Lab 3 Jacobian estimation estimating autoregressive models estimating

Lab #3: - Jacobian estimation - estimating autoregressive models - estimating spatial filter models - estimating random effects models - mapping spatial filters Spatial statistics in practice Center for Tropical Ecology and Biodiversity, Tunghai University & Fushan Botanical Garden

SAS code for the Jacobian approximation FILENAME EIGEN 'D: JYU-SUMMERSCHOOL 2006LAB#3PR-EIG. TXT'; TITLE 'MATRIX C OR W JACOBIAN APPROXIMATION, IRREGULAR LATTICE'; ************************ * THE APPROPRIATE EIGENVALES MUST BE SELECTED * ************************; DATA STEP 0; INFILE EIGEN; INPUT ID LAMBDAC LAMBDAW; LAMBDA=LAMBDAW; X 0=1; RUN; PROC UNIVARIATE NOPRINT; VAR LAMBDA; OUTPUT OUT=EXTREMEL MAX=LMAX MIN=LMIN; RUN; PROC PRINT DATA=EXTREMEL; VAR LMAX LMIN; RUN; PROC MEANS DATA=STEP 0 NOPRINT; VAR X 0; OUTPUT OUT=STEP 0 A SUM=N; RUN; DATA EXTREMEL(REPLACE=YES); SET EXTREMEL; SET STEP 0 A; FINISH = 0. 999/LMAX; START = 0. 999/LMIN; INC = (FINISH - START)/201; RUN; DATA STEP 1; IF _N_=1 THEN SET EXTREMEL; SET STEP 0; ARRAY JACOB{202} JAC 1 -JAC 202; RHO = START; DO I = 1 TO 202; JACOB{I} = -LOG(1 - RHO*LAMBDA); RHO = RHO + INC; END; RUN; PROC MEANS NOPRINT; VAR JAC 1 -JAC 202; OUTPUT OUT=JACOB 1 MEAN=; RUN;

PROC TRANSPOSE OUT=JACOB 2; VAR JAC 1 -JAC 202; RUN; DATA STEP 2 (REPLACE=YES); SET EXTREMEL; DO RHO = START TO FINISH BY INC; OUTPUT; END; DROP START FINISH INC; RUN; DATA STEP 2 (REPLACE=YES); SET JACOB 2; SET STEP 2; J = COL 1; DROP COL 1; RUN; PROC GPLOT; PLOT J*RHO; RUN; PROC NLIN DATA=STEP 2 NOITPRINT MAXITER=15000 METHOD=MARQUARDT; PARMS A 1=0. 15 A 2=0. 15 D 1=1. 1 D 2=1. 1; BOUNDS D 1<2 , D 2<2 ; MODEL J = A 1*(LOG(1 - RHO*LMIN)/(RHO*LMIN) + 1 - D 1*LOG(1 - RHO*LMIN))+ A 2*(LOG(1 - RHO*LMAX)/(RHO*LMAX) + 1 - D 2*LOG(1 - RHO*LMAX)); OUTPUT OUT=TEMP 3 ESS=ESS 2 P=JHAT PARMS=A 1 A 2 D 1 D 2; RUN; PROC MEANS; VAR A 1 A 2 D 1 D 2; RUN; PROC GPLOT; PLOT J*RHO JHAT*RHO/OVERLAY; RUN; PROC UNIVARIATE DATA=STEP 2 NOPRINT; VAR J; OUTPUT OUT=TOTALSS CSS=TSS; RUN; DATA TEMP 3(REPLACE=YES); SET TOTALSS; SET TEMP 3; RESS = ESS 2/TSS; RUN; PROC PRINT; VAR ESS 2 TSS RESS; RUN; one of three possible approximation equations

SAS code for simultaneous autoregressive (SAR) modeling FILENAME ATTRIBUT 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-DEM&QUAD-DATA. TXT'; FILENAME CONN 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-CON. TXT'; FILENAME EIGEN 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-EIG. TXT'; TITLE 'SAR FOR PUERTO RICO DEM'; DATA STEP 1; INFILE ATTRIBUT; INPUT IDDEM MELEV SELEV U V QUAD NAME$; U=U/1000; V=V/1000; Y=LOG(MELEV+17. 5); * Y=(SELEV-25)**0. 5; RUN; PROC SORT OUT=STEP 1(REPLACE=YES); BY IDDEM; RUN; PROC STANDARD MEAN=0 STD=1 OUT=STEP 1(REPLACE=YES); VAR Y; RUN; DATA STEP 1 (REPLACE=YES); SET STEP 1; INFILE CONN; INPUT MUNUM 2 C 1 -C 73; ARRAY CONY{73} CY 1 -CY 73; ARRAY CON{73} C 1 -C 73; CSUM = 0; DO I=1 TO 73; CSUM = CSUM + CON{I}; CONY{I} = Y*CON{I}; END; RUN; PROC MEANS SUM; VAR CSUM; RUN; PROC PRINT; VAR NAME IDDEM MUNUM 2; RUN; PROC MEANS DATA=STEP 1 NOPRINT; VAR CY 1 -CY 73; OUTPUT OUT=CYOUT 1 SUM=CY 1 -CY 73; RUN; PROC TRANSPOSE DATA=CYOUT 1 PREFIX=WY OUT=CYOUT 2; VAR CY 1 -CY 73; RUN; DATA STEP 1 (REPLACE=YES); SET STEP 1; SET CYOUT 2; WY = WY 1/CSUM; RUN; PROC REG; MODEL Y=WY; RUN;
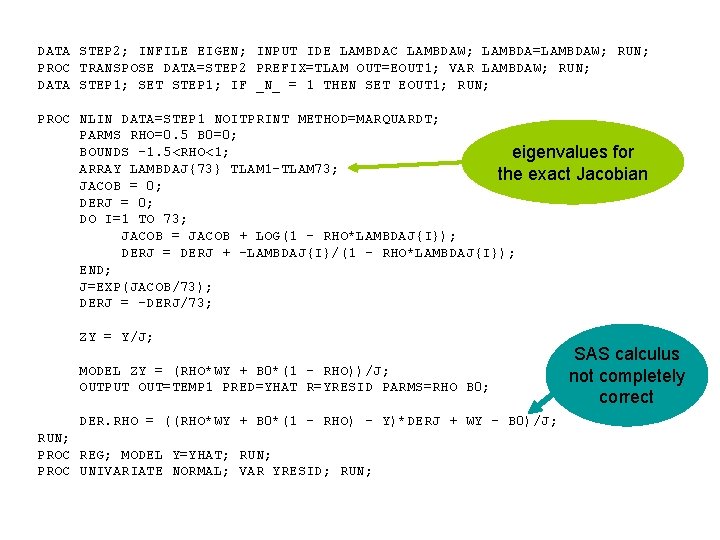
DATA STEP 2; INFILE EIGEN; INPUT IDE LAMBDAC LAMBDAW; LAMBDA=LAMBDAW; RUN; PROC TRANSPOSE DATA=STEP 2 PREFIX=TLAM OUT=EOUT 1; VAR LAMBDAW; RUN; DATA STEP 1; SET STEP 1; IF _N_ = 1 THEN SET EOUT 1; RUN; PROC NLIN DATA=STEP 1 NOITPRINT METHOD=MARQUARDT; PARMS RHO=0. 5 B 0=0; BOUNDS -1. 5<RHO<1; eigenvalues for ARRAY LAMBDAJ{73} TLAM 1 -TLAM 73; the exact Jacobian JACOB = 0; DERJ = 0; DO I=1 TO 73; JACOB = JACOB + LOG(1 - RHO*LAMBDAJ{I}); DERJ = DERJ + -LAMBDAJ{I}/(1 - RHO*LAMBDAJ{I}); END; J=EXP(JACOB/73); DERJ = -DERJ/73; ZY = Y/J; MODEL ZY = (RHO*WY + B 0*(1 - RHO))/J; OUTPUT OUT=TEMP 1 PRED=YHAT R=YRESID PARMS=RHO B 0; DER. RHO = ((RHO*WY + B 0*(1 - RHO) - Y)*DERJ + WY - B 0)/J; RUN; PROC REG; MODEL Y=YHAT; RUN; PROC UNIVARIATE NORMAL; VAR YRESID; RUN; SAS calculus not completely correct

from the Jacobian approximation ************* * * * JACOBIAN APPROXIMATION * **************; PROC MEANS NOPRINT DATA=STEP 2; VAR LAMBDA; OUTPUT OUT=EXTREMES MIN=LMIN MAX=LMAX; RUN; DATA STEP 1(REPLACE=YES); SET STEP 1; IF _N_=1 THEN SET EXTREMES(KEEP=LMIN LMAX); RUN; PROC NLIN DATA=STEP 1 NOITPRINT METHOD=MARQUARDT MAXITER=500; PARMS RHO=0. 5 B 0=0; BOUNDS -1<RHO<1; A 1 = 0. 4276827; A 2 = 0. 2998183; D 1 = 0. 9853629; D 2 IF RHO=0 THEN J=1; ELSE J = EXP(A 1*(LOG(1 - RHO*LMIN)/(RHO*LMIN) + A 2*(LOG(1 - RHO*LMAX)/(RHO*LMAX) + 1 IF RHO=0 THEN DERJ=0; ELSE DERJ = A 1*(-LOG(1 - RHO*LMIN)/(LMIN*RHO**2) + A 2*(-LOG(1 - RHO*LMAX)/(LMAX*RHO**2) + = 1. 0664149; 1 - D 1*LOG(1 - RHO*LMIN))+ D 2*LOG(1 - RHO*LMAX)) ); (RHO*D 1*LMIN - 1)/(RHO*(1 - RHO*LMIN)) ) + (RHO*D 2*LMAX - 1)/(RHO*(1 - RHO*LMAX)) ) ; ZY = Y*J; MODEL ZY = (RHO*WY + B 0*(1 - RHO))*J; OUTPUT OUT=TEMP 1 PRED=YHAT R=YRESID; DER. RHO = ( ( (RHO*WY + B 0*(1 - RHO) ) - Y)*DERJ + WY - B 0)*J; RUN;
The AR-SAR model specification 4 th order effect

SAS code for spatial filter: Gaussian RV FILENAME FILENAME ATTRIBUT 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-DEM&QUAD-DATA. TXT'; MC 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-MC-EIG. TXT'; CONN 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-CON. TXT'; EVEC 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-EIGENVECTORS. TXT'; OUTFILE 'D: JYU-SUMMERSCHOOL 2006LAB#3GAUSSIAN-SF. TXT'; OPTIONS LINESIZE=72; TITLE 'SPATIAL FILTER MODEL FOR PUERTO RICO: GAUSSIAN RV'; DATA STEP 1; INFILE ATTRIBUT; INPUT IDDEM MELEV SELEV U V QUAD NAME$; * Y=LOG(MELEV+17. 5); Y=(SELEV-25)**0. 5; Y 0=Y; RUN; PROC SORT DATA=STEP 1 OUT=STEP 1(REPLACE=YES); BY NAME; RUN; PROC UNIVARIATE NORMAL; VAR Y; RUN; PROC STANDARD MEAN=0 STD=1 OUT=STEP 1(REPLACE=YES); VAR Y 0; RUN; DATA STEP 1 (REPLACE=YES); SET STEP 1; INFILE EVEC LRECL=2048; INPUT IDE E 1 -E 73; RUN; ********************************* * THE SET OF CANDIDATE EIGENVECTORS IS DETERMINED BY MCADJ >= 0 * *********************************; PROC REG OUTEST=COEF; MODEL Y = E 1 -E 18/SELECTION=STEPWISE SLE=0. 10; OUTPUT OUT=TEMP P=YHAT R=YRESID; RUN; PROC UNIVARIATE NORMAL; VAR YRESID; RUN; may wish to make a Bonferroni adjustment here stepwise regression

PROC TRANSPOSE DATA=COEF PREFIX=B OUT=COEF 2; VAR E 1 -E 18; RUN; DATA COEF(REPLACE=YES); INFILE MC; INPUT ID LAM_MCM MC MCADJ; IF MCADJ<0. 25 THEN DELETE; RUN; DATA COEF(REPLACE=YES); SET COEF 2; IF B 1='. ' THEN DELETE; IF B 1=0 THEN DELETE; BSQ=B 1**2; EMC=BSQ*MC; P=1; RUN; PROC PRINT; RUN; PROC MEANS SUM NOPRINT; VAR LAM_MCM P EMC BSQ; OUTPUT OUT=EMC SUM=SUML P SPANUM SPADEN; RUN; PROC STANDARD DATA=TEMP MEAN=0 STD=1 OUT=TEMP(REPLACE=YES); VAR YRESID; RUN; DATA STEP 2(REPLACE=YES); INFILE CONN LRECL=1024; INPUT ID C 1 -C 73; RUN; DATA STEP 2(REPLACE=YES); SET STEP 2; SET TEMP(KEEP=YRESID); SET STEP 1(KEEP=Y 0); ARRAY CONN{73} C 1 -C 73; ARRAY ZC{73} ZC 1 -ZC 73; ARRAY Y 0 C{73} Y 0 C 1 -Y 0 C 73; CSUM=0; DO I=1 TO 73; CSUM = CSUM + CONN{I}; ZC{I} = YRESID*CONN{I}; Y 0 C{I} = Y 0*CONN{I}; END; Z=YRESID; X 0=1; RUN; PROC MEANS SUM NOPRINT; VAR CSUM X 0; OUTPUT OUT=CSUM SUM=CSUM N; RUN; PROC PRINT; VAR CSUM; RUN;

PROC DATA MEANS DATA=STEP 2 NOPRINT; VAR Y 0 C 1 -Y 0 C 73; OUTPUT OUT=Y 0 COUT 1 SUM=Y 0 C 1 -Y 0 C 73; RUN; TRANSPOSE DATA=Y 0 COUT 1 PREFIX=Y 0 C OUT=Y 0 COUT 2; VAR Y 0 C 1 -Y 0 C 73; RUN; MEANS DATA=STEP 2 NOPRINT; VAR ZC 1 -ZC 73; OUTPUT OUT=ZCOUT 1 SUM=ZC 1 -ZC 73; RUN; TRANSPOSE DATA=ZCOUT 1 PREFIX=ZC OUT=ZCOUT 2; VAR ZC 1 -ZC 73; RUN; STEP 2(REPLACE=YES); SET STEP 2(KEEP=X 0 Z CSUM Y 0); SET ZCOUT 2(KEEP=ZC 1); SET Y 0 COUT 2(KEEP=Y 0 C 1); Y 0 C=Y 0 C 1; ZC=ZC 1; DROP ZC 1; RUN; PROC REG DATA=STEP 2 OUTEST=DEN NOPRINT; MODEL CSUM=X 0/NOINT; RUN; PROC REG DATA=STEP 2 OUTEST=NUM 0 NOPRINT; MODEL Y 0 C=Y 0/NOINT; RUN; PROC REG DATA=STEP 2 OUTEST=NUM NOPRINT; MODEL ZC=Z/NOINT; RUN; DATA STEP 3; SET DEN; SET NUM 0; SET NUM; SET CSUM(KEEP=CSUM N); SET EMC(KEEP=SUML SPANUM SPADEN P); MC 0=Y 0/X 0; MC=Z/X 0; EMC = -(1+(N/CSUM)*SUML)/(N-P-1); ZMC = (MC-EMC)/SQRT(2/CSUM); MCSPA=SPANUM/SPADEN; RUN; PROC PRINT; VAR MC 0 MC EMC ZMC MCSPA; RUN; Residual diagnostics; MC distribution theory known for this case PROC STANDARD DATA=TEMP OUT=TEMP(REPLACE=YES) MEAN=0; VAR YHAT; RUN; DATA _NULL_; SET TEMP; FILE OUTFILE; PUT IDDEM Y YHAT; RUN; YHAT is the spatial filter; output for mapping purposes

SAS code for spatial filter: binomial RV FILENAME INDATA 'D: JYU-SUMMERSCHOOL 2006LAB#3PR-POP-1899&2000. TXT'; FILENAME EVEC 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-EIGENVECTORS. TXT'; FILENAME OUTFILE 'D: JYU-SUMMERSCHOOL 2006LAB#3BINOMIAL-SF. TXT'; TITLE 'SPATIAL FILTER MODEL FOR PUERTO RICO: BINOMIAL RV'; DATA STEP 1; INFILE INDATA; INPUT ID P 1899 U 1988 P 2000 U 2000 QUAD NAME$; Y=U 2000/P 2000; RUN; PROC SORT DATA=STEP 1 OUT=STEP 1(REPLACE=YES); BY NAME; RUN; DATA STEP 1 (REPLACE=YES); SET STEP 1; INFILE EVEC LRECL=2048; INPUT IDE E 1 -E 73; RUN; ********************************** * THE SET OF CANDIDATE EIGENVECTORS IS DETERMINED BY MCADJ >= 0. 25 * **********************************; PROC LOGISTIC; MODEL U 2000/P 2000 = E 1 -E 18/SELECTION=STEPWISE SLE=0. 10 SCALE=WILLIAMS; OUTPUT OUT=TEMP P=YHAT; RUN; PROC REG; MODEL Y=YHAT; RUN; stepwise selection PROC GENMOD; MODEL U 2000/P 2000 = E 1 E 3 E 4 E 10 E 14 E 18/DIST=BINOMIAL SCALE=DEVIANCE; OUTPUT OUT=SF XBETA=XBETA P=YHAT; RUN; DATA STEP 1(REPLACE=YES); SET STEP 1; Y=(U 2000 -1280)/(P 2000 -1057); TRY=LOG(Y/(1 -Y)); W=1/((P 2000 -1057)*Y*(1 -Y)); RUN; PROC REG; MODEL TRY=E 1 -E 18/NOINT SELECTION=STEPWISE SLE=0. 10; WEIGHT W; RUN; quasilikelihood normal approximation PROC STANDARD DATA=SF OUT=SF(REPLACE=YES) MEAN=0; VAR XBETA; RUN; DATA _NULL_; SET SF; FILE OUTFILE; YRESID=Y-YHAT; PUT ID Y XBETA YRESID; RUN; XBETA is the spatial filter; output for mapping purposes

SAS code for spatial filter: Poisson RV FILENAME INDATA 1 'D: JYU-SUMMERSCHOOL 2006LAB#3PR-POP-1899&2000. TXT'; INDATA 2 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-AREAS-COMPETITION. TXT'; EVEC 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-EIGENVECTORS. TXT'; OUTFILE 'D: JYU-SUMMERSCHOOL 2006LAB#3POISSON-SF. TXT'; OPTIONS LINESIZE=72; TITLE 'SPATIAL FILTER MODEL FOR PUERTO RICO: POISSON RV'; DATA STEP 1 A; INFILE INDATA 1; INPUT ID 1 P 1899 U 1988 P 2000 U 2000 QUAD NAME$; DENOM=1000000; RUN; PROC SORT DATA=STEP 1 A OUT=STEP 1 A(REPLACE=YES); BY NAME; RUN; DATA STEP 1 B; INFILE INDATA 2; INPUT ID 2 AREAM AREACT MCT_RATIO TRMCT_RATIO AREAPT MPT_RATIO TRMPT_RATIO NAME$; LNAREAM=LOG(AREAM); RUN; PROC SORT DATA=STEP 1 B OUT=STEP 1 B(REPLACE=YES); BY NAME; RUN; DATA STEP 1; MERGE STEP 1 A STEP 1 B; BY NAME; RUN; denominator for binomial approximation DATA STEP 1(REPLACE=YES); SET STEP 1; INFILE EVEC LRECL=2048; INPUT IDE E 1 -E 73; TRY=LOG(P 2000/AREAM - 441830); RUN; ********************************** * * * THE SET OF CANDIDATE EIGENVECTORS IS DETERMINED BY MCADJ >= 0. 25 * ***********************************; PROC GENMOD; MODEL P 2000 = E 1 -E 18/DIST=POISSON OFFSET=LNAREAM; RUN; offset variable must be converted to its log form no stepwise Poisson regression option in SAS

*************************************** * ORDER OF REMOVAL (BACKWARD ELIMINATION): 13, 5, 17, 14, 11, 15, 16, 9, 18 * ***************************************; PROC GENMOD; MODEL P 2000 = E 1 -E 4 E 6 -E 8 E 10 E 12/DIST=POISSON OFFSET=LNAREAM SCALE=DEVIANCE; RUN; quasi- likelihood *********************************** * ORDER OF REMOVAL (BACKWARD ELIMINATION): 17, 14, 11, 13, 15, 5, 9 * ***********************************; PROC GENMOD; MODEL P 2000 = E 1 -E 4 E 6 -E 8 E 10 E 12 E 16 E 18/DIST=NB OFFSET=LNAREAM; OUTPUT OUT=TEMP XBETA=XBETA P=YHAT; RUN; DATA TEMP(REPLACE=YES); SET TEMP; negative Y=P 2000/AREAM; binomial YHAT=YHAT/AREAM; RUN; binomial PROC REG; MODEL Y=YHAT; RUN; approximation PROC LOGISTIC; MODEL P 2000/DENOM = E 1 -E 18/SELECTION=BACKWARD OFFSET=LNAREAM SCALE=WILLIAMS SLS=0. 10; RUN; PROC REG; MODEL TRY=E 1 -E 18/SELECTION=BACKWARD SLS=0. 10; RUN; normal approximation PROC STANDARD DATA=TEMP OUT=TEMP(REPLACE=YES) MEAN=0; VAR XBETA; RUN; DATA _NULL_; SET TEMP; FILE OUTFILE; XBETA=XBETA-LNAREAM; XBETA is the spatial filter; YRESID=Y-YHAT; PUT ID 1 Y XBETA YRESID; output for mapping purposes RUN;
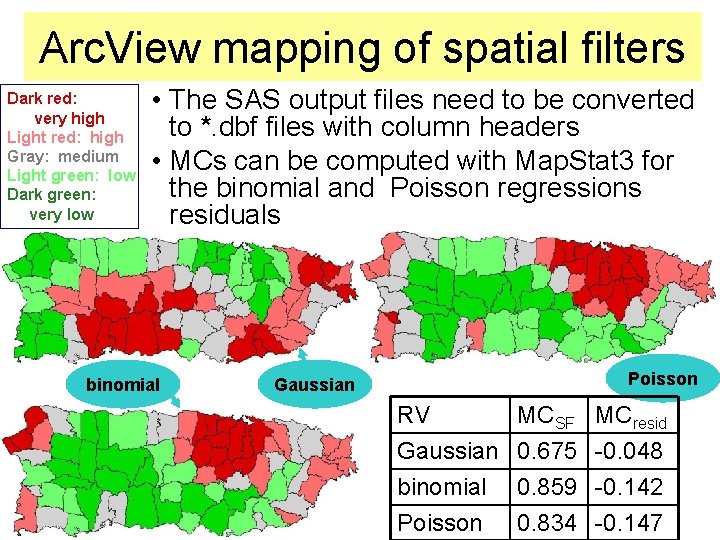
Arc. View mapping of spatial filters Dark red: very high Light red: high Gray: medium Light green: low Dark green: very low • The SAS output files need to be converted to *. dbf files with column headers • MCs can be computed with Map. Stat 3 for the binomial and Poisson regressions residuals binomial Poisson Gaussian RV Gaussian binomial Poisson MCSF 0. 675 0. 859 0. 834 MCresid -0. 048 -0. 142 -0. 147

SAS code for spatially structured random effects linear modeling FILENAME INDATA 1 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-AREAS-COMPETITION. TXT'; FILENAME INDATA 2 'D: JYU-SUMMERSCHOOL 2006LAB#1PR-DEM&QUAD-DATA. TXT'; TITLE 'MIXED MODEL FOR THE PR DATA'; ************************************* * READ IN GEOREFERENCED DATA; THEN CENTER THE SELEDTED ATTRIBUTE VARIABLE * *************************************; DATA STEP 1; INFILE INDATA 1 LRECL=1024; INPUT ID AREAM AREACT MCT_RATIO TRMCT_RATIO AREAPT MPT_RATIO TRMPT_RATIO NAME$; * Y = LOG(AREAM + 0. 001); * Y = TRMPT_RATIO; RUN; PROC SORT DATA=STEP 1 OUT=STEP 1(REPLACE=YES); BY NAME; RUN; DATA STEP 2; INFILE INDATA 2 LRECL=1024; INPUT IDDEM MELEV SELEV U V QUAD NAME$; U=U/1000; V=V/1000; * Y=LOG(MELEV+17. 5); Y=(SELEV-25)**0. 5; X 0=1; RUN; PROC SORT DATA=STEP 2 OUT=STEP 2(REPLACE=YES); BY NAME; RUN; DATA STEP 3; MERGE STEP 1 STEP 2; BY NAME; RUN; PROC REG; MODEL Y=/; RUN; PROC MIXED METHOD=ML MAXITER=500 COVTEST SCORING=10; MODEL Y=X 0/NOINT S; REPEATED/SUB=INTERCEPT; RUN;

maximum likelihood estimation PROC MIXED METHOD=ML MAXITER=500 COVTEST SCORING=10; MODEL Y=X 0/NOINT S; PARMS (13, 10); REPEATED/SUB=INTERCEPT TYPE=SP(EXP) (U V); RUN; PROC MIXED METHOD=ML MAXITER=500 COVTEST SCORING=10; MODEL Y=X 0/NOINT S; PARMS (13, 10); REPEATED/SUB=INTERCEPT TYPE=SP(GAU) (U V); RUN; PROC MIXED METHOD=ML MAXITER=500 COVTEST SCORING=10; MODEL Y=X 0/NOINT S; PARMS (13, 10); REPEATED/SUB=INTERCEPT TYPE=SP(SPH) (U V); RUN; PROC MIXED METHOD=ML MAXITER=500 COVTEST SCORING=10; MODEL Y=X 0/NOINT S; PARMS (0. 1, 13); REPEATED/SUB=INTERCEPT TYPE=SP(POW) (U V); RUN; PROC MIXED METHOD=ML MAXITER=500 COVTEST SCORING=10; MODEL Y=X 0/NOINT S; PARMS (13, 10, 1); REPEATED/SUB=INTERCEPT TYPE=SP(MATERN) (U V); RUN; initial parameter values random effect semivariogram model geocodings
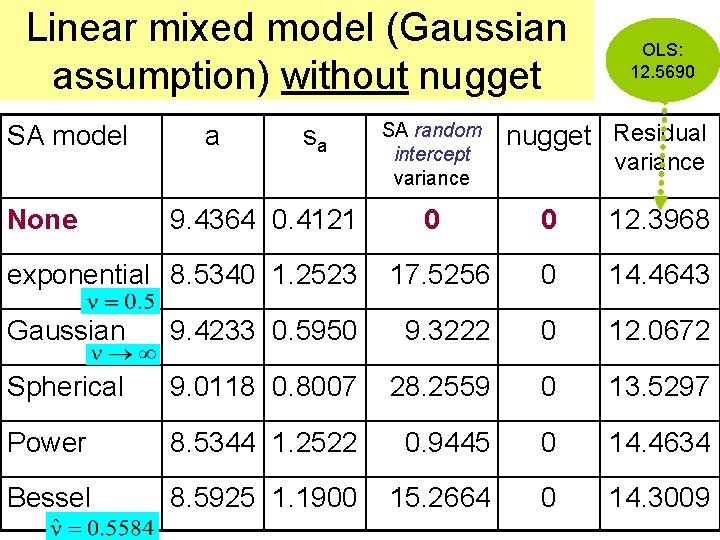
Linear mixed model (Gaussian assumption) without nugget SA model None a sa 9. 4364 0. 4121 exponential 8. 5340 1. 2523 SA random intercept variance OLS: 12. 5690 nugget Residual variance 0 0 12. 3968 17. 5256 0 14. 4643 Gaussian 9. 4233 0. 5950 9. 3222 0 12. 0672 Spherical 9. 0118 0. 8007 28. 2559 0 13. 5297 Power 8. 5344 1. 2522 0. 9445 0 14. 4634 Bessel 8. 5925 1. 1900 15. 2664 0 14. 3009
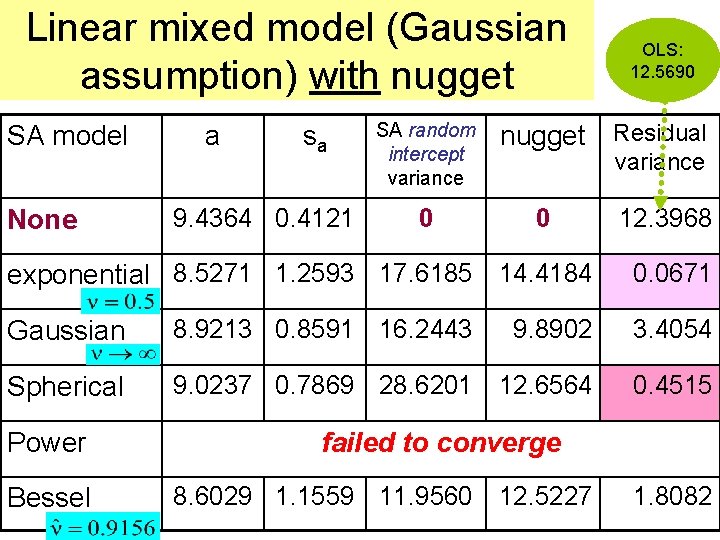
Linear mixed model (Gaussian assumption) with nugget SA model None a sa 9. 4364 0. 4121 SA random intercept variance 0 OLS: 12. 5690 nugget Residual variance 0 exponential 8. 5271 1. 2593 17. 6185 14. 4184 12. 3968 0. 0671 Gaussian 8. 9213 0. 8591 16. 2443 9. 8902 3. 4054 Spherical 9. 0237 0. 7869 28. 6201 12. 6564 0. 4515 Power Bessel failed to converge 8. 6029 1. 1559 11. 9560 12. 5227 1. 8082

SAS code for spatially structured random effects generalized linear modeling FILENAME INDATA 'D: JYU-SUMMERSCHOOL 2006LAB#3PR-URBAN-DATA. TXT'; FILENAME EVECS 'D: JYU-SUMMERSCHOOL 2006LAB#3PR-EXPANDED-EVECS. TXT'; OPTIONS LINESIZE=72; TITLE 'GENERALIZED LINEAR MIXED MODEL FOR THE PR DATA'; ********************************** * READ IN GEOREFERENCED DATA; THEN CENTER THE ELEVATION VARIABLE * **********************************; DATA STEP 1; INFILE INDATA; INPUT ID Y 1899 Y 1910 Y 1920 Y 1935 Y 1940 Y 1950 Y 1960 Y 1970 Y 1980 Y 1990 Y 2000 MELEV SELEV U V AAR NAME$; NUM=Y 1899; U=U/1000; V=V/1000; ELEV=LOG(MELEV/SELEV-0. 6); UTELEV=MELEV/SELEV-0. 6; IF AAR=1 THEN ISJ=1; ELSE ISJ=0; IF AAR=2 THEN IA =1; ELSE IA =0; IF AAR=3 THEN IM =1; ELSE IM =0; IF AAR=4 THEN IP =1; ELSE IP =0; IF AAR=5 THEN IC =1; ELSE IC =0; IA=IA-ISJ; IM=IM-ISJ; IP=IP-ISJ; IC=IC-ISJ; DENOM=100; Y=NUM/DENOM; P=Y; RUN;

PROC STANDARD MEAN=0 STD=1 OUT=STEP 1(REPLACE=YES); VAR ELEV; RUN; DATA STEP 2; INFILE EVECS LRECL=1024; INPUT IDE E 1 -E 45; RUN; DATA STEP 2(REPLACE=YES); SET STEP 2; SET STEP 1; RUN; DATA STEP 4; SET STEP 2(KEEP=ID DENOM ELEV UTELEV IA IP RUN; DATA STEP 3; SET STEP 2(KEEP=ID DENOM ELEV UTELEV IA IP RUN; DATA STEP 4(REPLACE=YES); SET STEP 4 STEP 3; RUN; DATA STEP 3(REPLACE=YES); SET STEP 2(KEEP=ID DENOM ELEV Y 1920; RUN; DATA STEP 4(REPLACE=YES); SET STEP 4 STEP 3; RUN; DATA STEP 3(REPLACE=YES); SET STEP 2(KEEP=ID DENOM ELEV Y 1935; RUN; DATA STEP 4(REPLACE=YES); SET STEP 4 STEP 3; RUN; DATA STEP 3(REPLACE=YES); SET STEP 2(KEEP=ID DENOM ELEV Y 1940; RUN; DATA STEP 4(REPLACE=YES); SET STEP 4 STEP 3; RUN; DATA STEP 3(REPLACE=YES); SET STEP 2(KEEP=ID DENOM ELEV Y 1950; RUN; DATA STEP 4(REPLACE=YES); SET STEP 4 STEP 3; RUN; DATA STEP 3(REPLACE=YES); SET STEP 2(KEEP=ID DENOM ELEV Y 1960; RUN; DATA STEP 4(REPLACE=YES); SET STEP 4 STEP 3; RUN; DATA STEP 3(REPLACE=YES); SET STEP 2(KEEP=ID DENOM ELEV Y 1970; RUN; DATA STEP 4(REPLACE=YES); SET STEP 4 STEP 3; RUN; DATA STEP 3(REPLACE=YES); SET STEP 2(KEEP=ID DENOM ELEV Y 1980; RUN; DATA STEP 4(REPLACE=YES); SET STEP 4 STEP 3; RUN; DATA STEP 3(REPLACE=YES); SET STEP 2(KEEP=ID DENOM ELEV Y 1990; RUN; DATA STEP 4(REPLACE=YES); SET STEP 4 STEP 3; RUN; DATA STEP 3(REPLACE=YES); SET STEP 2(KEEP=ID DENOM ELEV Y 2000; RUN; DATA STEP 4(REPLACE=YES); SET STEP 4 STEP 3; RUN; PROC SORT OUT=STEP 5; BY ID DESCENDING TIME; RUN; IM IC Y 1899 E 1 -E 45); NUM=Y 1899; TIME=1899; DROP Y 1899; IM IC Y 1910 E 1 -E 45); NUM=Y 1910; TIME=1910; DROP Y 1910; UTELEV IP IM IC Y 1920 E 1 -E 45); NUM=Y 1920; TIME=1920; DROP UTELEV IP IM IC Y 1930 E 1 -E 45); NUM=Y 1930; TIME=1930; DROP UTELEV IP IM IC Y 1935 E 1 -E 45); NUM=Y 1935; TIME=1935; DROP UTELEV IP IM IC Y 1940 E 1 -E 45); NUM=Y 1940; TIME=1940; DROP UTELEV IP IM IC Y 1950 E 1 -E 45); NUM=Y 1950; TIME=1950; DROP UTELEV IP IM IC Y 1960 E 1 -E 45); NUM=Y 1960; TIME=1960; DROP UTELEV IP IM IC Y 1970 E 1 -E 45); NUM=Y 1970; TIME=1970; DROP UTELEV IP IM IC Y 1980 E 1 -E 45); NUM=Y 1980; TIME=1980; DROP UTELEV IP IM IC Y 1990 E 1 -E 45); NUM=Y 1990; TIME=1990; DROP UTELEV IP IM IC Y 2000 E 1 -E 45); NUM=Y 2000; TIME=2000; DROP

PROC NLMIXED DATA=STEP 5; PARMS B 0=0 BE=0 BIM=0 BIP=0 BIC=0 S 2 U=1 BE 1 =0 BE 2 =0 BE 3 =0 BE 4 =0 BE 5 =0 BE 6 =0 BE 7 =0 BE 9 =0 BE 10=0 BE 11=0 BE 16=0 BE 17=0 BE 18=0 BE 19=0 BE 20=0 BE 21=0 BE 23=0 BE 26=0 BE 27=0 BE 33=0 BE 35=0; ETA = B 0 + ALPHA + BE*ELEV + BIM*IM + BIP*IP + BIC*IC + BE 1*E 1+BE 2*E 2+BE 3*E 3+BE 4*E 4+BE 5*E 5+BE 6*E 6+BE 7*E 7+BE 9*E 9+BE 10*E 10+BE 11*E 11+BE 16*E 16+ BE 17*E 17+BE 18*E 18+BE 19*E 19+BE 20*E 20+BE 21*E 21+BE 23*E 23+BE 26*E 26+BE 27*E 27+BE 33*E 33+BE 35*E 35; P = EXP(ETA)/(1+EXP(ETA)); MODEL NUM ~ BINOMIAL(100, P); RANDOM ALPHA ~ NORMAL(0, S 2 U) SUBJECT=ID; PREDICT P OUT=POUT; RUN; PROC NLMIXED DATA=STEP 5; PARMS B 0=0 BE=0 BIM=0 BIC=0 S 2 U=1 BE 2 =0 BE 19=0 ETA = B 0 + ALPHA + BE*ELEV + BIM*IM + BIC*IC + BE 2*E 2+ BE 19*E 19+ P = EXP(ETA)/(1+EXP(ETA)); MODEL NUM ~ BINOMIAL(100, P); RANDOM ALPHA ~ NORMAL(0, S 2 U) SUBJECT=ID; PREDICT P OUT=POUT; RUN; PROC DATA PROC BE 6 =0 BE 10=0 BE 11=0 BE 35=0; BE 6*E 6+ BE 10*E 10+BE 11*E 11+ SORT OUT=POUT(REPLACE=YES); BY TIME ID; RUN; POUT(REPLACE=YES); SET POUT; SET STEP 4(KEEP=NUM); P=NUM/100; RUN; REG; MODEL P=PRED; BY TIME; RUN; GPLOT; PLOT P*PRED; BY TIME; RUN; PROC GENMOD DATA=STEP 2; MODEL NUM/DENOM= ELEV E 1 E 4 E 6 E 7 E 11 E 17 E 23 E 26 E 33/DIST=B SCALE=P; OUTPUT OUT=POUT P=PRED; RUN; BE 35*E 35;
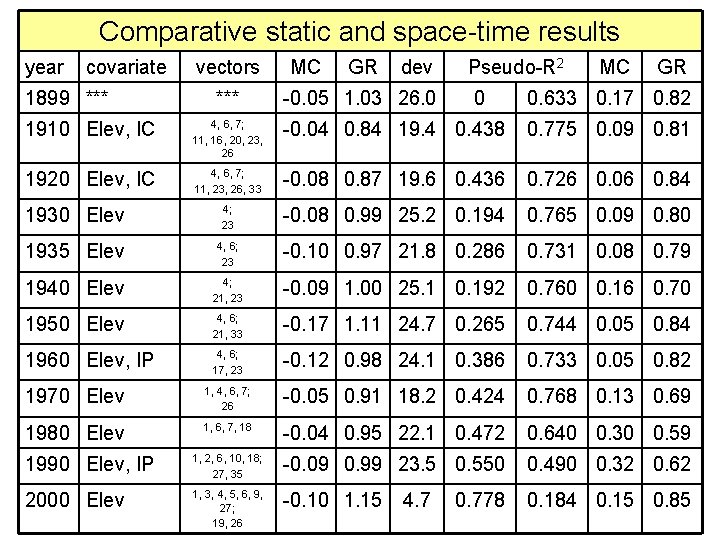
Comparative static and space-time results year covariate 1899 *** vectors *** MC GR dev -0. 05 1. 03 26. 0 Pseudo-R 2 0 MC GR 0. 633 0. 17 0. 82 1910 Elev, IC 4, 6, 7; 11, 16, 20, 23, 26 -0. 04 0. 84 19. 4 0. 438 0. 775 0. 09 0. 81 1920 Elev, IC 4, 6, 7; 11, 23, 26, 33 -0. 08 0. 87 19. 6 0. 436 0. 726 0. 06 0. 84 1930 Elev 4; 23 -0. 08 0. 99 25. 2 0. 194 0. 765 0. 09 0. 80 1935 Elev 4, 6; 23 -0. 10 0. 97 21. 8 0. 286 0. 731 0. 08 0. 79 1940 Elev 4; 21, 23 -0. 09 1. 00 25. 1 0. 192 0. 760 0. 16 0. 70 1950 Elev 4, 6; 21, 33 -0. 17 1. 11 24. 7 0. 265 0. 744 0. 05 0. 84 1960 Elev, IP 4, 6; 17, 23 -0. 12 0. 98 24. 1 0. 386 0. 733 0. 05 0. 82 1970 Elev 1, 4, 6, 7; 26 -0. 05 0. 91 18. 2 0. 424 0. 768 0. 13 0. 69 1980 Elev 1, 6, 7, 18 -0. 04 0. 95 22. 1 0. 472 0. 640 0. 30 0. 59 1990 Elev, IP 1, 2, 6, 10, 18; 27, 35 -0. 09 0. 99 23. 5 0. 550 0. 490 0. 32 0. 62 2000 Elev 1, 3, 4, 5, 6, 9, 27; 19, 26 -0. 10 1. 15 0. 184 0. 15 0. 85 4. 7 0. 778
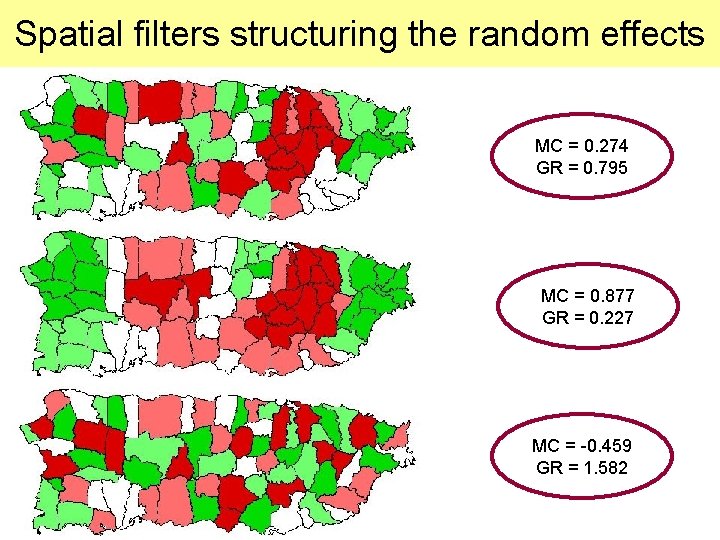
Spatial filters structuring the random effects MC = 0. 274 GR = 0. 795 MC = 0. 877 GR = 0. 227 MC = -0. 459 GR = 1. 582
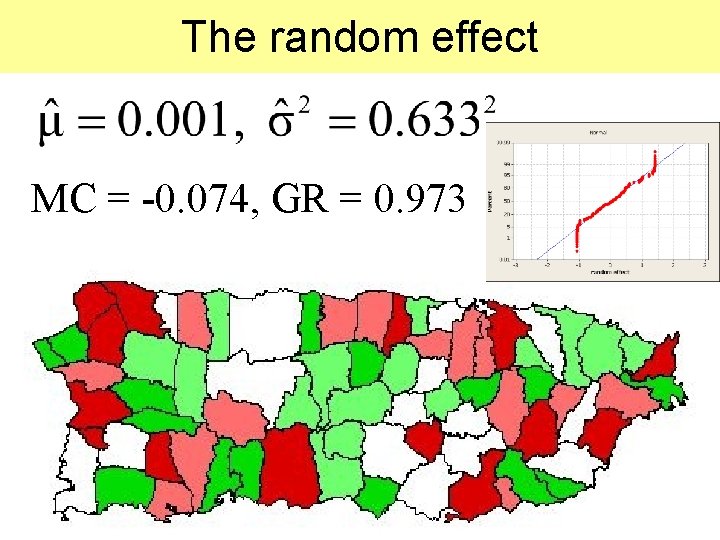
The random effect MC = -0. 074, GR = 0. 973

What you should have learned in today’s lab: 1. Calculate the J approximation for a given connectivity matrix (both C and W) 2. Normal approximation: estimate AR, SAR, AR -SAR and SF models 3. Estimate a SF GLM (Poisson or binomial) 4. Estimate a univariate LMM 5. Estimate a bivariate (time) SF-GLMM 6. Construct maps of the SFs 7. Construct maps of the random effects
- Slides: 25