Laboratoire dInformatique Fondamentale de Lille Parallel Cooperative Optimization

Laboratoire d’Informatique Fondamentale de Lille Parallel Cooperative Optimization Research Group Evolutionary Aglorithms for the Protein Structure Prediction Problem and Molecular Docking A. -A. Tantar, N. Melab and E-G. Talbi {tantar, melab, talbi}@lifl. fr INRIA DOLPHIN Project Supported by the French Research Agency

Outline n n Protein Structure Prediction, Molecular Docking: modeling and complexity analysis Parallel Hybrid Metaheuristics for the PSP Problem n Grid experimentation on GRID 5000 n Conclusion and Future Work
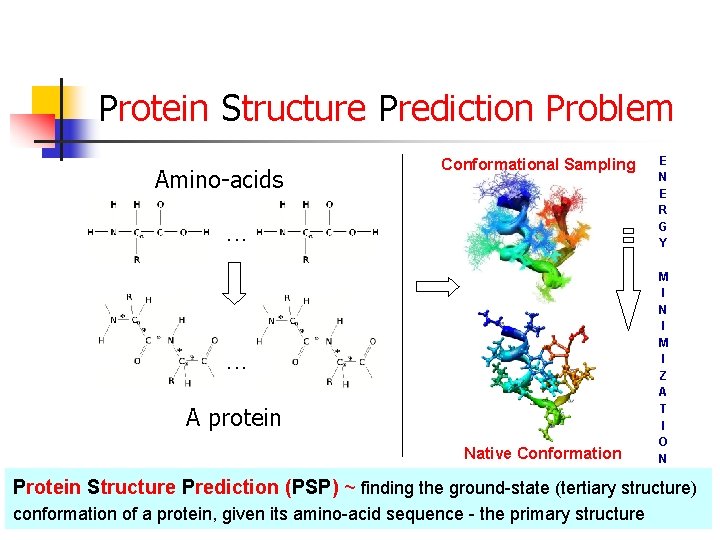
Protein Structure Prediction Problem Amino-acids Conformational Sampling . . . A protein Native Conformation E N E R G Y M I N I M I Z A T I O N Protein Structure Prediction (PSP) ~ finding the ground-state (tertiary structure) conformation of a protein, given its amino-acid sequence - the primary structure
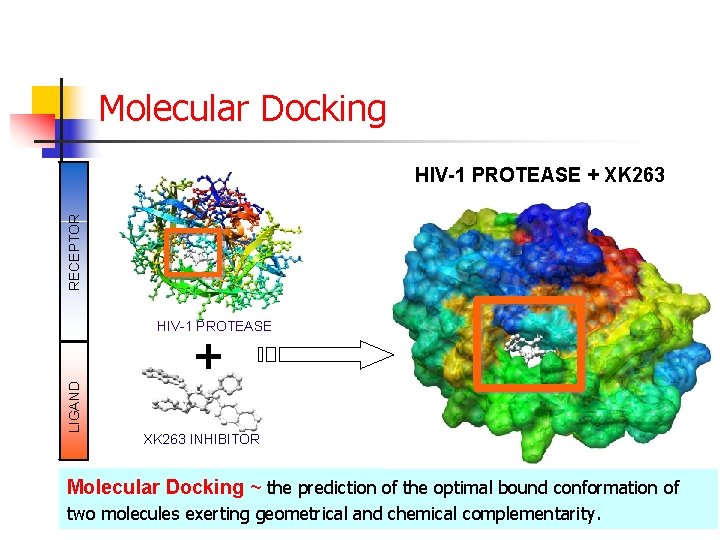
Molecular Docking RECEPTOR HIV-1 PROTEASE + XK 263 HIV-1 PROTEASE LIGAND + XK 263 INHIBITOR Molecular Docking ~ the prediction of the optimal bound conformation of two molecules exerting geometrical and chemical complementarity.

Scoring Energy Function (A) Empirical Force Fields – Classical Mechanics n AMBER – Assisted Model Building with Energy Refinement n CHARMM – Chemistry at HARvard Molecular Mechanics n OPLS – Optimized Potentials for Liquid Simulations n GROMOS – GROningen MOlecular Simulation n SYBYL n D-Score - DOCK n F-Score - Flex. X n G-Score - Gold n Auto. Dock/Auto. Grid – suite of toos for automated docking n . . . WEAK London Forces Van der Waals dipole-dipole stacking hydrogen bond STRONG ionic interactions From recent results Empirical Methods OUTPERFORM Ab Initio Methods (!!) § Neumaier, A. (1997). Molecular modeling of proteins and mathematical prediction of protein structure. SIAM Review, 39(3): 407 – 460.

Scoring Energy Function (B) § Auto. Dock § DOCK - SYBYL/D-Score

Scoring Energy Function (C) § GOLD - SYBYL/G-Score § Flex. X - SYBYL/F-Score § Renxiao Wang. Yipin Lu, and Shaomeng Wang, Comparative Evaluation of 11 Scoring Functions for Molecular Modeling, J. Med. Chem. , 10. 1021/jm 0203783, 2003, 46, 2287 -2303

Scoring Energy Function (D) the factors model oscillating entities – forces simulated by interconnecting springs between atoms; offer an easy concept and fast energy evaluations. constants derived from higher-level calculations (e. g. Ab initio) – difficult to obtain and to fit. empirical force fields have the DRAWBACK of offering results not directly comparable with results obtained through another differently parameterized force field. RMSD (Root Mean Squared Deviation) – typical distance measure between conformations.
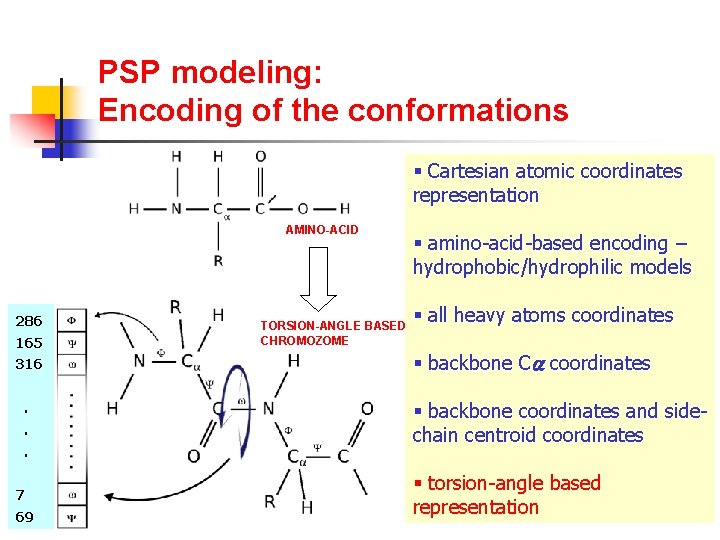
PSP modeling: Encoding of the conformations § Cartesian atomic coordinates representation AMINO-ACID 286 165 316. . . 7 69 TORSION-ANGLE BASED CHROMOZOME § amino-acid-based encoding – hydrophobic/hydrophilic models § all heavy atoms coordinates § backbone C coordinates § backbone coordinates and sidechain centroid coordinates § torsion-angle based representation
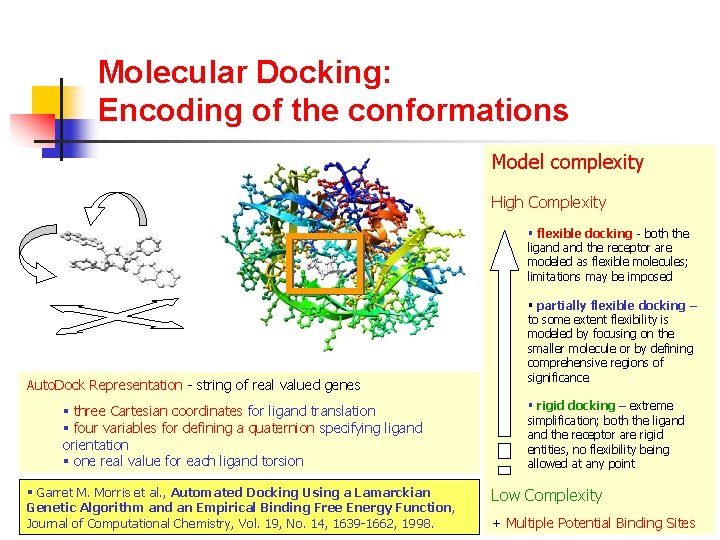
Molecular Docking: Encoding of the conformations Model complexity High Complexity § flexible docking - both the ligand the receptor are modeled as flexible molecules; limitations may be imposed Auto. Dock Representation - string of real valued genes § three Cartesian coordinates for ligand translation § four variables for defining a quaternion specifying ligand orientation § one real value for each ligand torsion § Garret M. Morris et al. , Automated Docking Using a Lamarckian Genetic Algorithm and an Empirical Binding Free Energy Function, Journal of Computational Chemistry, Vol. 19, No. 14, 1639 -1662, 1998. § partially flexible docking – to some extent flexibility is modeled by focusing on the smaller molecule or by defining comprehensive regions of significance § rigid docking – extreme simplification; both the ligand the receptor are rigid entities, no flexibility being allowed at any point Low Complexity + Multiple Potential Binding Sites
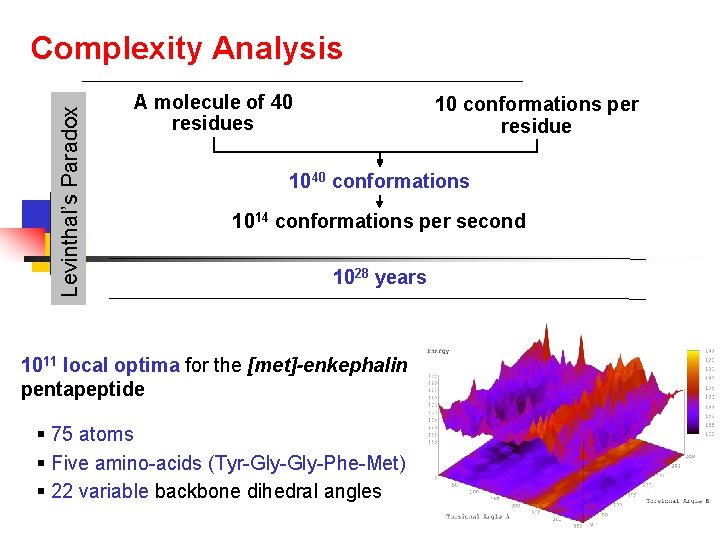
Levinthal’s Paradox Complexity Analysis A molecule of 40 residues 10 conformations per residue 1040 conformations 1014 conformations per second 1028 years 1011 local optima for the [met]-enkephalin pentapeptide § 75 atoms § Five amino-acids (Tyr-Gly-Phe-Met) § 22 variable backbone dihedral angles

Motivation and objectives n PSP and Molecular Docking can be modeled as a optimization problems … n … which are multi-modal n … and require a huge amount of resources for large proteins n Need of hybrid metaheuristics n Need of large scale parallelism (Grid computing) n $$$ : $150000 -$250000 per X-ray structure n Study of two hybrid metaheuristics for PSP: n Genetic Algorithms and Simulated Annealing

Classification of the Existing Methods n Energy minimization vs. Geometry complementarity approaches n Ab initio, de nuovo electronic structure calculations n n Semi-empirical methods n n n make use of approximations as substitution for ab initio techniques employ simplified models for electron-electron interactions Empirical methods n n rely on quantum mechanics for determining different molecular characteristics comprise no approximations and no a priori experimental data is required high computational complexity => reduced size systems rely upon molecular dynamics (classical mechanics based methods) often the only applicable methods for large molecular systems, i. e. proteins and polymers do not dissociate atoms into electrons and nuclei - indivisible entities Continuous vs. Discretization - Combinatorial Optimization

Classification – Algorithmic Overview (A) n Global Optimization – Energy Landscape Projections, Convex Global Underestimation, etc. n n n construct a metamodel of the energy surface - avoid most of the inconvenients determined by the funnel with bumps structure of the landscape tend to require less computational time than classical techniques – a few orders of magnitude apparently have a lower probability of remaining blocked in kinetic traps employ suppositions regarding the landscape which may not extend to different classes require the construction of a metamodel, underestimation, etc. which offers no optimality/corectness guarantee Molecular Dynamics n n n offer accurate results as far as the force field model fits the real energy landscape the method can be applied straightforward – apart the formalism, no parameters to be tuned require extensive computational resources – may only be applied to reduced models the design and the development of the formalism may be far from trivial given the high computational complexity, stand-alone approaches cannot be practical bond vibration 10 -15 femto MD step isomerization, water dynamics 10 -12 pico 10 -9 nano, MD long run fastest folders 10 -6 micro typical folders 10 -3 mili slow folders 100 seconds

Classification – Algorithmic Overview (B) n Monte Carlo / Simulated Annealing / Metropolis-based algorihtms n n n n do not require a large number of parameters – adaptive versions overcome to an extent the inconvenients determined by the initial specified values offer a performance guarantee for ataining the optimal value compared to gradient based methods, are less prone to getting traped in local minima require a non feasible number of sampling points for having the performance guarantee appropiate for local search optimization or for the refinement of specific areas might employ specific distributions for performing the conformational sampling depending on the nature of the method, it may pose important parallelization problems Evolutionary algoritmhs – Genetic Algorithms n n n exhibit strong exploration characteristics while lacking intensification capabilities – hybridization approaches tend to be more appropiate offer the base for complex parallelization techniques require implementing different components, having different parameters the design of the operators may require to consider structural aspects (i. e. xover operator should not mix angles from different amino-acids when performing the recombination, etc. ) do not offer any performance guarantee – statistical convergence
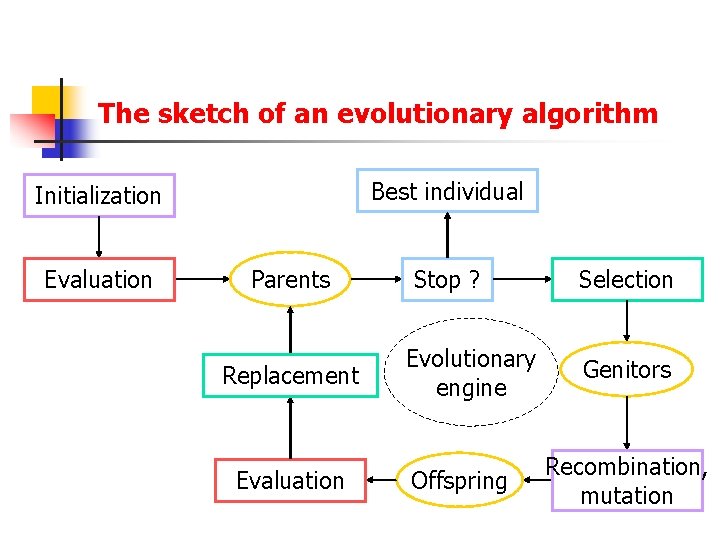
The sketch of an evolutionary algorithm Best individual Initialization Evaluation Parents Replacement Evaluation Stop ? Evolutionary engine Offspring Selection Genitors Recombination, mutation

Generation of the Initial population n Seeding - information inserted from structural databases n n n Ab initio approach – no initial structural information is given n ensure that the population is seeded with the substructures that lead to the optimal conformation does not represent the desired approach as strucutural information may not be available in all the cases uniform random initialization of the dihedral angles – the most common option depending on structural data different distributions may be employed for offering a bias The size of the population n n may have an impact on the convergence rate: n reduced number of chromosomes -> premature convergence n larger number of chromosomes -> extra computational time should be related to the cardinality of the alphabet used for encoding, chromosome length, algorithm parameters, etc. ; empirical, restricted to special cases - no rigorous specifications § DUSAN P. DJURDJEVIC, MARK J. BIGGS, Ab Initio Protein Fold Prediction Using Evolutionary Aglorithms: Influence of Design and Control Parameters on Performance, 10. 1002/jcc. 20440, Wiley Inter. Science, 2005.

Selection and Replacement (A) Selection pressure the probability of selecting the best chromosome as compared to the average selection probability of all the chromosomes Selection intensity population expected average fitness value after performing a selection step having as base a normalized Gaussian distribution Selection variance population expected variance of the fitness distribution after performing a selection step having as base a normalized Gaussian distribution Loss of diversity the percentage of non-selected individuals during the selection phase Bias the absolute difference between the expected reproduction probability of a specific chromozome and the chromosome’s normalized fitness Spread range of possible values for the offsprings of a given chromosome

Selection and Replacement (B) Selection strategy n fitness-proportionate selection n n rank-based selection n n selection relative to rank and not to the chromosome’s real fitness value stochastic tournament selection n n no bias, does not guarantee a minimum spread tournament among randomly chosen individuals selection pressure can be adjusted by modifying the size of the tournament offer a constant selection pressure uniform selection n n selection bias towards sparse fitness levels and not towards best fit chromosomes might not behave well under extensive genetic diversity conditions (i. e, initial phases)

Selection and Replacement (C) Replacement strategy n Global replacement vs. Local replacement n n n n generational replacement – replace all the parents with the offsprings uniform replacement - less offsprings than parents, replace at random elitist replacement - less offsprings than parents, replace worst parents fitness-based replacement - more offsprings than parents, insert only the best offsprings random, first-in-first out – the simplest ones, do not exert strong performace characteristics worst-fit replacement – the least fit chromosome(s) are replaced exponential replacement – chromosomes are ranked from the least fit to the most fit, the replacement process employing an associated probability distribution. . .

Recombination Operators n n discrete recombination – exchange of chromosome values; for each locus in the offspring, the parent chromosomes have equal probabilities of contributing intermediate recombination – offsprings obtained by interpolation of the parents Offspring[ loci ] = ui * A[ loci ] + ( 1 -ui ) * B [ loci ], ui [ - , 1+ ], i 1, n n line recombination – particular case of intermediat recombination, a single uniformly random generated variable being considered Offspring[ loci ] = u * A[ loci ] + ( 1 -u ) * B [ loci ], u [ - , 1+ ], i 1, n n one/multi point crossover – implies the generation of one/multiple cutting points marking the segments to be combined into offsprings n uniform crossover – each locus is consider as potential cutting point n . . .

Mutation Operators n n real valued mutation – the main parameters to adjust are the mutation step size and the mutation probability – usually chosen to be 1/n, n – no. of variables variable step mutation – small step mutations are accepted with high probability while large step mutations are accepted with low probability M[ loci ] = M[ loci ] + si * ri * pi , 1, n n si { -1, +1 } n ri = r * |Bi – Ai|, r ~ 0. 1, M[ loci ] [Ai, Bi] n pi =2 -u*k, u [ 0, 1 ] n n step-size adaptation mutation – n step-sizes / one direction / n-directions; do not offer consistent improvements for extreme multi-modal / high noise functions. . .
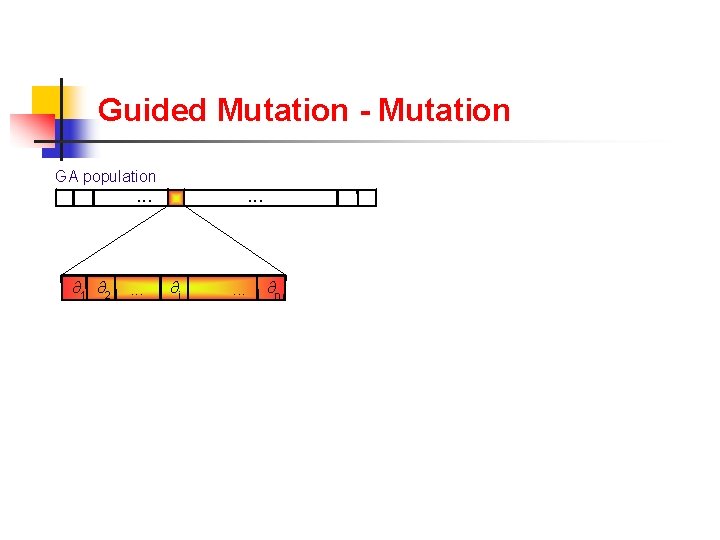
Guided Mutation - Mutation + LS GA population . . . ∂1 ∂2 . . . ∂i . . . ∂n ∂i ' . . . ∂n Angle Mutation ∂1 ∂2 . . . Local Search ∂'1 ∂'2 . . . Optimized Individual ∂'n

Termination Criteria n preset number of fitness function evaluations / number of generations n termination once the evolution ceases n optimal value has been found (!!) n . . .
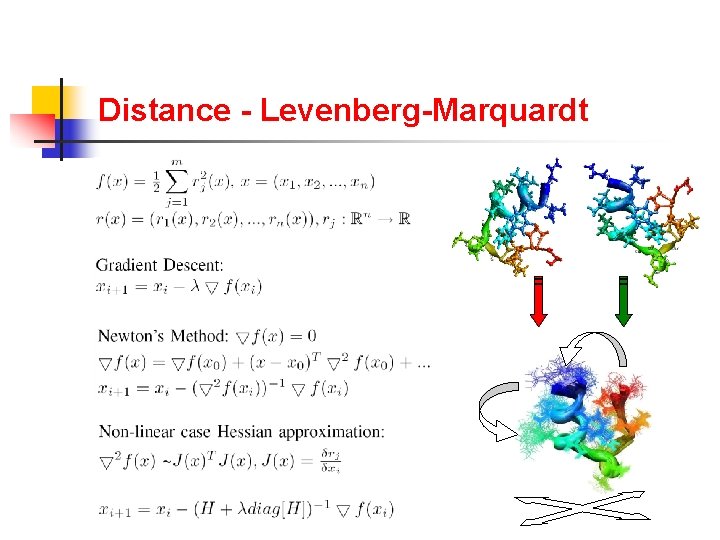
Distance - Levenberg-Marquardt
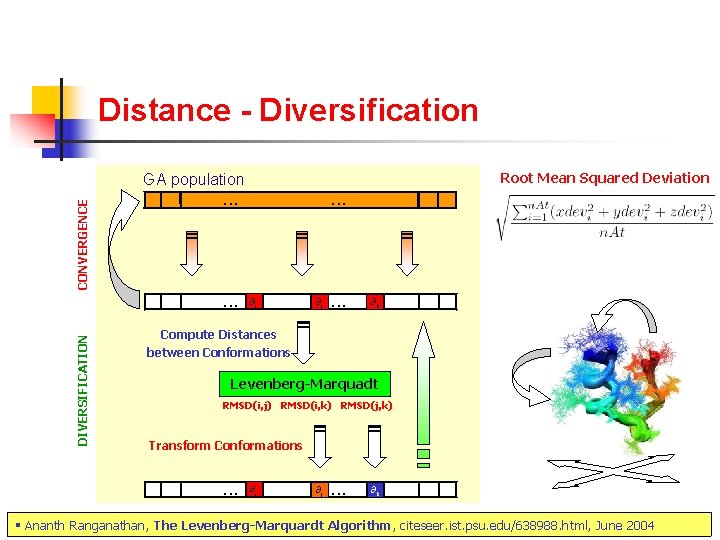
Distance - Diversification Root Mean Squared Deviation DIVERSIFICATION CONVERGENCE GA population . . ∂i ∂j . . . ∂k Compute Distances between Conformations Levenberg-Marquadt RMSD(i, j) RMSD(i, k) RMSD(j, k) Transform Conformations . . . ∂i ∂j . . . ∂k § Ananth Ranganathan, The Levenberg-Marquardt Algorithm, citeseer. ist. psu. edu/638988. html, June 2004

Literature examples (A) n n Dusan P. Djurdjevic, Mark J. Biggs, Ab Initio Protein Fold Prediction Using Evolutionary Algorithms: Influence of Design and Control Parameters on Performance, DOI 10. 1002/jcc. 20440, Wiley Interscience n Generational Evolutionary Algorithm n Steady-State Evolutionary Algorithm n tournament selection, single-point/multi-point and uniform crossover, uniform distribution mutation with angles within the range of 0 -360, termination in case of lack of improvement for a specified number of generations Christopher D. Rosin University of California, San Diego, A Comparison of Global and Local Search Methods in Drug Docking, Proceedings of the Seventh International Conference on Genetic Algorithms, ICGA, 1997 n Simulated Annealing – 100 tests with 1. 5~1. 8 billion function evaluations per test, linearly reduced temperature n Solis and Wet’s Algorithm (a class of local and global search algorithms) – part of the GA+LS hybrid, deviations chosen from a normal distribution - does not require gradient information n Genetic Algorithm+LS – local search applied to only 7% of the population in each generation; stochastic selection, two-points crossover, Cauchy-deviate mutation

Literature examples (B) n Garrett M. Morris et al, Automated Docking Using a Lamarckian Genetic Algorithm and an Empirical Binding Free Energy Function, Journal of Computational Chemistry, Vol. 19, No. 14, 1639 -1662, 1998. n Monte Carlo – applied on a pre-calculated grid of interaction energies n Simulated Annealing n Genetic Algorithm, Lamarckian Genetic Algorithm n n n uniform random value initialization (-180. 0, 180. 0), maximum number of generations/function evaluations, proportional elitist selection, constant-size population, two-point crossover with breaks between genes, Cauchy distribution-based mutation, local search at the end of each generation on a user-defined percentage of the population each method is given ~1. 5 million function evaluations / up to 41. 5 minutes on a 200 MHz Silicon Graphics MIPS with 128 Mb of RAM

Outline n n Protein Structure Prediction (PSP): modeling and complexity analysis Parallel Hybrid Metaheuristics for the PSP Problem n Grid experimentation on GRID 5000 n Conclusion and Future Work

PARAllel and DIStributed Evolving Objects http: //paradiseo. gforge. inria. fr/ Paradis. EO for computational Grids Evolving Objects framework (EO) MO MOEO European project (Geneura Team, http: //eodev. sourceforge. net INRIA, LIACS) PVM, PThreads MPI (LAM, CH) Transparent use § § Our contributions § Multi-Objective EO (MOEO) for the design of multi-objective evolutionary algorithms § Moving Objects (MO) for the design of local search algorithms § Paradis. EO for parallel hybrid metaheuristics Condor-MW Globus § § Message passing (MPI, PVM) § Clusters, Networks of Workstations, § Shared Memory Multi-processors (SMP) § Clusters of SMPs (CLUMPS) Multi-programming (PThreads) Parallel distributed computing Grid computing § Condor-MW and Globus (MPICH-G 2) § S. Cahon, N. Melab and E-G. Talbi. Paradis. EO: A Framework for the Reusable Design of Parallel and Distributed Metaheuristics. Journal of Heuristics, Elsevier Science, Vol. 10(3), pages 357 -380, May 2004.
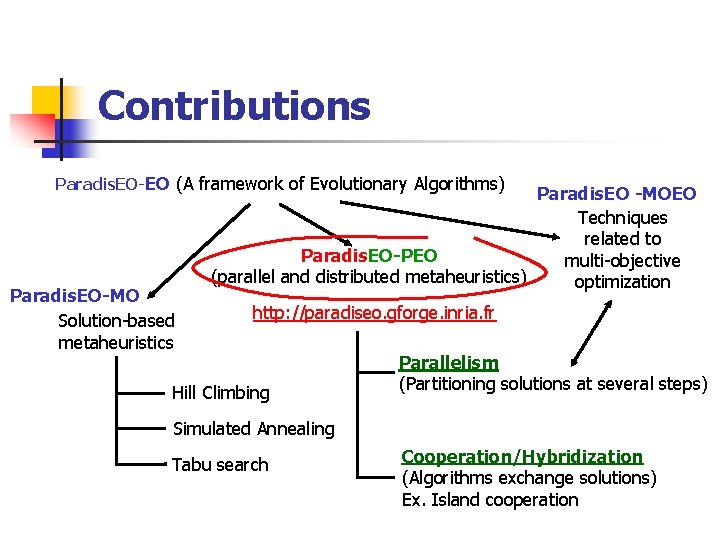
Contributions Paradis. EO-EO (A framework of Evolutionary Algorithms) Paradis. EO-MO Solution-based metaheuristics Paradis. EO -MOEO Techniques related to Paradis. EO-PEO multi-objective (parallel and distributed metaheuristics) optimization http: //paradiseo. gforge. inria. fr Hill Climbing Parallelism (Partitioning solutions at several steps) Simulated Annealing Tabu search Cooperation/Hybridization (Algorithms exchange solutions) Ex. Island cooperation
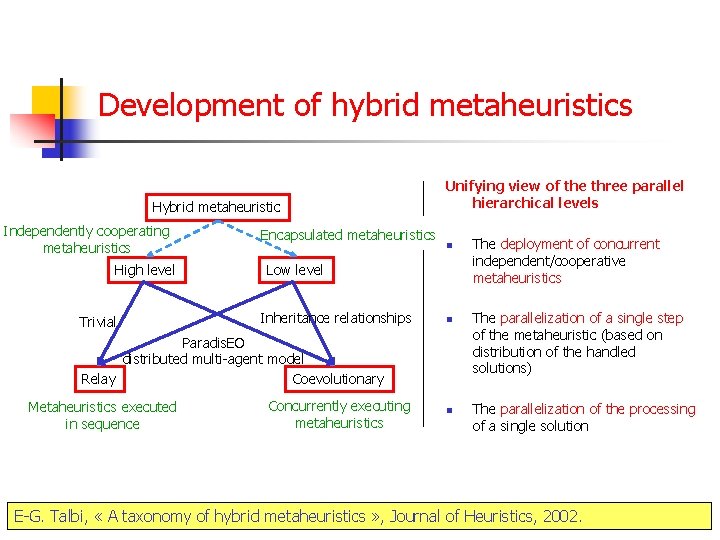
Development of hybrid metaheuristics Hybrid metaheuristic Independently cooperating metaheuristics High level Trivial Encapsulated metaheuristics Unifying view of the three parallel hierarchical levels n Low level Inheritance relationships n Paradis. EO distributed multi-agent model Relay Coevolutionary Metaheuristics executed in sequence Concurrently executing metaheuristics n The deployment of concurrent independent/cooperative metaheuristics The parallelization of a single step of the metaheuristic (based on distribution of the handled solutions) The parallelization of the processing of a single solution E-G. Talbi, « A taxonomy of hybrid metaheuristics » , Journal of Heuristics, 2002.
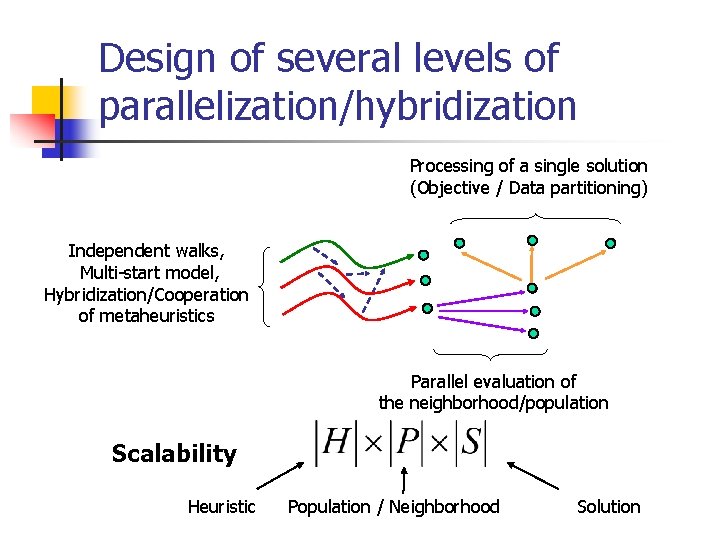
Design of several levels of parallelization/hybridization Processing of a single solution (Objective / Data partitioning) Independent walks, Multi-start model, Hybridization/Cooperation of metaheuristics Parallel evaluation of the neighborhood/population Scalability Heuristic Population / Neighborhood Solution
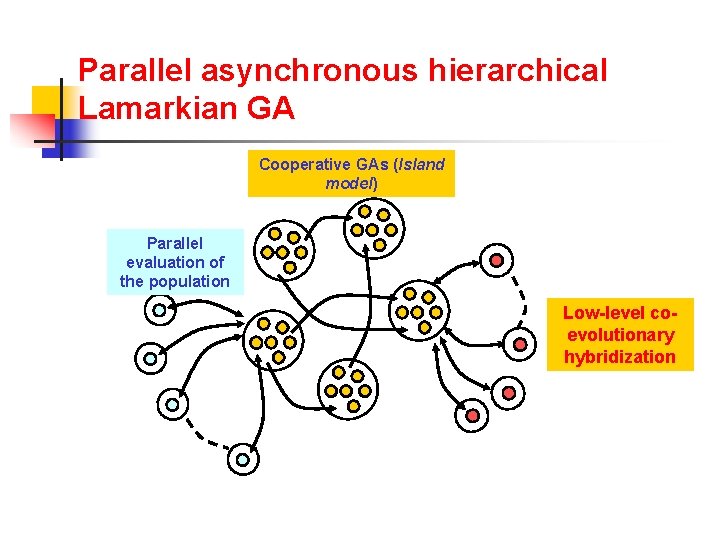
Parallel asynchronous hierarchical Lamarkian GA Cooperative GAs (Island model) Parallel evaluation of the population Low-level coevolutionary hybridization
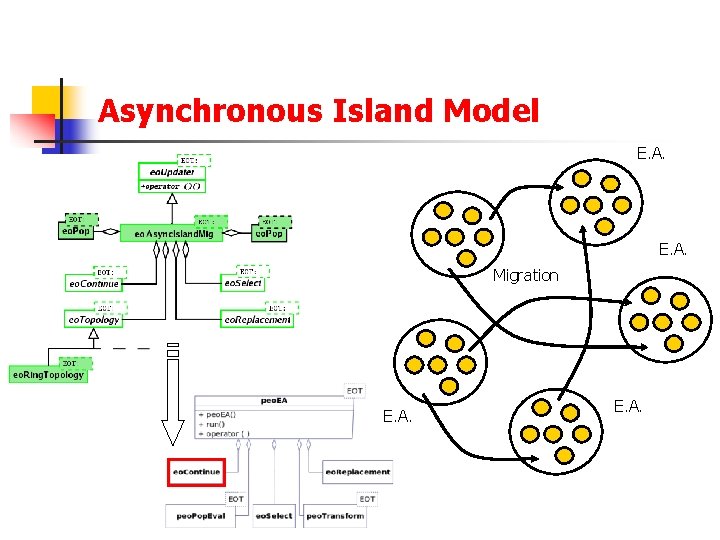
Asynchronous Island Model E. A. Migration E. A.
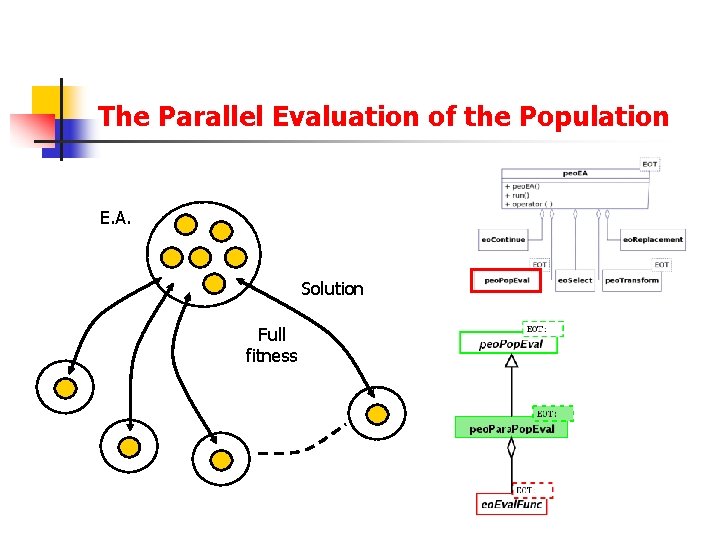
The Parallel Evaluation of the Population E. A. Solution Full fitness
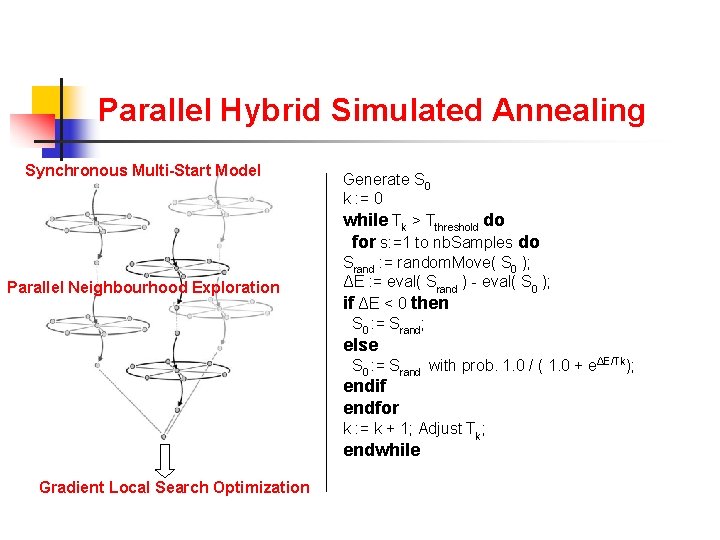
Parallel Hybrid Simulated Annealing Synchronous Multi-Start Model Parallel Neighbourhood Exploration Generate S 0 k : = 0 while Tk > Tthreshold do for s: =1 to nb. Samples do Srand : = random. Move( S 0 ); ΔE : = eval( Srand ) - eval( S 0 ); if ΔE < 0 then S 0 : = Srand; else S 0 : = Srand with prob. 1. 0 / ( 1. 0 + eΔE/Tk); endif endfor k : = k + 1; Adjust Tk; endwhile Gradient Local Search Optimization

Outline n n Protein Structure Prediction (PSP): modeling and complexity analysis Parallel Hybrid Metaheuristics for the PSP Problem n Grid experimentation on GRID 5000 n Conclusion and Future Work
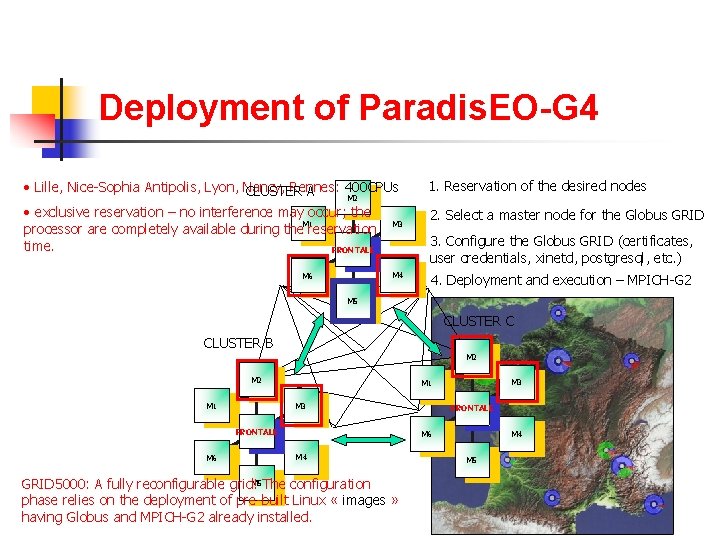
Deployment of Paradis. EO-G 4 • Lille, Nice-Sophia Antipolis, Lyon, Nancy, Rennes: 400 CPUs CLUSTER A 1. Reservation of the desired nodes • exclusive reservation – no interference may occur; the M 1 processor are completely available during the reservation time. FRONTALE 2. Select a master node for the Globus GRID M 2 M 6 M 3 3. Configure the Globus GRID (certificates, user credentials, xinetd, postgresql, etc. ) M 4 4. Deployment and execution – MPICH-G 2 M 5 CLUSTER C CLUSTER B M 2 M 3 M 1 FRONTALE M 6 M 3 M 1 FRONTALE M 4 M 6 M 4 M 5 GRID 5000: A fully reconfigurable grid! The configuration phase relies on the deployment of pre-built Linux « images » having Globus and MPICH-G 2 already installed. M 5
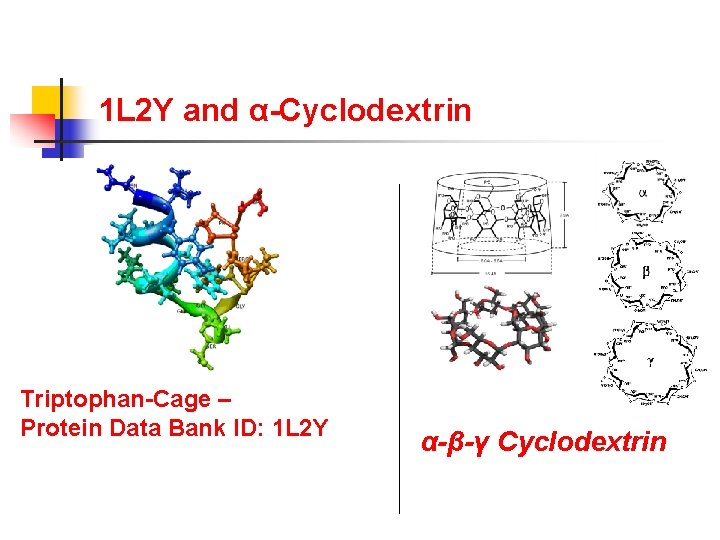
1 L 2 Y and α-Cyclodextrin Triptophan-Cage – Protein Data Bank ID: 1 L 2 Y α-β-γ Cyclodextrin
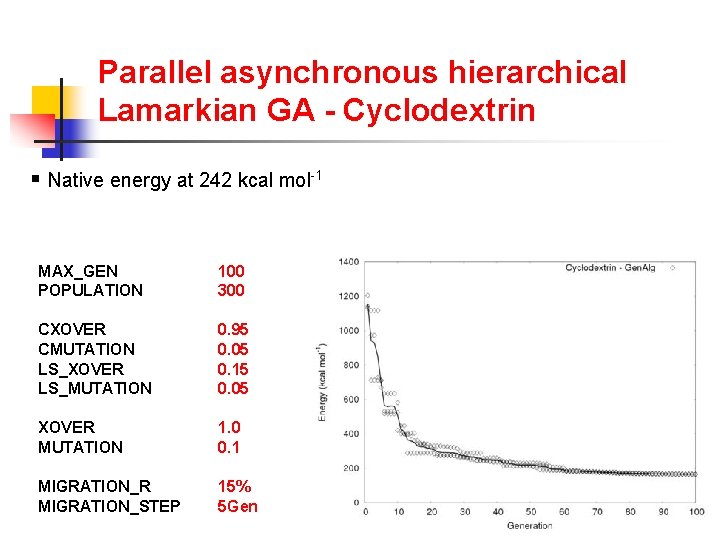
Parallel asynchronous hierarchical Lamarkian GA - Cyclodextrin § Native energy at 242 kcal mol-1 MAX_GEN POPULATION 100 300 CXOVER CMUTATION LS_XOVER LS_MUTATION 0. 95 0. 05 0. 15 0. 05 XOVER MUTATION 1. 0 0. 1 MIGRATION_R MIGRATION_STEP 15% 5 Gen
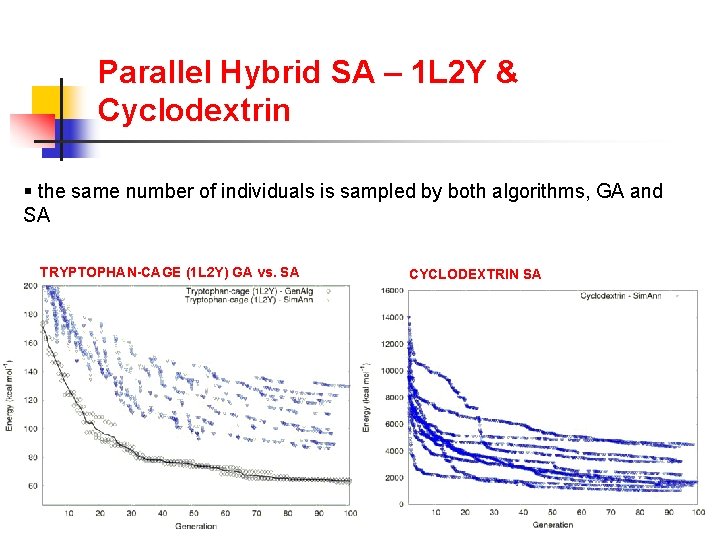
Parallel Hybrid SA – 1 L 2 Y & Cyclodextrin § the same number of individuals is sampled by both algorithms, GA and SA TRYPTOPHAN-CAGE (1 L 2 Y) GA vs. SA CYCLODEXTRIN SA
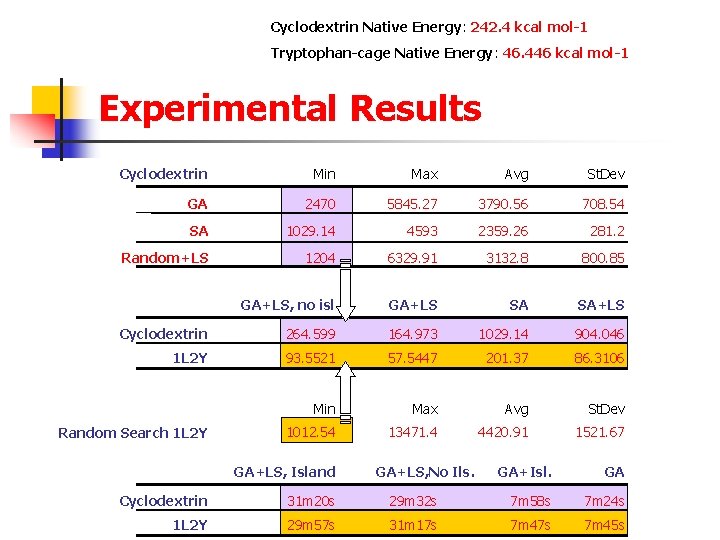
Cyclodextrin Native Energy: 242. 4 kcal mol-1 Tryptophan-cage Native Energy: 46. 446 kcal mol-1 Experimental Results Cyclodextrin Max Avg St. Dev GA 2470 5845. 27 3790. 56 708. 54 SA 1029. 14 4593 2359. 26 281. 2 Random+LS 1204 6329. 91 3132. 8 800. 85 GA+LS SA SA+LS GA+LS, no isl. Cyclodextrin 264. 599 164. 973 1029. 14 904. 046 1 L 2 Y 93. 5521 57. 5447 201. 37 86. 3106 Random Search 1 L 2 Y Min Max Avg St. Dev 1012. 54 13471. 4 4420. 91 1521. 67 GA+LS, Island GA+LS, No Ils. GA+Isl. GA Cyclodextrin 31 m 20 s 29 m 32 s 7 m 58 s 7 m 24 s 1 L 2 Y 29 m 57 s 31 m 17 s 7 m 47 s 7 m 45 s

Outline n n Protein Structure Prediction (PSP): modeling and complexity analysis Parallel Hybrid Metaheuristics for the PSP Problem n Grid experimentation on GRID 5000 n Conclusion and Future Work

Conclusion and Future Work (1) n The GA behaves better on the considered benchmarks, especially on Cyclodextrin n n Bellow native-conformation energy results were obtained The SA results are comparable to the results obtained by the GA with no hybridization The SA allows for little intrinsic parallelism … … but, it is not getting easily trapped in local minima (like directed LS, i. e. gradient LS)

Conclusion and Future Work (2) n n Hierarchical multi-stage parallel models may prove to be efficient on the considered benchmarks Hybridization schemes combining … n n … the GA (strong exploration capabilities) … and the SA (intensification + less prone to getting trapped in local minima than gradient methods)

Paradis. EO-PEO Implementation

Paradis. EO-PEO Evolutionary Algorithm peo. EA ( eo. Continue< EOT >& __cont, peo. Pop. Eval< EOT >& __pop_eval, eo. Select< EOT >& __select, peo. Transform< EOT >& __transform, eo. Replacement< EOT >& __replace ); pop_eval ( population ); do { eo. Pop< EOT > offsprings; select ( population, offsprings ); § evaluation function - the designed class has to be derived from the peo. Pop. Eval class § § peo. Seq. Pop. Eval peo. Para. Pop. Eval § selection strategy – applied at each iteration in order to obtain the offspring population § transformation operators - crossover operator(s), mutation operator(s); peo. EA requires a peo. Transform derived object § § peo. Seq. Transform peo. Para. SGATransform transform ( offsprings ); pop_eval ( offsprings ); § replacement strategy – for integrating the offsprings back into the initial population § continuation criterion - maximum number of generations, checkpoints, etc. replace ( population, offsprings ); } while ( cont ( population ) );

Paradis. EO-PEO EA Representation of the Individuals #include <EO. h>. . . class Conformation : public EO< eo. Scalar. Fitness< double, greater< double > > > {. . . int operator[]( const int& index ) const { }. . . }; return chromosome->get. Chromo( index+1 ); Individu* chromosome; . . . static Molecule* molecule; static Hamiltonian* hamiltonian; static Population* population; extern void pack( const docking. At. Grid: : Conformation& conformation ); extern void unpack( docking. At. Grid: : Conformation& conformation );

Unpack Recv. Data Pack Data to Send Paradis. EO-PEO EA Representation – Packing & Unpacking void pack( const docking. At. Grid: : Conformation& conformation ) { if ( conformation. invalid() ) pack( (unsigned int)0 ); else { pack( (unsigned int)1 ); pack( conformation. fitness() ); } } for ( unsigned int index = 0; index < conformation. size(); index++ ) pack( conformation[ index ] ); void unpack( docking. At. Grid: : Conformation& conformation ) { eo. Scalar. Fitness<double, std: : greater<double> > fitness; unsigned int valid. Fitness; } unpack( valid. Fitness ); if ( valid. Fitness ) { unpack( fitness ); conformation. fitness( fitness ); } else {. . . } M I D L E W A R E

Paradis. EO-PEO EA Random chromozome initialization Random initialization for each of the active torsional angles with a value within a specified angle domain (typically 0. . 360). #include <eo. Init. h>. . . class Conformation. Init : public eo. Init< Conformation > { public: . . . void operator()( Conformation& conf ) { for ( unsigned int i =1; i <= conf. molecule->get. Nvars(); i++ ) { } }; . . . conf. chromosome->set. Chromo( i, eo: : rng. random( ANGLE_DOMAIN ) ); }

Asynchronous Island Model Parameters // migrations will occur periodically at every N generations eo. Periodic. Continue < Conformation > mig_cont ( MIGRATIONS_AT_N_GENERATIONS ); // selection of the emigrant conformations is performed as a stochastic tournament eo. Stoch. Tournament. Select < Conformation > mig_select_one; eo. Select. Number < Conformation > mig_select ( mig_select_one, NUMBER_OF_MIGRANTS ); // migrations are merged with the population - the worse individuals are discarded eo. Deterministic. Sa. DReplacement < Conformation > mig_elit_replace ( 0. 3, 0. 0, 0. 1, 0. 0 ); Async. Ils. peo. Async. Island. Mig < Conformation > island_mig ( mig_cont, mig_select, mig_elit_replace, ring_topology, alg. population ); EA // the migration stream will follow a ring topology Ring. Topology ring_topology; // to be activated at the end of every generation checkpoint->add ( island_mig ); peo. EA < Conformation > e. Alg (checkpoint, pop_eval, select, para. Transform, replacement ); island_mig. set. Owner ( e. Alg );

The Parallel Evaluation of the Population // functor for performing the evaluation of a specified conformation Conformation. Eval conformation. Eval; // wrapper for the evaluation functor – in addition it offers parallel evaluation peo. Para. Pop. Eval< Conformation > pop_eval ( conformation. Eval ); peo. Para. SGATransform< Conformation > para. Transform ( crossover, XOVER_RATE, mutation, MUTATION_RATE ); XML Grid Mapping … peo. EA < Conformation > e. Alg ( checkpoint, pop_eval, select, para. Transform, replacement ); <? xml version="1. 0"? > <schema> <group scheduler="0"> <node name="0" num_workers="0"> </node> <node name="1" num_workers="0"> <runner>1</runner>. . . </node> <node name="2" num_workers="1"> </node> <node name="3" num_workers="1">. . . </group> </schema> SCHEDULER ALGORITHMS WORKER NODES

End of story. . .
- Slides: 54