KNOWME KNOWledgeMapExplorer Semantic Browsing of Integrated Data using
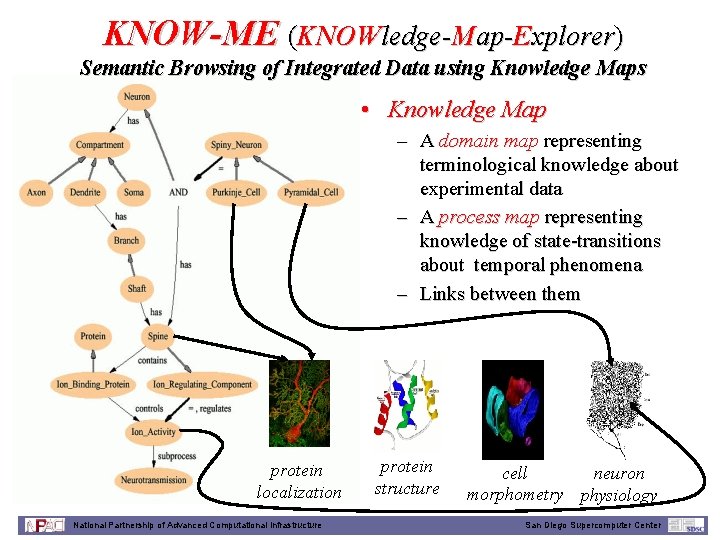
KNOW-ME (KNOWledge-Map-Explorer) Semantic Browsing of Integrated Data using Knowledge Maps • Knowledge Map – A domain map representing terminological knowledge about experimental data – A process map representing knowledge of state-transitions about temporal phenomena – Links between them protein localization National Partnership of Advanced Computational Infrastructure protein structure cell neuron morphometry physiology San Diego Supercomputer Center
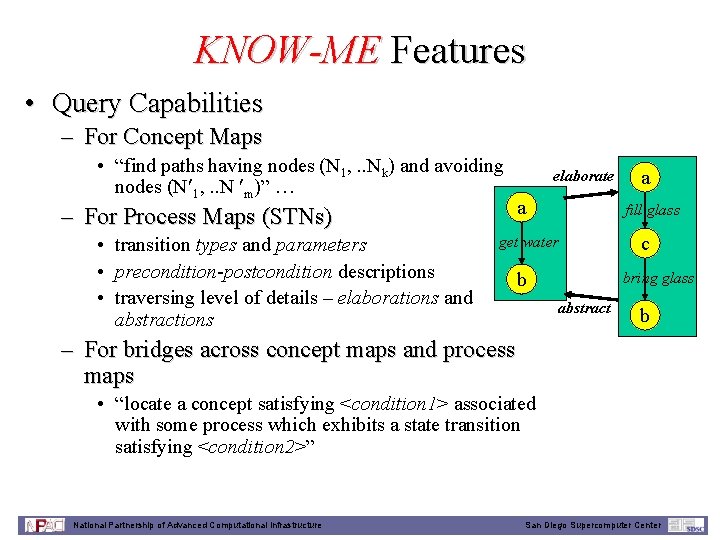
KNOW-ME Features • Query Capabilities – For Concept Maps • “find paths having nodes (N 1, . . Nk) and avoiding nodes (N 1, . . N m)” … – For Process Maps (STNs) • transition types and parameters • precondition-postcondition descriptions • traversing level of details – elaborations and abstractions elaborate a a fill glass get water b c bring glass abstract b – For bridges across concept maps and process maps • “locate a concept satisfying <condition 1> associated with some process which exhibits a state transition satisfying <condition 2>” National Partnership of Advanced Computational Infrastructure San Diego Supercomputer Center
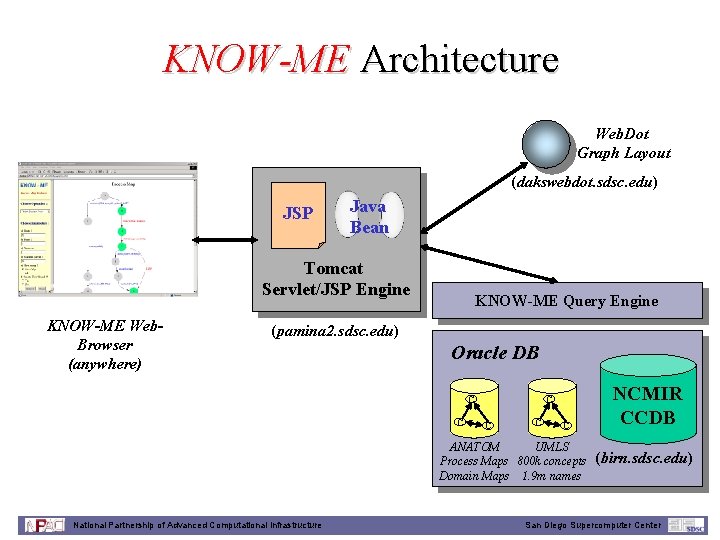
KNOW-ME Architecture Web. Dot Graph Layout (dakswebdot. sdsc. edu) JSP Java Bean Tomcat Servlet/JSP Engine KNOW-ME Web. Browser (anywhere) KNOW-ME Query Engine (pamina 2. sdsc. edu) Oracle DB C C C UMLS ANATOM Process Maps 800 k concepts Domain Maps 1. 9 m names National Partnership of Advanced Computational Infrastructure NCMIR CCDB (birn. sdsc. edu) San Diego Supercomputer Center
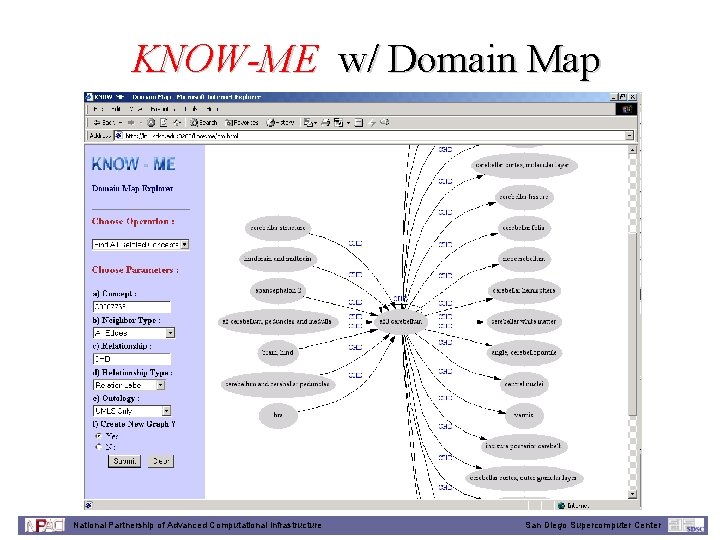
KNOW-ME w/ Domain Map National Partnership of Advanced Computational Infrastructure San Diego Supercomputer Center
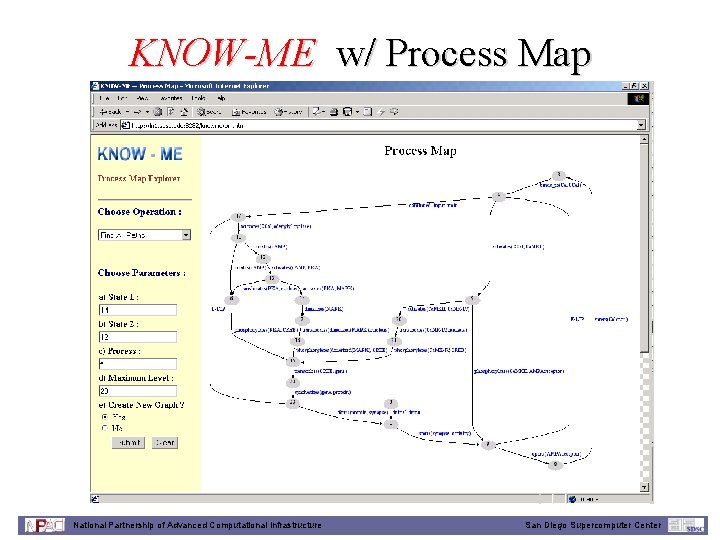
KNOW-ME w/ Process Map National Partnership of Advanced Computational Infrastructure San Diego Supercomputer Center
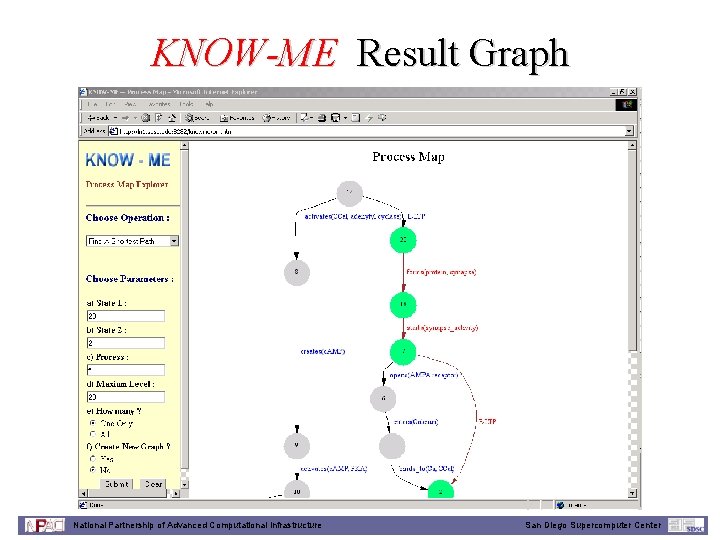
KNOW-ME Result Graph National Partnership of Advanced Computational Infrastructure San Diego Supercomputer Center
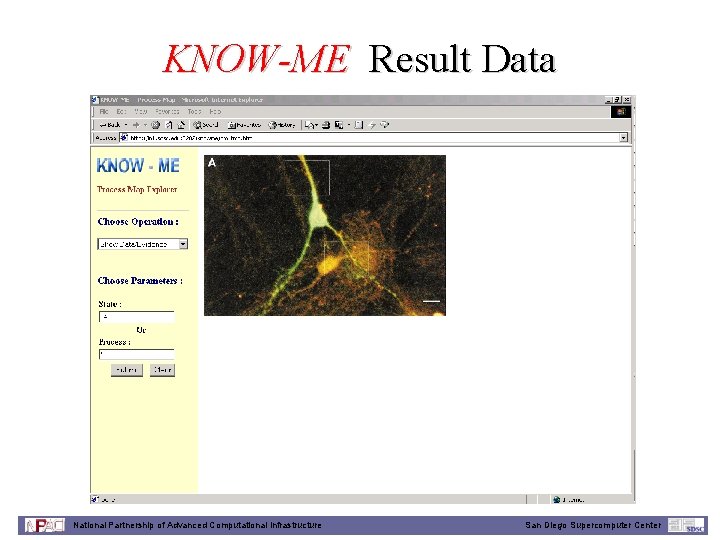
KNOW-ME Result Data National Partnership of Advanced Computational Infrastructure San Diego Supercomputer Center

Summary: KNOW-ME Features • “Semantic” Browsing of Data: – Data “about” a concept • “All electromicrographs of spiny dendrites” – Data “about” a process • “All electrode recordings that show ‘long-term-potentiation’ ” – Data that is “evidence of” a process • “All electromicrographs that suggest ‘synapse generation’ occurs during LTP” Xufei Qian, Bertram Ludäscher, Maryann E. Martone, Amarnath Gupta San Diego Supercomputer Center & Department of Neurosciences University of California, San Diego National Partnership of Advanced Computational Infrastructure San Diego Supercomputer Center

Some Related References: Mediation of Neuroscience Data • • • Model-Based Mediation with Domain Maps, B. Ludäscher, A. Gupta, M. E. Martone, 17 th Intl. Conference on Data Engineering (ICDE), Heidelberg, Germany, IEEE Computer Society, April 2001. Navigating Virtual Information Sources with Know-ME, X. Qian, B. Ludäscher, M. E. Martone, A. Gupta, demonstration track, Intl. Conference on Extending Database Technology (EDBT), Prague, Czech Republic, March 2002. Model-Based Information Integration in a Neuroscience Mediator System, B. Ludäscher, A. Gupta, M. E. Martone, demonstration track, 26 th Intl. Conference on Very Large Databases (VLDB), Cairo, Egypt, September 2000. Knowledge-Based Integration of Neuroscience Data Sources, A. Gupta, B. Ludäscher, M. E. Martone, 12 th Intl. Conference on Scientific and Statistical Database Management (SSDBM), Berlin, Germany, IEEE Computer Society, July 2000. A Cell-Centered Database for Electron Tomographic Data, M. E. Martone, A. Gupta, M. Wong, X. Qian, G. Sosinsky, S. Lamont, B. Ludäscher , and M. H. Ellisman. Journal of Structural Biology, 2002. to appear National Partnership of Advanced Computational Infrastructure San Diego Supercomputer Center
- Slides: 9