 5 Gb/2. 5 day run Helicos Heli. Scope 25 -55 bp reads >1 Gb/day

UHTS applications n n UHTS is a transformative technology Numerous applications besides de novo genome sequencing: n n n n RNA-Seq Non-coding RNAs Ch. IP-Seq Epigenetics Structural variation Metagenomics Paleogenomics …
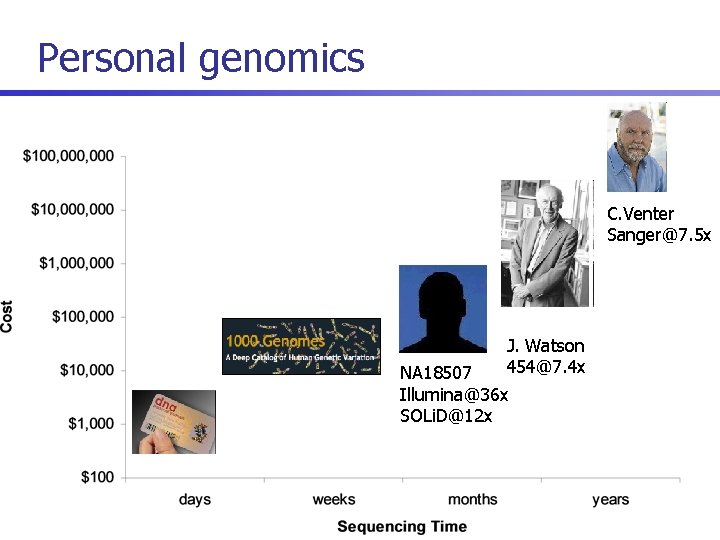
Personal genomics C. Venter Sanger@7. 5 x J. Watson 454@7. 4 x NA 18507 Illumina@36 x SOLi. D@12 x

Challenges for medical applications of sequencing n Sequencing provides single-base resolution of genetic variation (SNPs, CNVs, genome rearrangements) n n n However, interpretation requires determination of both alleles at variable loci This is limited by coverage depth due to random nature of shotgun sequencing For the Venter and Watson genomes (both sequenced at ~7. 5 x average coverage), comparison with SNP genotyping chips has shown only ~75% accuracy for sequencing based calls of heterozygous SNPs [Levy et al 07, Wheeler et al 08]
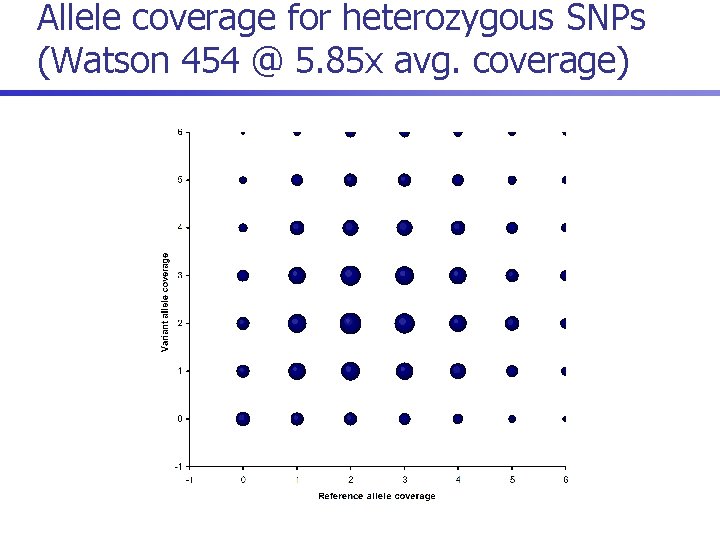
Allele coverage for heterozygous SNPs (Watson 454 @ 5. 85 x avg. coverage)
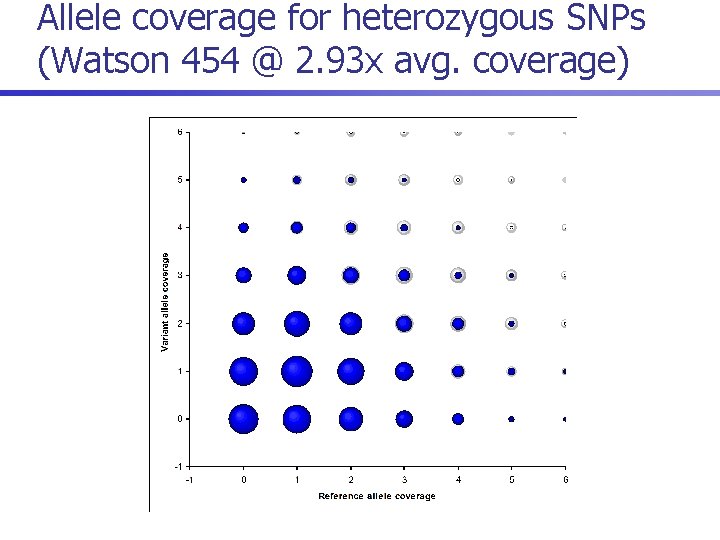
Allele coverage for heterozygous SNPs (Watson 454 @ 2. 93 x avg. coverage)
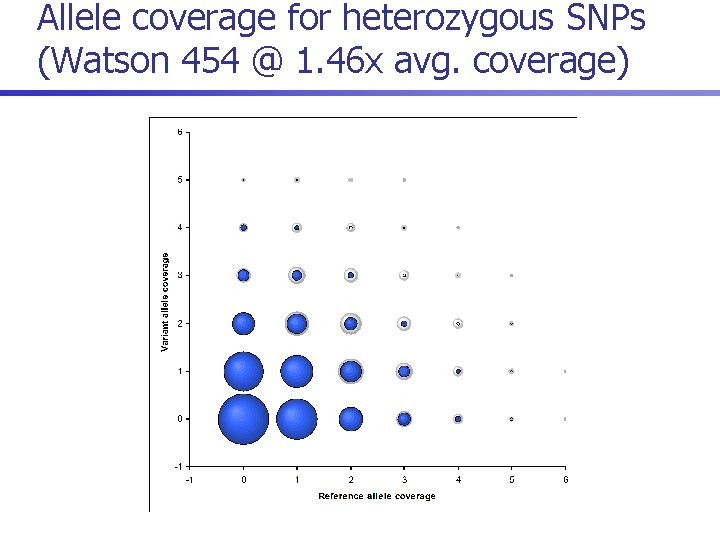
Allele coverage for heterozygous SNPs (Watson 454 @ 1. 46 x avg. coverage)
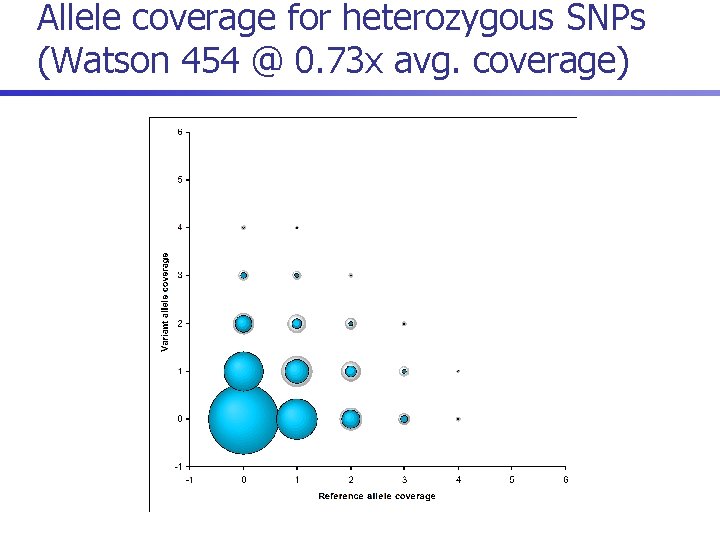
Allele coverage for heterozygous SNPs (Watson 454 @ 0. 73 x avg. coverage)

Prior work n Most work devoted to de novo variation discovery from sequencing data, e. g. , SNPs, CNVs n n Unlike genotying known variation, de novo discovery requires very stringent detection criteria Prior genotyping methods are based on allele coverage n n [Levy et al 07] and [Wheeler et al 08] require that each allele be covered by at least 2 reads in order to be called Combined with hypothesis testing based on the binomial distribution when calling hets n n Binomial probability for the observed number of 0 and 1 alleles must be at least 0. 01 [Wendl&Wilson 08] generalize coverage methods to allow an arbitrary minimum allele coverage k
![What coverage is required? n [Wendl&Wilson 08] estimate that 21 x coverage will be What coverage is required? n [Wendl&Wilson 08] estimate that 21 x coverage will be](http://slidetodoc.com/presentation_image/201e20431914aae3b7c7e088d6026c8a/image-12.jpg)
What coverage is required? n [Wendl&Wilson 08] estimate that 21 x coverage will be required for sequencing of normal tissue samples based on idealized theory that “neglects any heuristic inputs”

Do heuristic inputs help? n We propose methods incorporating additional sources of information: n n n Quality scores reflecting uncertainty in sequencing data Allele/genotype frequency and linkage disequilibrium (LD) info extracted from a reference panel such as Hapmap Experimental results show significantly improved genotyping accuracy

Outline n Introduction n Single SNP Genotype Calling n Multilocus Genotyping Problem n HMM-Posterior Algorithm n Experimental Results n Conclusion

Basic notations n n Biallelic SNPs: 0 = major allele, 1 = minor allele SNP genotypes: 0/2 = homozygous major/minor, 1=heterozygous 012100120 Mapped reads with allele 0 Inferred genotypes Mapped reads with allele 1 Sequencing errors

Incorporating base call uncertainty n Let ri denote the set of mapped reads covering SNP locus i and ci =| ri | n n n For a read r in ri , r(i) denotes the allele observed at locus i If qr(i) is the phred quality score of r(i), the probability that r(i) is incorrect is given by Probability of observing read set ri conditional on Gi:

Single SNP genotype calling n Applying Bayes’ formula: n Where are genotype frequencies inferred from a representative panel

Outline n Introduction n Single SNP Genotype Calling n Multilocus Genotyping Problem n HMM-Posterior Algorithm n Experimental Results n Conclusion
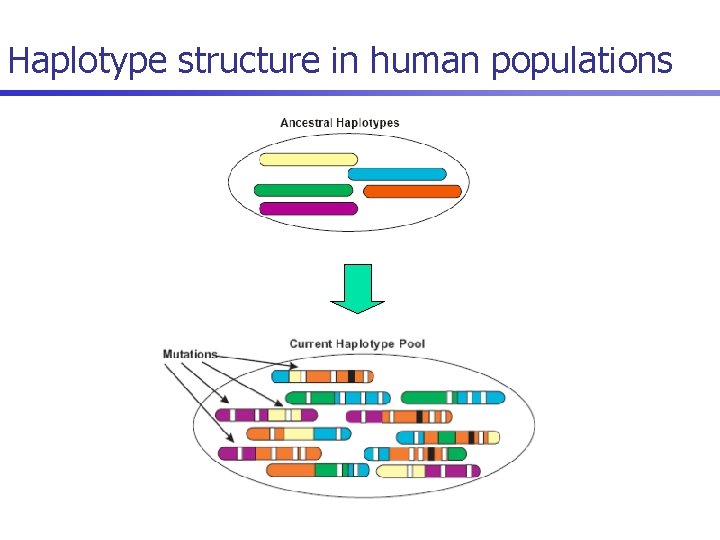
Haplotype structure in human populations
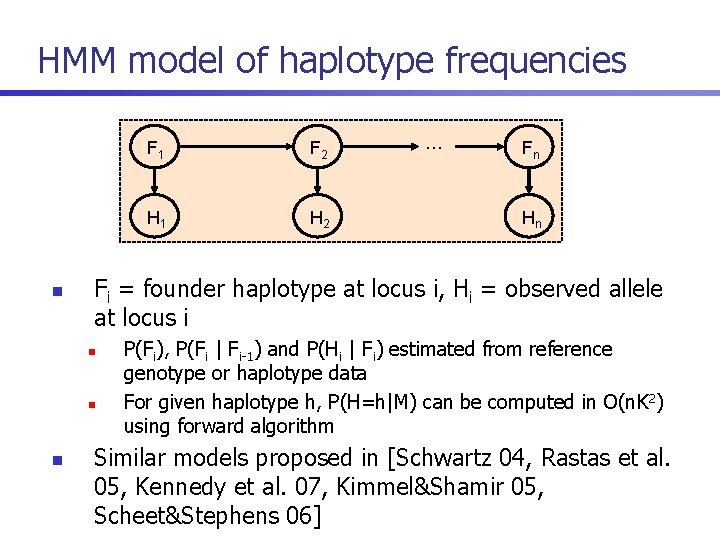
HMM model of haplotype frequencies n F 2 H 1 H 2 … Fn Hn Fi = founder haplotype at locus i, Hi = observed allele at locus i n n n F 1 P(Fi), P(Fi | Fi-1) and P(Hi | Fi) estimated from reference genotype or haplotype data For given haplotype h, P(H=h|M) can be computed in O(n. K 2) using forward algorithm Similar models proposed in [Schwartz 04, Rastas et al. 05, Kennedy et al. 07, Kimmel&Shamir 05, Scheet&Stephens 06]
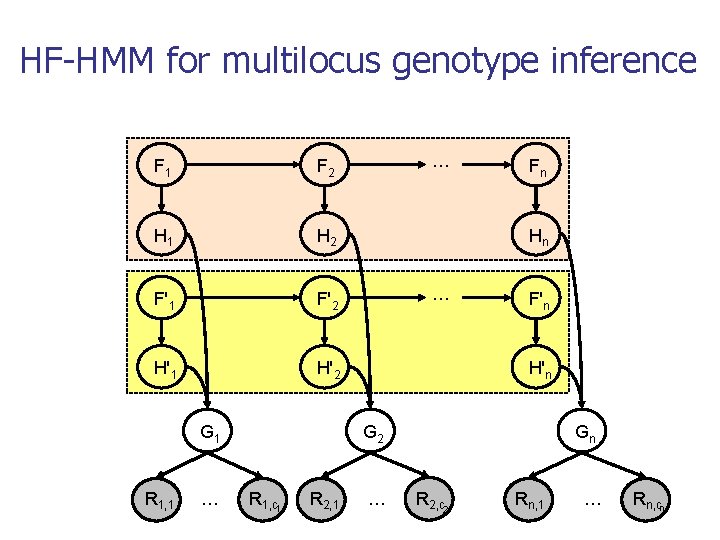
HF-HMM for multilocus genotype inference F 1 F 2 H 1 H 2 F'1 F'2 H'1 H'2 G 1 R 1, 1 … … Fn Hn … F'n H'n G 2 R 1, c 1 R 2, 1 … Gn R 2, c 2 Rn, 1 … Rn, cn

Model training n n P(f 1), P(f’ 1), P(fi+1|fi), P(f’i+1|f’i), P(hi|fi), P(h’i|f’i) trained using Baum-Welch algorithm on haplotypes inferred from the populations of origin for mother/father P(gi|hi, h’i) set to 1 if h+h’i=gi and to 0 otherwise n n This gives

Multilocus genotyping problem GIVEN: • Shotgun read sets r=(r 1, r 2, … , rn) • Quality scores • Trained HMM models representing LD in populations of origin for mother/father FIND: • Multilocus genotype g*=(g*1, g*2, …, g*n) with maximum posterior probability, i. e. , g*=argmaxg P(g | r)

Computational complexity of MGP Theorem: maxg. P(g | r) cannot be approximated within unless ZPP=NP Idea: reduction from the clique problem

Outline n Introduction n Single SNP Genotype Calling n Multilocus Genotyping Problem n HMM-Posterior Algorithm n Experimental Results n Conclusion

Posterior decoding algorithm 1. For each i = 1. . n, compute 2. Return
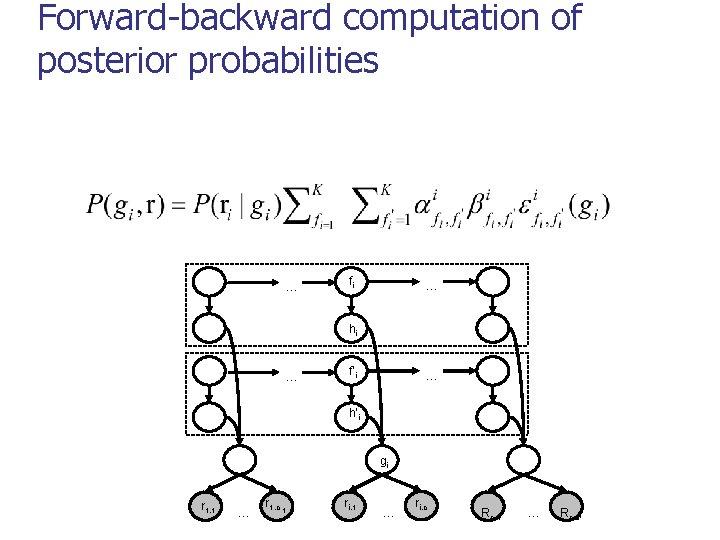
Forward-backward computation of posterior probabilities … fi … hi … f’i … h’i gi r 1, 1 … r 1, c 1 ri, 1 … ri, c i Rn, 1 … Rn, cn
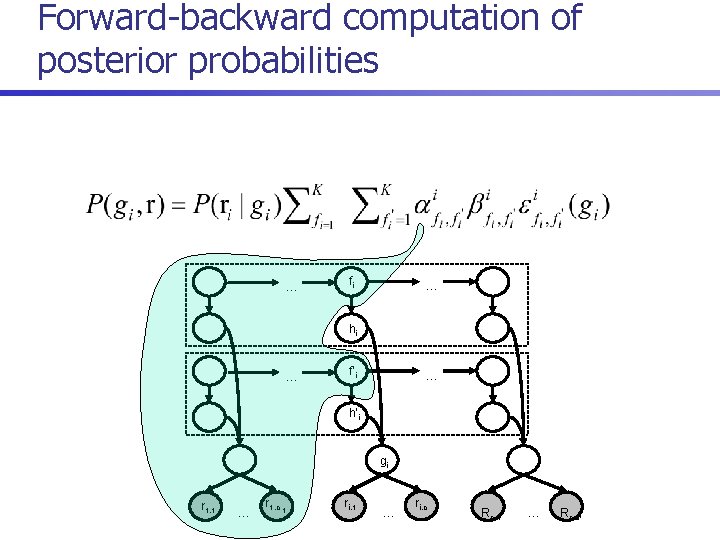
Forward-backward computation of posterior probabilities … fi … hi … f’i … h’i gi r 1, 1 … r 1, c 1 ri, 1 … ri, c i Rn, 1 … Rn, cn
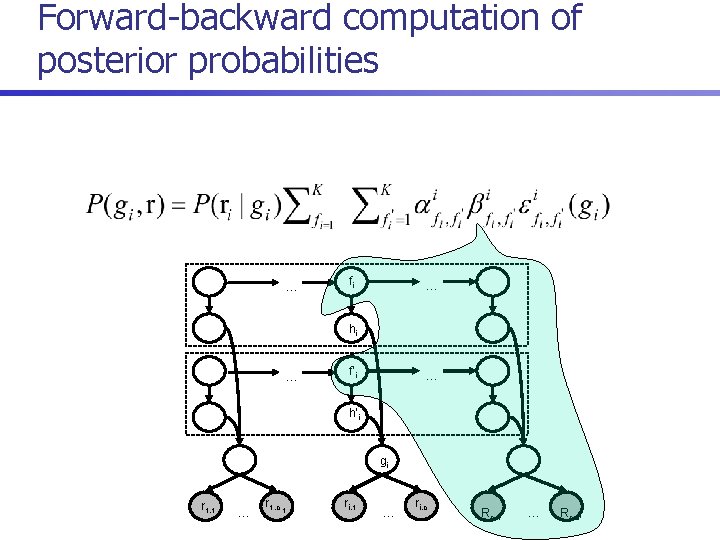
Forward-backward computation of posterior probabilities … fi … hi … f’i … h’i gi r 1, 1 … r 1, c 1 ri, 1 … ri, c i Rn, 1 … Rn, cn
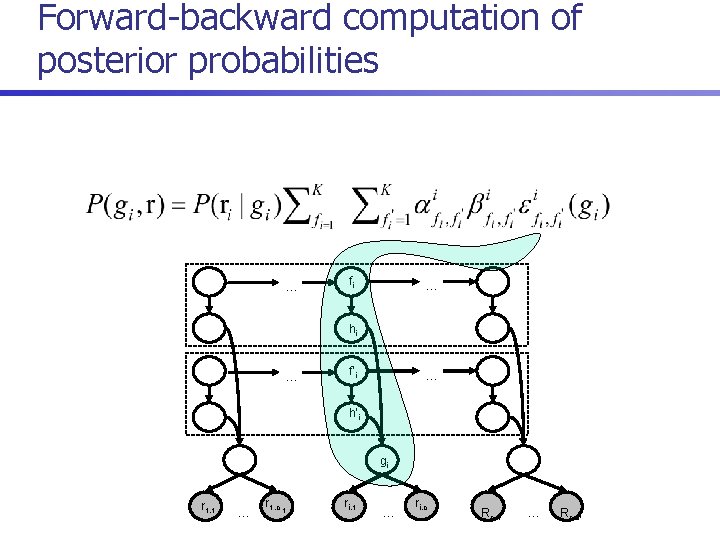
Forward-backward computation of posterior probabilities … fi … hi … f’i … h’i gi r 1, 1 … r 1, c 1 ri, 1 … ri, c i Rn, 1 … Rn, cn
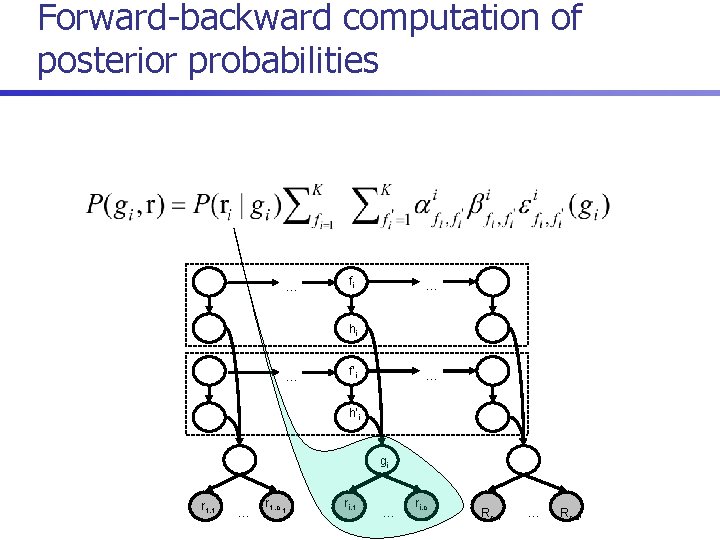
Forward-backward computation of posterior probabilities … fi … hi … f’i … h’i gi r 1, 1 … r 1, c 1 ri, 1 … ri, c i Rn, 1 … Rn, cn

Runtime n Direct recurrences for computing forward probabilities: n Runtime reduced to O(m+n. K 3) by reusing common terms: where

Outline n Introduction n Single SNP Genotype Calling n Multilocus Genotyping Problem n HMM-Posterior Algorithm n Experimental Results n Conclusion
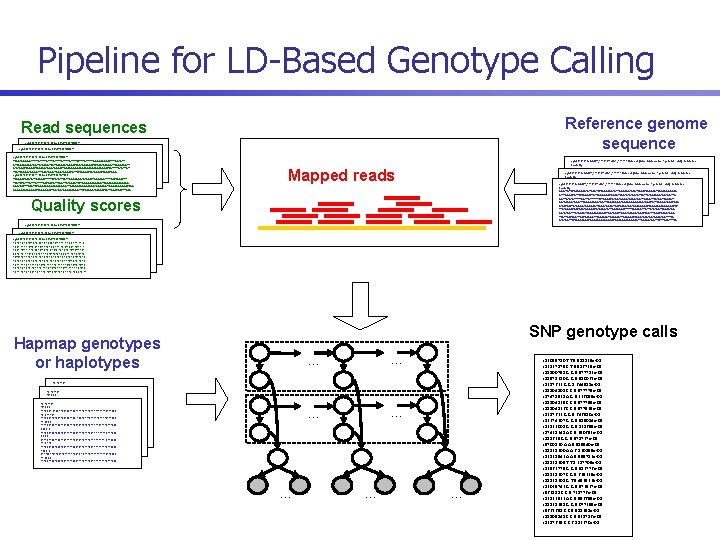
Pipeline for LD-Based Genotype Calling Reference genome sequence Read sequences >gnl|ti|1779718824 name: EI 1 W 3 PE 02 ILQXT TCAGTGAGGGTTTTTGTTTTGTTTTGTTTTGTTTTTGAGACAGAATTTTGCTCTT >gnl|ti|1779718824 name: EI 1 W 3 PE 02 ILQXT GTCGCCCAGGCTGGTGTGCAGTGGTGCAACCTCAGCTCACTGCAACCTCTGCCTCCAGGTTCAAGCAATT TCAGTGAGGGTTTTTGTTTTGTTTTGTTTTGTTTTTGAGACAGAATTTTGCTCTT CTCTGCCTCAGCCTCCCAAGTAGCTGGGATTACAGGCGCCACCACGCCCAGCTAATTTTGTATTGT GTCGCCCAGGCTGGTGTGCAGTGGTGCAACCTCAGCTCACTGCAACCTCTGCCTCCAGGTTCAAGCAATT >gnl|ti|1779718824 name: EI 1 W 3 PE 02 ILQXT TAGTAAAGATGGGGTTTCACTACGTTGGCTGAGCTGTTCTCGAACTCCTGACCTCAAATGAC CTCTGCCTCAGCCTCCCAAGTAGCTGGGATTACAGGCGCCACCACGCCCAGCTAATTTTGTATTGT TCAGTGAGGGTTTTTGTTTTGTTTTGTTTTGTTTTTGAGACAGAATTTTGCTCTT >gnl|ti|1779718825 name: EI 1 W 3 PE 02 GTXK 0 TAGTAAAGATGGGGTTTCACTACGTTGGCTGAGCTGTTCTCGAACTCCTGACCTCAAATGAC GTCGCCCAGGCTGGTGTGCAGTGGTGCAACCTCAGCTCACTGCAACCTCTGCCTCCAGGTTCAAGCAATT TCAGAATACCTGTTGCCCATTTTTATATGTTCCTTGGAGAAATGTCAATTCAGAGCTTTTGCTCAGCTTT >gnl|ti|1779718825 name: EI 1 W 3 PE 02 GTXK 0 CTCTGCCTCAGCCTCCCAAGTAGCTGGGATTACAGGCGCCACCACGCCCAGCTAATTTTGTATTGT TAATATGTTTATTTGTTTTGCTGCTGTTGAGTTGTACAATGTTGGGGAAAACAGTCGCACAACACCCGGC TCAGAATACCTGTTGCCCATTTTTATATGTTCCTTGGAGAAATGTCAATTCAGAGCTTTTGCTCAGCTTT TAGTAAAGATGGGGTTTCACTACGTTGGCTGAGCTGTTCTCGAACTCCTGACCTCAAATGAC AGGTACTTTGAGTCTGGGGGAGACAAAGGAGTTAGAAAGAGAATAAGCACTTAAAAGGCGGGTCCA TAATATGTTTATTTGTTTTGCTGCTGTTGAGTTGTACAATGTTGGGGAAAACAGTCGCACAACACCCGGC >gnl|ti|1779718825 name: EI 1 W 3 PE 02 GTXK 0 GGGGGCCCGAGCATCGGAGGGTTGCTCATGGCCCACAGTTGTCAGGCTCCACCTAATTAAATGGTTTACA AGGTACTTTGAGTCTGGGGGAGACAAAGGAGTTAGAAAGAGAATAAGCACTTAAAAGGCGGGTCCA TCAGAATACCTGTTGCCCATTTTTATATGTTCCTTGGAGAAATGTCAATTCAGAGCTTTTGCTCAGCTTT GGGGGCCCGAGCATCGGAGGGTTGCTCATGGCCCACAGTTGTCAGGCTCCACCTAATTAAATGGTTTACA TAATATGTTTATTTGTTTTGCTGCTGTTGAGTTGTACAATGTTGGGGAAAACAGTCGCACAACACCCGGC AGGTACTTTGAGTCTGGGGGAGACAAAGGAGTTAGAAAGAGAATAAGCACTTAAAAGGCGGGTCCA GGGGGCCCGAGCATCGGAGGGTTGCTCATGGCCCACAGTTGTCAGGCTCCACCTAATTAAATGGTTTACA >gi|88943037|ref|NT_113796. 1|Hs 1_111515 Homo sapiens chromosome 1 genomiccontig, reference assembly GAATTCTGTGAAAGCCTGTAGCTATAAAAAAATGTTGAGCCATAAATACCATCAGAAATAACAAAGGGAG CTTTGAAGTATTCTGAGACTTGTAGGAAGGTGAAGTAAATATCTAATATAATTGTAACAAGTAGTGCTTG >gi|88943037|ref|NT_113796. 1|Hs 1_111515 Homo sapiens chromosome 1 genomiccontig, reference GATTGTATGTTTTTGATTATTTTTTGTTAGGCTGTGATGGGCTCAAGTAATTGAAATTCCTGATGCAAGT assembly AATACAGATGGATTCAGGAGAGGTACTTCCAGGGGGTCAAGGGGAGAAATACCTGTTGGGGGTCAATGCC GAATTCTGTGAAAGCCTGTAGCTATAAAAAAATGTTGAGCCATAAATACCATCAGAAATAACAAAGGGAG CTCCTAATTCTGGAGTAGGGGCTAGAATGGTAGAATGCTCAAAAGAATCCAGCGAAGAGGAATAT >gi|88943037|ref|NT_113796. 1|Hs 1_111515 Homo sapiens chromosome 1 genomiccontig, reference CTTTGAAGTATTCTGAGACTTGTAGGAAGGTGAAGTAAATATCTAATATAATTGTAACAAGTAGTGCTTG TTCTGAGATAATAGGACTGTCCCATATTGGAGGCCTTTTTGAACAGTTGTTGTATGGTGACCCTGA assembly GATTGTATGTTTTTGATTATTTTTTGTTAGGCTGTGATGGGCTCAAGTAATTGAAATTCCTGATGCAAGT AATGTACTTTCTCAGATACAGAACACCCTTGGTCAATTGAATACAGATCACTTTAAGCTAAG GAATTCTGTGAAAGCCTGTAGCTATAAAAAAATGTTGAGCCATAAATACCATCAGAAATAACAAAGGGAG AATACAGATGGATTCAGGAGAGGTACTTCCAGGGGGTCAAGGGGAGAAATACCTGTTGGGGGTCAATGCC TCCTTACTAAATTGATGAGACTTAAACCCATGAAAACTTAACAGCTAAACTCCCTAGTCAACTGGTTTGA CTTTGAAGTATTCTGAGACTTGTAGGAAGGTGAAGTAAATATCTAATATAATTGTAACAAGTAGTGCTTG CTCCTAATTCTGGAGTAGGGGCTAGAATGGTAGAATGCTCAAAAGAATCCAGCGAAGAGGAATAT ATCTACTTCTCCAGCAGCTGGGGGAAAAAAGGTGAGAGAAGCAGGATTGAAGCTGCTTCTTTGAATTTAC GATTGTATGTTTTTGATTATTTTTTGTTAGGCTGTGATGGGCTCAAGTAATTGAAATTCCTGATGCAAGT TTCTGAGATAATAGGACTGTCCCATATTGGAGGCCTTTTTGAACAGTTGTTGTATGGTGACCCTGA Mapped reads Quality scores AATACAGATGGATTCAGGAGAGGTACTTCCAGGGGGTCAAGGGGAGAAATACCTGTTGGGGGTCAATGCC AATGTACTTTCTCAGATACAGAACACCCTTGGTCAATTGAATACAGATCACTTTAAGCTAAG CTCCTAATTCTGGAGTAGGGGCTAGAATGGTAGAATGCTCAAAAGAATCCAGCGAAGAGGAATAT TCCTTACTAAATTGATGAGACTTAAACCCATGAAAACTTAACAGCTAAACTCCCTAGTCAACTGGTTTGA TTCTGAGATAATAGGACTGTCCCATATTGGAGGCCTTTTTGAACAGTTGTTGTATGGTGACCCTGA ATCTACTTCTCCAGCAGCTGGGGGAAAAAAGGTGAGAGAAGCAGGATTGAAGCTGCTTCTTTGAATTTAC AATGTACTTTCTCAGATACAGAACACCCTTGGTCAATTGAATACAGATCACTTTAAGCTAAG TCCTTACTAAATTGATGAGACTTAAACCCATGAAAACTTAACAGCTAAACTCCCTAGTCAACTGGTTTGA ATCTACTTCTCCAGCAGCTGGGGGAAAAAAGGTGAGAGAAGCAGGATTGAAGCTGCTTCTTTGAATTTAC >gnl|ti|1779718824 name: EI 1 W 3 PE 02 ILQXT 28 28 26 28 28 40 34 14 44 36 23 13 2 27 42 35 21 7 27 42 35 21 6 28 name: EI 1 W 3 PE 02 ILQXT 43 36 22 10 27 42 35 20 6 28 43 36 22 9 >gnl|ti|1779718824 2813 28 28 2742 2835 2621 26 35 28 2843 2836 2622 28 92828404434361424441436423 2 27 7 26 >gnl|ti|1779718824 3 name: EI 1 W 3 PE 02 ILQXT 28 28 6282840433336142228936 27 27 4240 3534 21186 28 432836282227103327244226352820 28 2843 28 26 28 2 27 728 26 26 26 37928 29 28 28 27423 28 28 3742 2835 2721 27 35 36 28 37 36 22 2840 4434 3614 2444 1436 2813 28 27 28 26 26 27 4240 3534 21 6 28 43 36 22 10 28 28 28 27 28 28 24 28 28 27 28 37 29 36 18 3 28 28 28 27 28 3327 2442 2635 2820 28 628 2828 4043 3336 1422 28927 3627 2728 27 28 4326 3626 22 9 28 44 36 24 14 4 28 28 28 27 28 26 26 35 26 28 33 23 28 36 27 33 23 35 25 28 28 36 27 37 29 28 28 27 28 28 28 37 28 27 27 28 36 28 3736 27 40 3428 1828328 2828 2824 2728 3337 2429 2628 2819 2828 2826 4037 3329 1426 2839 3633 2713 28 27 28 28 28 24 28 28 27 28 28 37 29 36 27 27 28 37 27 2828 26 2628 3733 29 28 28 27 28 2815 2828 3736 2827 2726 2728 2824 3635 2827 3728 28 21 24 28 27 41 34 23 28 33 23 28 36 27 33 23 28 35 25 28 28 36 27 36 40 27 34 15 28 2828 2728 2824 2828 2837 2429 2828 2819 2728 2826 28 37 37 29 29 26 36 39 27 33 27 13 28 37 27 2828 28 3328 2321 2824 3328 2327 2841 3634 2715 3328 2336 2827 3526 2528 2824 28 35 36 27 27 28 36 40 27 34 15 28 28 28 24 28 37 29 28 19 28 26 37 29 26 39 33 13 37 2828 28 21 24 28 27 41 34 15 28 36 27 26 28 24 35 27 28 40 34 15 SNP genotype calls Hapmap genotypes or haplotypes 90 209342 16 F 0 0 2110001? 0100210010011002122201210211? 1221220212000 90 209342 16 F 0180 F 15 16 21100012010021001001100? 100201? 10111110111? 0212000 2110001? 0100210010011002122201210211? 1221220212000 15 16 M 00 18 F 15 90 2093422112001001201001120010110101011110212000 16 F 021100012010021001001100? 100201? 10111110111? 0212000 0 15 M 70 M 000 2110001? 0100210010011002122201210211? 1221220212000 211000100100020012211000111100111? 121210222000 2112001001201001120010110101011110212000 18 F 15 168 F 0 0 7 M 00 21100012010021001001100? 100201? 10111110111? 0212000 011202100120022012211200101101210211122111? 0120000 15 M 211000100100020012211000111100111? 121210222000 00 8 F 0 12 0 F 9 10 2112001001201001120010110101011110212000 21100010010002001221100010110111001112121210220000 7 M 0011202100120022012211200101101210211122111? 0120000 0 M 00 12 F 99 10 211000100100020012211000111100111? 121210222000 011? 001? 012002201221120010? 10121021112211110120000 21100010010002001221100010110111001112121210220000 8 F 0 0 11 M 7 8 9 M 0 0 011202100120022012211200101101210211122111? 0120000 21100210010002001221100012110111001112121210222000 011? 001? 012002201221120010? 10121021112211110120000 12 F 9 10 11 M 7 8 21100010010002001221100010110111001112121210220000 9 M 021100210010002001221100012110111001112121210222000 0 … … 011? 001? 012002201221120010? 10121021112211110120000 11 M 7 8 21100210010002001221100012110111001112121210222000 … … … rs 12095710 T T 9. 988139 e-01 rs 12127179 C T 9. 986735 e-01 rs 11800791 G G 9. 977713 e-01 rs 11578310 G G 9. 980062 e-01 rs 1287622 G G 8. 644588 e-01 rs 11804808 C C 9. 977779 e-01 rs 17471528 A G 5. 236099 e-01 rs 11804835 C C 9. 977759 e-01 rs 11804836 C C 9. 977925 e-01 rs 1287623 G G 9. 646510 e-01 rs 13374307 G G 9. 989084 e-01 rs 12122008 G G 5. 121655 e-01 rs 17431341 A C 5. 290652 e-01 rs 881635 G G 9. 978737 e-01 rs 9700130 A A 9. 989940 e-01 rs 11121600 A A 6. 160199 e-01 rs 12121542 A A 5. 555713 e-01 rs 11121605 T T 8. 387705 e-01 rs 12563779 G G 9. 982776 e-01 rs 11121607 C G 5. 639239 e-01 rs 11121608 G T 5. 452936 e-01 rs 12029742 G G 9. 973527 e-01 rs 562118 C C 9. 738776 e-01 rs 12133533 A C 9. 956655 e-01 rs 11121648 G G 9. 077355 e-01 rs 9662691 C C 9. 988648 e-01 rs 11805141 C C 9. 928786 e-01 rs 1287635 C C 6. 113270 e-01

Datasets n Watson n Sequencing data: 74. 4 million 454 reads (average length 265 bp) Reference panel: CEU genotypes from Hapmap r 23 a phased using the ENT algorithm [Gusev et al. 08] Ground truth: duplicate Affymetrix 500 k SNP genotypes n n Excluded discordant genotypes and SNPs for which Hapmap and Affymetrix annotations have more than 5% difference in same-strand CEU allele frequency NA 18507 (Illumina & SOLi. D) n n n Sequencing data: 525 million Illumina reads (36 bp, paired) and 764 million SOLi. D reads (24 - 44 bp, unpaired) Reference panel: YRI haplotypes from Hapmap r 22 excluding NA 18507 haplotypes Ground truth: Hapmap r 22 genotypes for NA 18507

Mapping Procedure n 454 reads mapped on human genome build 36. 3 using the NUCMER tool of the MUMmer package [Kurtz et al 04] with default parameters n n n Additional filtering: at least 90% of the read length matched to the genome, no more than 10 errors (mismatches or indels) Reads meeting above conditions at multiple genome positions (likely coming from genomic repeats) were discarded Illumina and SOLi. D reads mapped using MAQ [Li et al 08] with default parameters n n For reads mapped at multiple positions MAQ returns best position (breaking ties arbitrarily) together with mapping confidence We filtered bad alignments and discarded paired end reads that are not mapped in pairs using the “submap -p” command
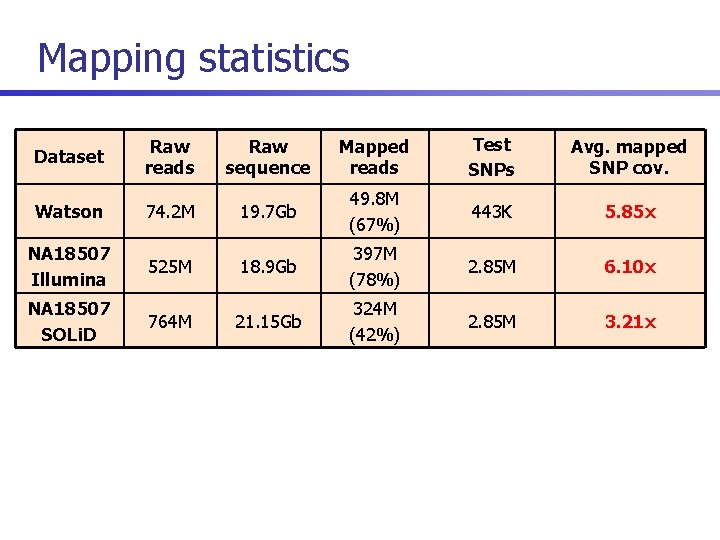
Mapping statistics Dataset Raw reads Raw sequence Mapped reads Test SNPs Avg. mapped SNP cov. Watson 74. 2 M 19. 7 Gb 49. 8 M (67%) 443 K 5. 85 x 2. 85 M 6. 10 x 2. 85 M 3. 21 x NA 18507 Illumina 525 M 18. 9 Gb 397 M (78%) NA 18507 SOLi. D 764 M 21. 15 Gb 324 M (42%)
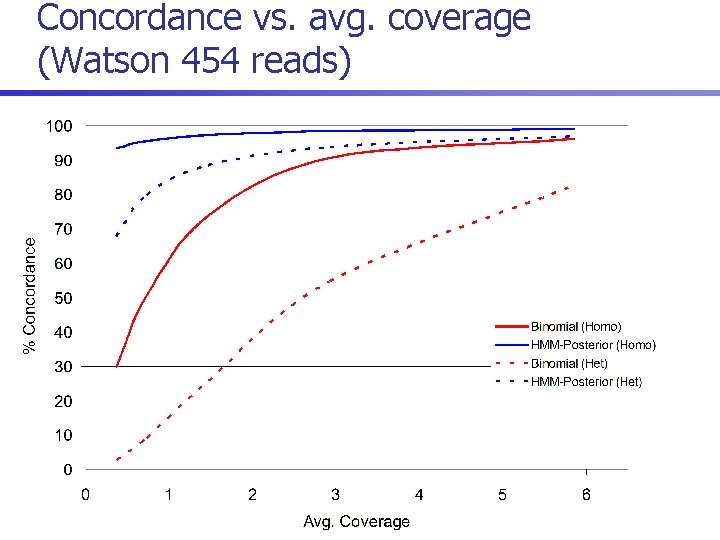
Concordance vs. avg. coverage (Watson 454 reads)
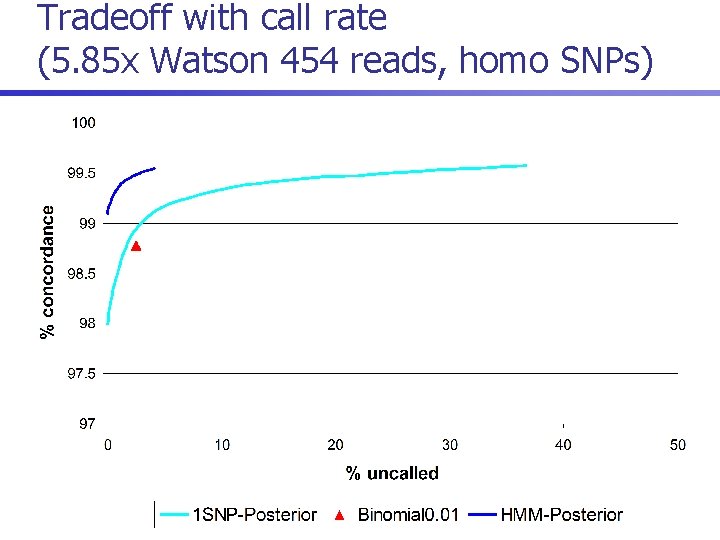
Tradeoff with call rate (5. 85 x Watson 454 reads, homo SNPs)

Tradeoff with call rate (5. 85 x Watson 454 reads, het SNPs)

Concordance vs. avg. coverage for NA 18507 (Illumina & SOLi. D reads)
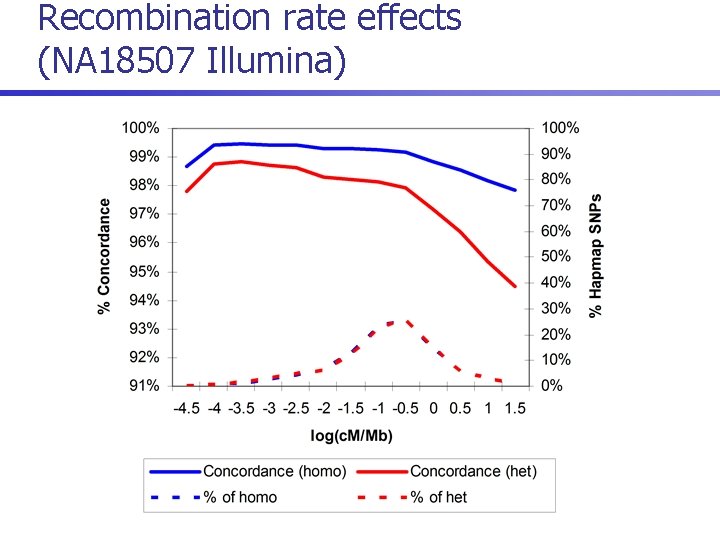
Recombination rate effects (NA 18507 Illumina)
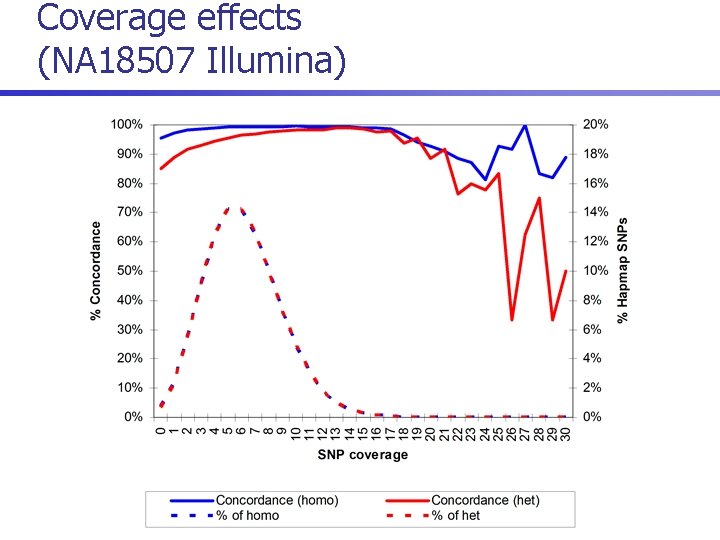
Coverage effects (NA 18507 Illumina)

Conclusions & ongoing work n Exploiting LD information yields significant improvements in genotyping calling accuracy and/or cost reductions n n Improvement depends on the coverage depth (higher at lower coverage), e. g. , accuracy achieved by previously proposed binomial test at 5 -6 x average coverage is achieved by HMM-based posterior decoding algorithm using less than 1/4 of the reads Ongoing work n n Extension to population sequencing data (removing need for reference panels) Mapping repetitive reads & haplotype inferrence

Acknowledgments n Work supported in part by NSF awards IIS-0546457 and DBI-0543365 to IM and IIS-0803440 to YW. SD and YH performed this research as part of the Summer REU program “Bio-Grid Initiatives for Interdisciplinary Research and Education" funded by NSF award CCF-0755373.
- Slides: 45