KMe D A KnowledgeBased Multimedia Medical Database System

KMe. D: A Knowledge-Based Multimedia Medical Database System Wesley W. Chu Computer Science Department University of California, Los Angeles http: //www. cobase. cs. ucla. edu 1

KMe. D A Knowledge-Based Multimedia Medical Distributed Database System October 1, 1991 to September 30, 1993 A Cooperative, Spatial, Evolutionary Medical Database System July 1, 1993 to June 30, 1997 Knowledge-Based Image Retrieval with Spatial and Temporal Constructs May 1, 1997 to April 30, 2001 Wesley W. Chu Alfonso F. Cardenas Ricky K. Taira Computer Science Department of Radiological Sciences 2

Research Team Students John David N. Dionisio Chih-Cheng Hsu David Johnson Christine Chih Collaborators Computer Science Department Alfonso F. Cardenas UCLA Medical School Denise Aberle, MD Robert Lufkin, MD Ricky K. Taira, MD 3

Significance Query multimedia data based on image content and spatial predicates Use domain knowledge to relax and interpret medical queries Present integrated view of multiple temporal and evolutionary data in a timeline metaphor 4

Overview Image retrieval by feature and content Query relaxation Spatial query answering Similarity query answering Visual query interface Timeline interface Sample cases 5

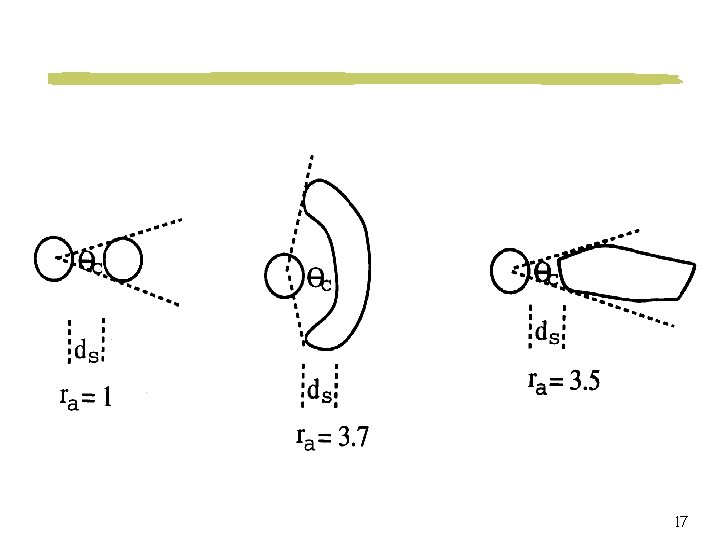
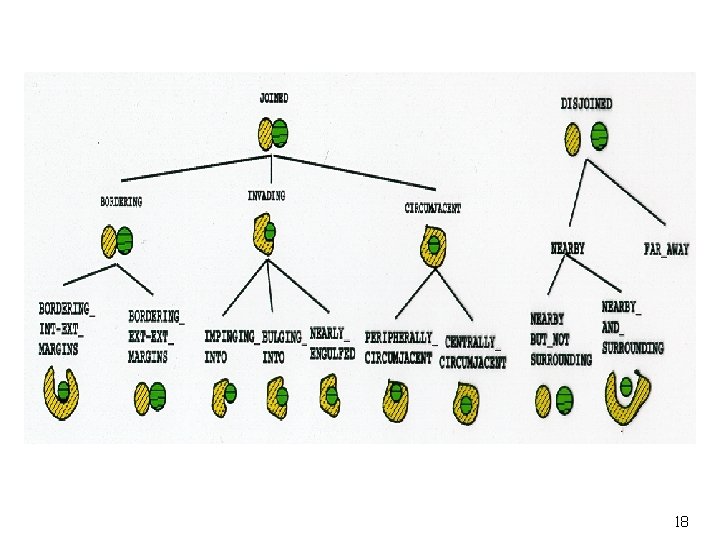
Image Retrieval by Content Features size, shape, texture, density, histology Spatial Relations angle of coverage, shortest distance, overlapping ratio, contact ratio, relative direction Evolution of Object Growth fusion, fission 6
7
8
9
10

Characteristics of Medical Queries Multimedia Temporal Evolutionary Spatial Imprecise 11
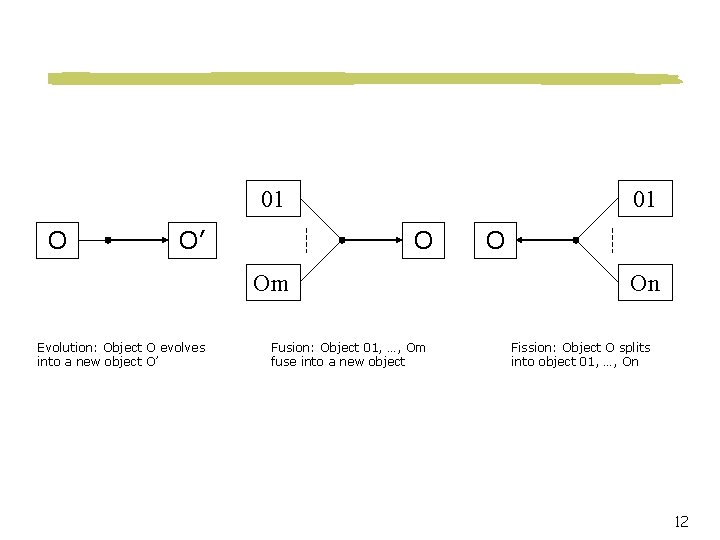
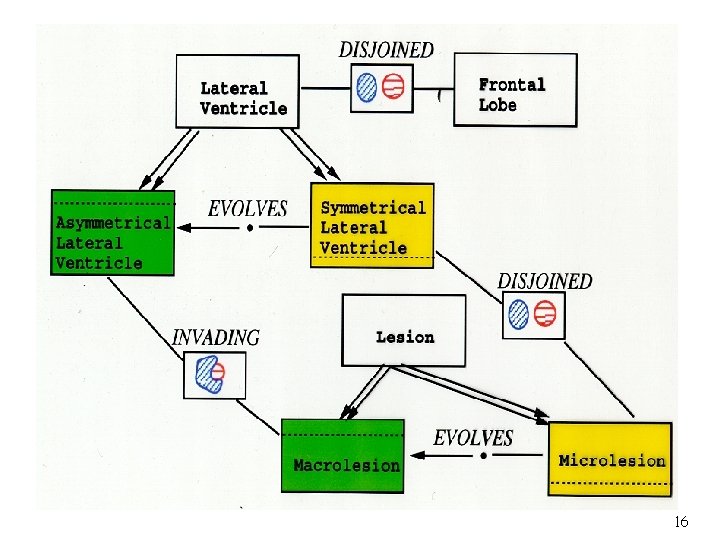
01 O O’ 01 O Om Evolution: Object O evolves into a new object O’ Fusion: Object 01, …, Om fuse into a new object O On Fission: Object O splits into object 01, …, On 12
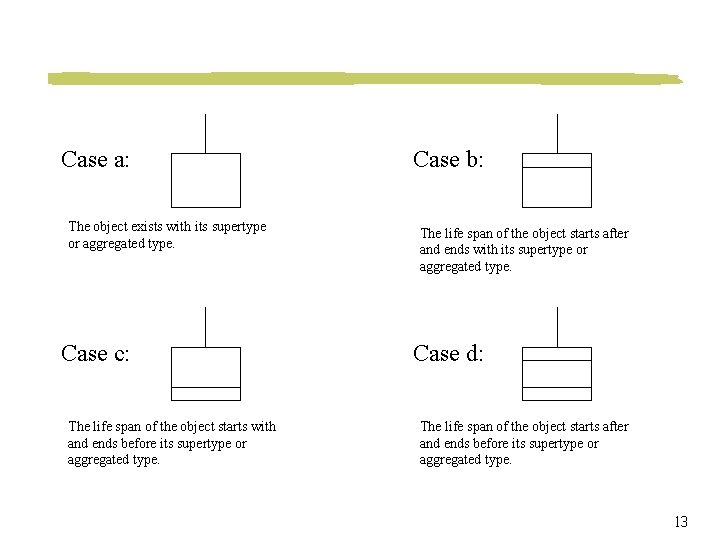
Case a: The object exists with its supertype or aggregated type. Case c: The life span of the object starts with and ends before its supertype or aggregated type. Case b: The life span of the object starts after and ends with its supertype or aggregated type. Case d: The life span of the object starts after and ends before its supertype or aggregated type. 13
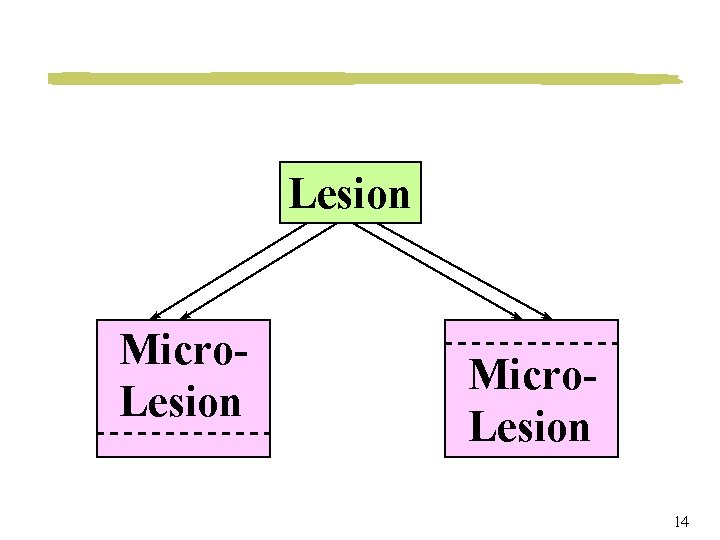
Lesion Micro. Lesion 14
15
16
17
18

Query Modification Techniques Relaxation Generalization Specialization Association 19
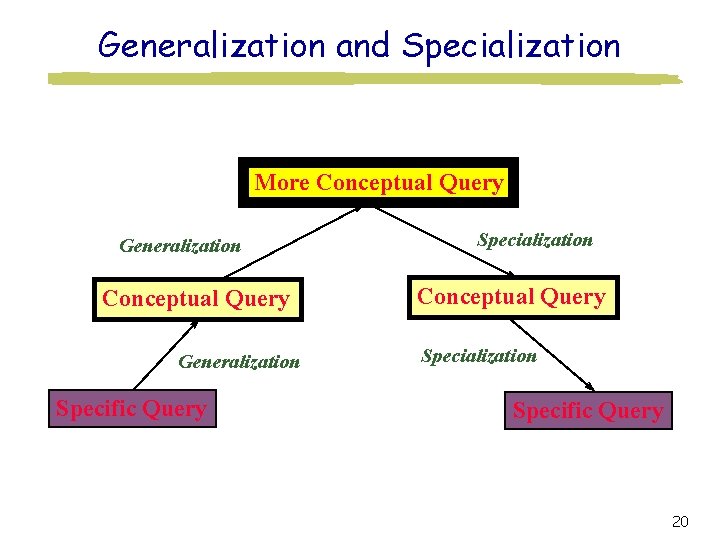
Generalization and Specialization More Conceptual Query Generalization Specific Query Specialization Conceptual Query Specialization Specific Query 20

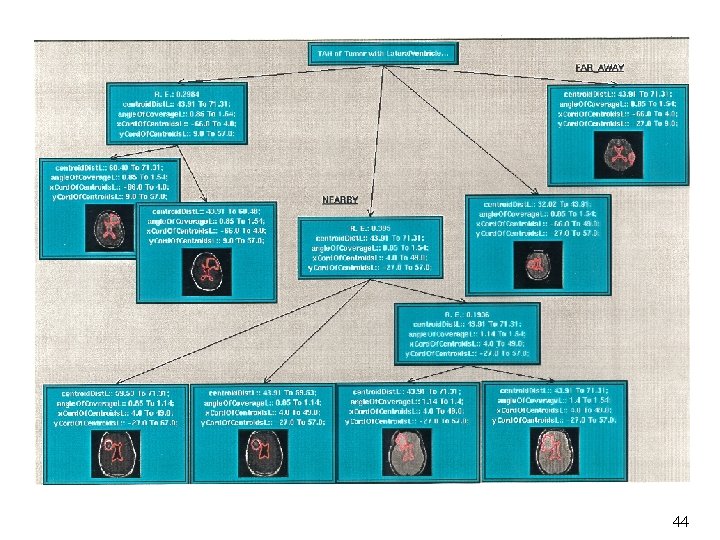
Type Abstraction Hierarchy Presents abstract view of Types Attribute values Image features Temporal and evolutionary behavior Spatial relationships among objects Provides multi-level knowledge representation 21

TAH Generation for Numerical Attribute Values Relaxation Error Difference between the exact value and the returned approximate value The expected error is weighted by the probability of occurrence of each value DISC (Distribution Sensitive Clustering) is based on the attribute values and frequency distribution of the data 22

TAH Generation for Numerical Attribute Values (cont. ) 2 Computation Complexity: O(n ), where n is the number of distinct value in a cluster DISC performs better than Biggest Cap (value only) or Max Entropy (frequency only) methods MDISC is developed for multiple attribute TAHs. Computation Complexity: O(mn 2), where m is the number of attributes 23
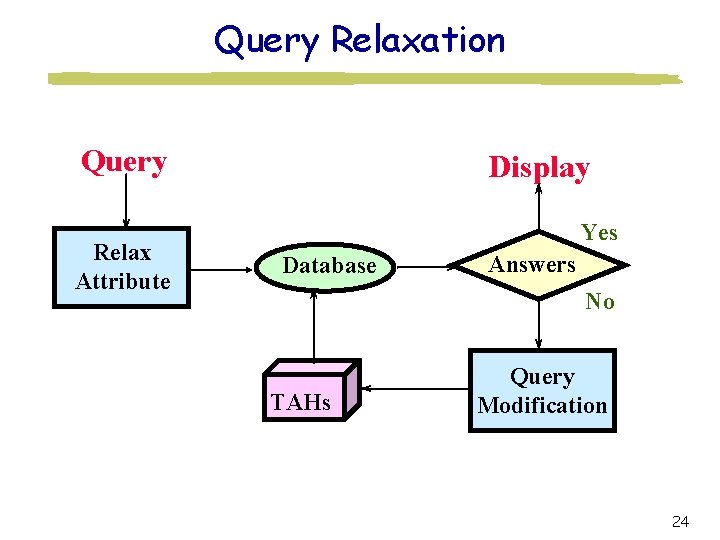
Query Relaxation Query Relax Attribute Display Yes Database Answers No TAHs Query Modification 24

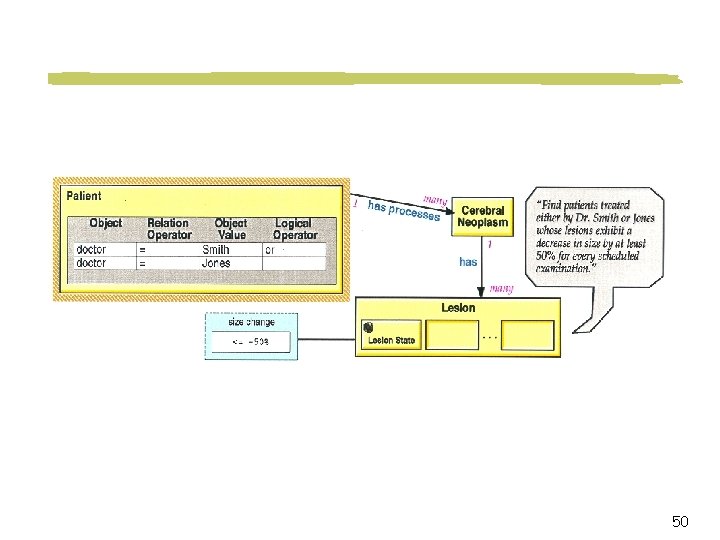
Cooperative Querying for Medical Applications Query Find the treatment used for the tumor similar-to (loc, size) X 1 on 12 year-old Korean males. Relaxed Query Find the treatment used for the tumor Class X on preteen Asians. Association The success rate, side effects, and cost of the treatment. 25
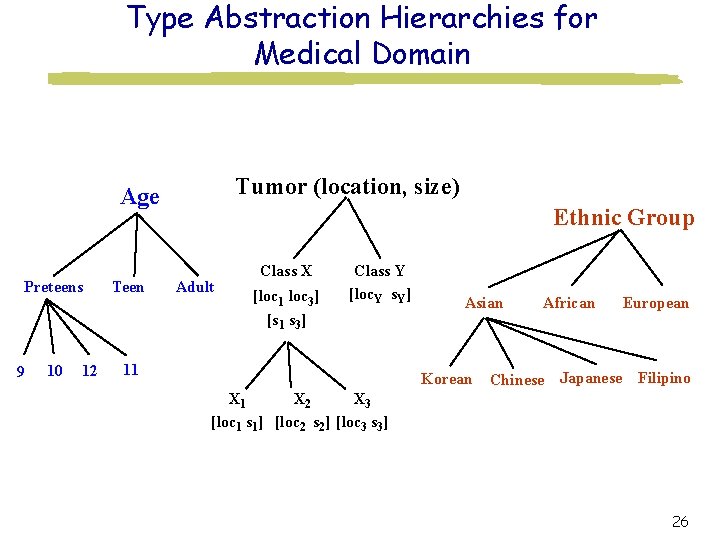
Type Abstraction Hierarchies for Medical Domain Tumor (location, size) Age Preteens 9 10 12 Teen Ethnic Group Adult Class X [loc 1 loc 3] [s 1 s 3] Class Y [loc. Y s. Y] 11 Asian Korean African Chinese Japanese European Filipino X 3 X 1 X 2 [loc 1 s 1] [loc 2 s 2] [loc 3 s 3] 26
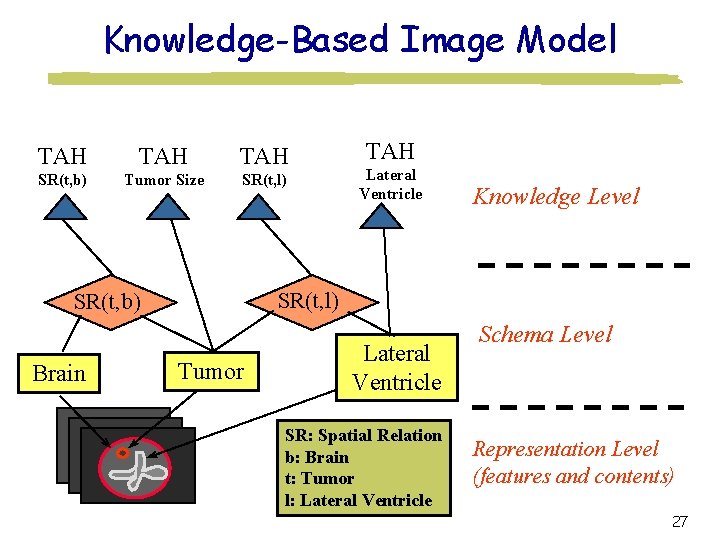
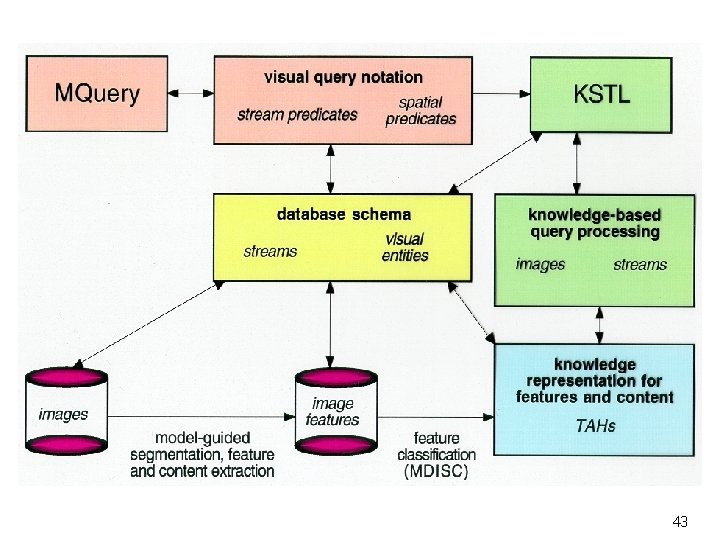
Knowledge-Based Image Model TAH TAH SR(t, b) Tumor Size SR(t, l) Lateral Ventricle Knowledge Level SR(t, l) SR(t, b) Brain TAH Tumor Lateral Ventricle SR: Spatial Relation b: Brain t: Tumor l: Lateral Ventricle Schema Level Representation Level (features and contents) 27
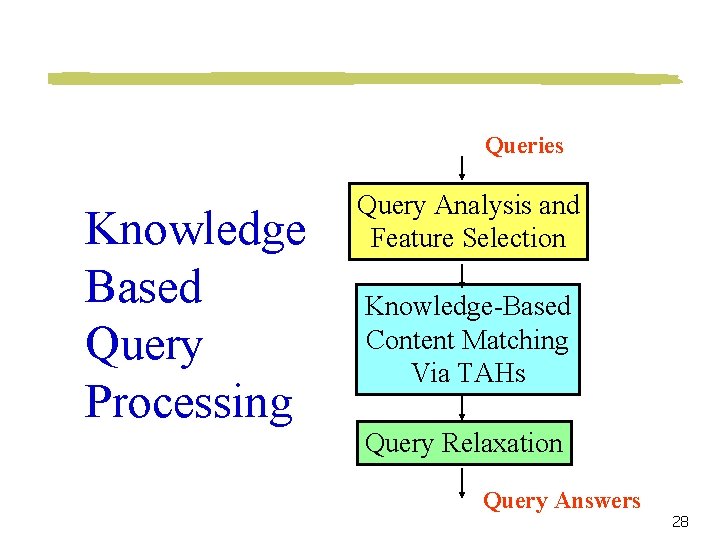
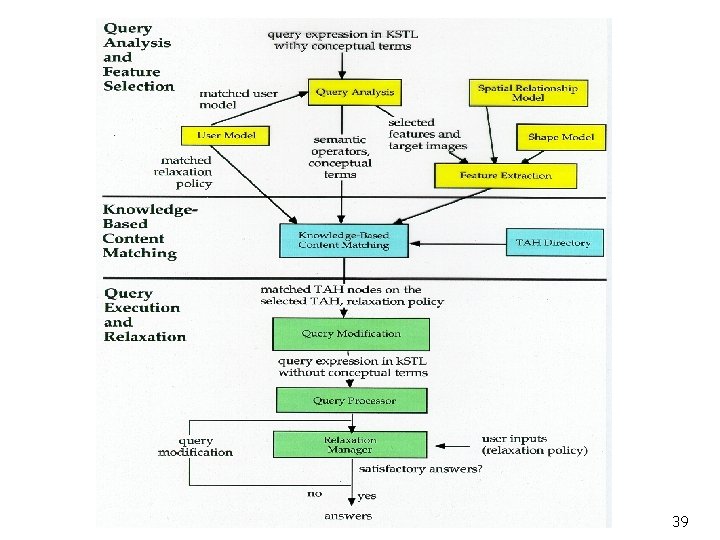
Queries Knowledge Based Query Processing Query Analysis and Feature Selection Knowledge-Based Content Matching Via TAHs Query Relaxation Query Answers 28

User Model To customize query conditions and knowledge-based query processing User type Default Parameter Values Feature and Content Matching Policies Complete Match Partial Match 29
User Model (cont. ) Relaxation Control Policies Relaxation Order Unrelaxable Object Preference List Measure for Ranking 30
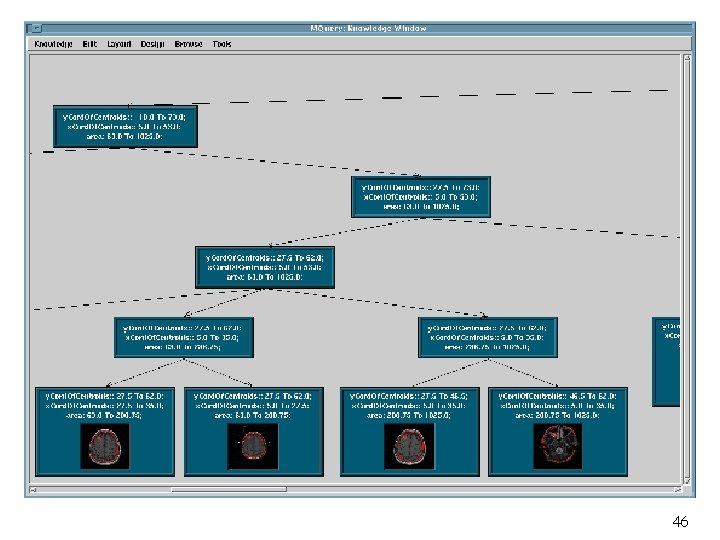
31
32

Query Preprocessing Segment and label contours for objects of interest Determine relevant features and spatial relationships (e. g. , location, containment, intersection) of the selected objects Organize the features and spatial relationships of objects into a feature database Classify the feature database into a Type Abstraction Hierarchy (TAH) 33

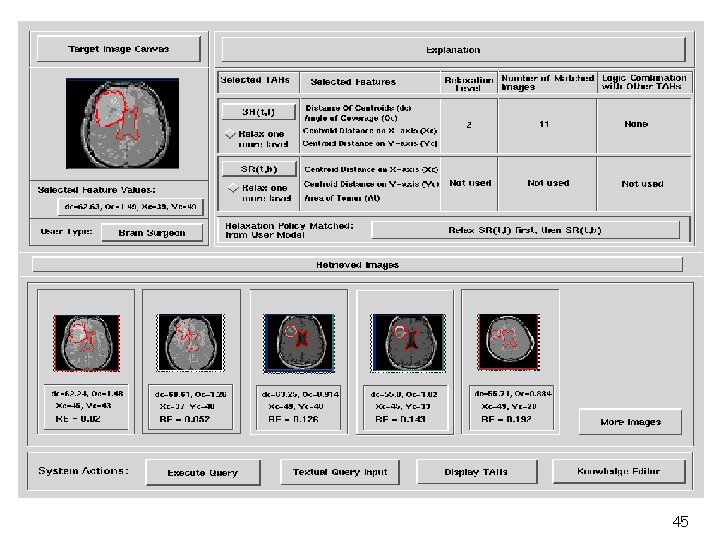
Similarity Query Answering Determine relevant features based on query input Select TAH based on these features Traverse through the TAH nodes to match all the images with similar features in the database Present the images and rank their similarity (e. g. , by mean square error) 34

Spatial Query Answering Preprocessing Draw and label contours for objects of interest Determine relevant features and spatial relationships (e. g. , location, containment, intersection) of the selected objects Organize the features and spatial relationships of objects into a feature database Classify the feature database into a type abstraction hierarchy (TAH) 35

Spatial Query Answering (cont. ) Processing Select TAH based on t he query conditions and context Search nodes to match the query conditions Return images linked to the TAH node 36

Similarity Query Answering Preprocessing Select objects and specify features of interest in the image Create a feature database of the selected objects for all images Classify the feature databases as type abstraction hierarchies 37

Similarity Query Answering (cont. ) Processing Determine relevant features based on query input Select TAH based on these features (interact with user to resolve ambiguity) Traverse through the TAH nodes to match all the images with similar features in the databases Present the images and rank their similarity (e. g. , by mean square error) 38
39

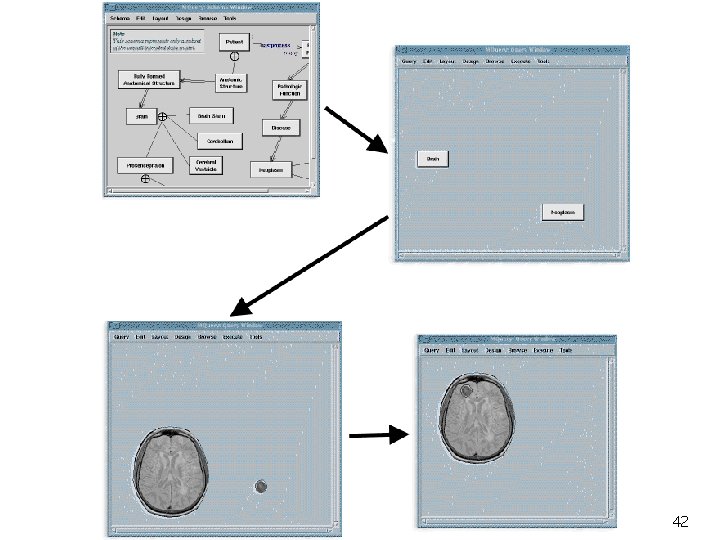
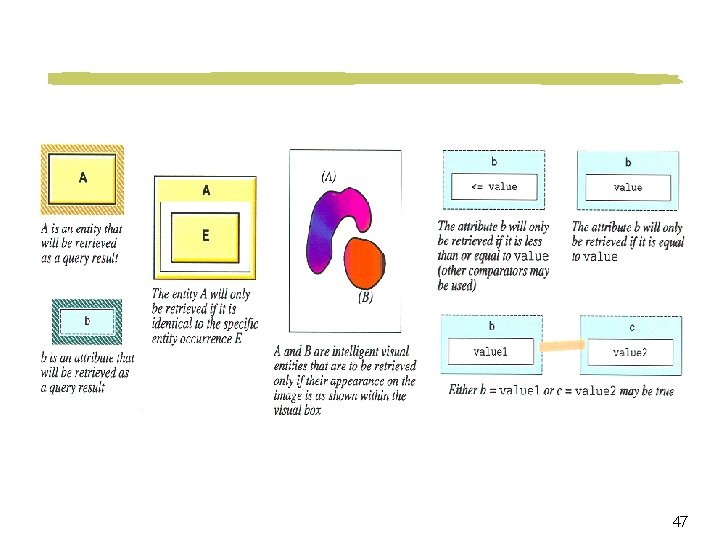
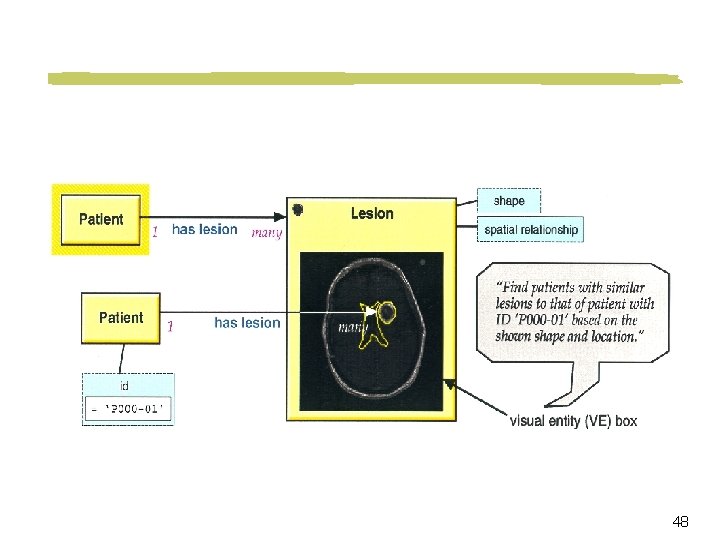
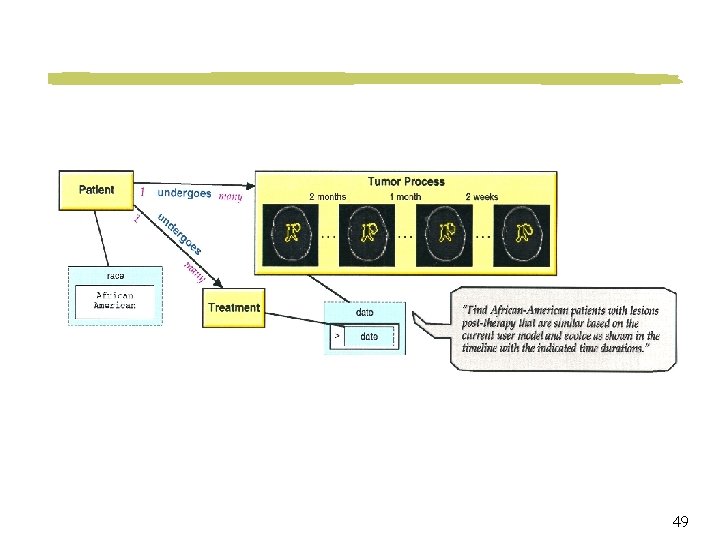
Visual Query Language and Interface Point-click-drag interface Objects may be represented iconically Spatial relationships among objects are represented graphically 40

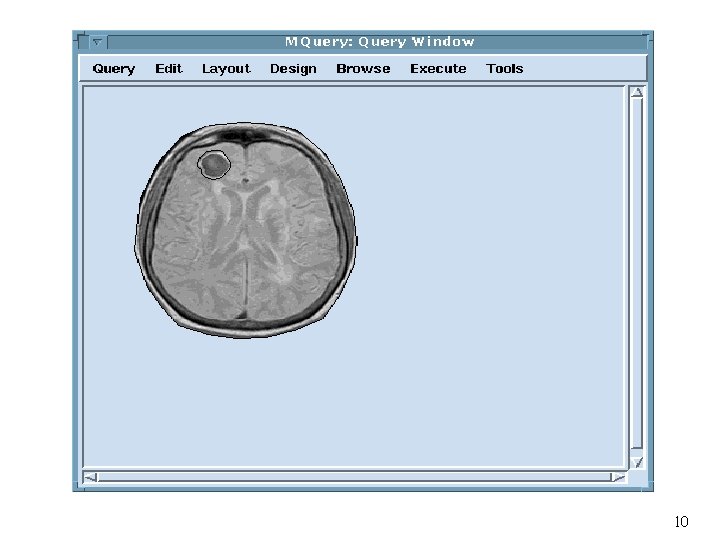
Visual Query Example Retrieve brain tumor cases where a tumor is located in the region as indicated in the picture 41
42
43
44
45
46
47
48
49
50
51

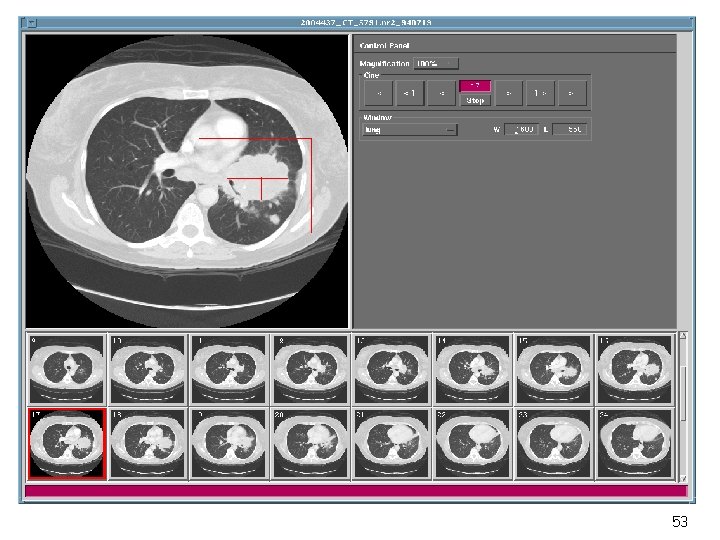
Implementation Sun Sparc 20 workstations (128 MB RAM, 24 -bit frame buffer) Oracle Database Management System X/Motif Development Environment, C++ Mass Storage of Images (9 GB) 52
53
54
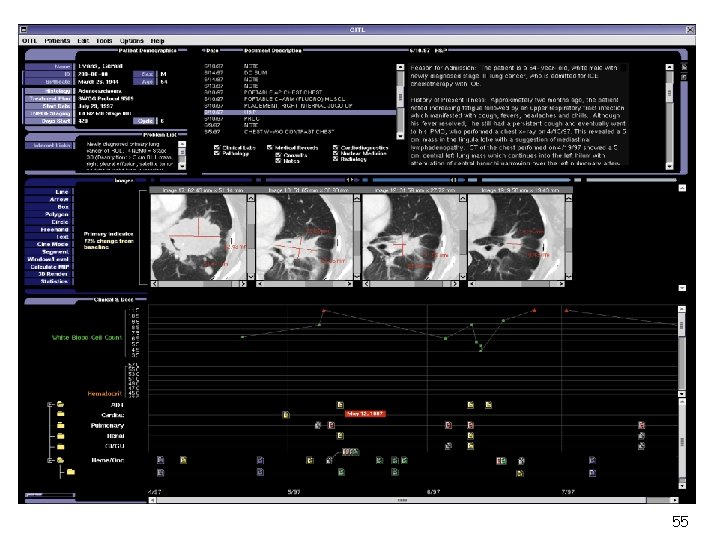
55
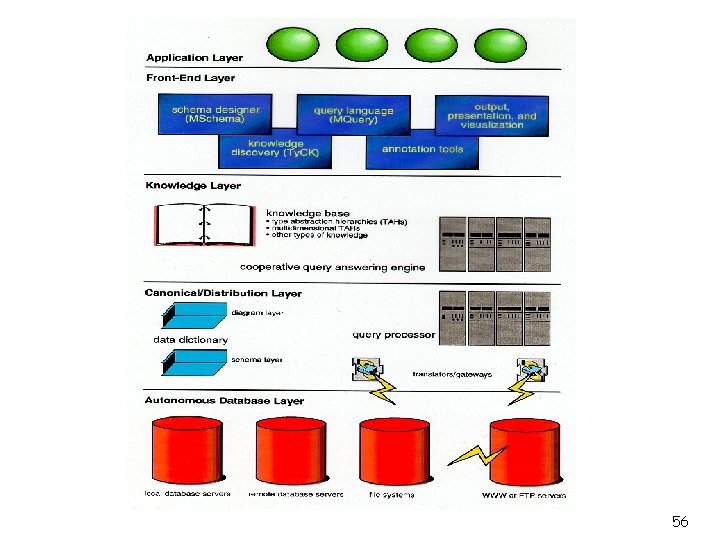
56

Conclusions Image retrieval by feature and content Matching and relaxation images based on features Processing of queries based on spatial relationships among objects Answering of imprecise queries Expression of queries via visual query language Integrated view of temporal multimedia data in a timeline metaphor 57

58
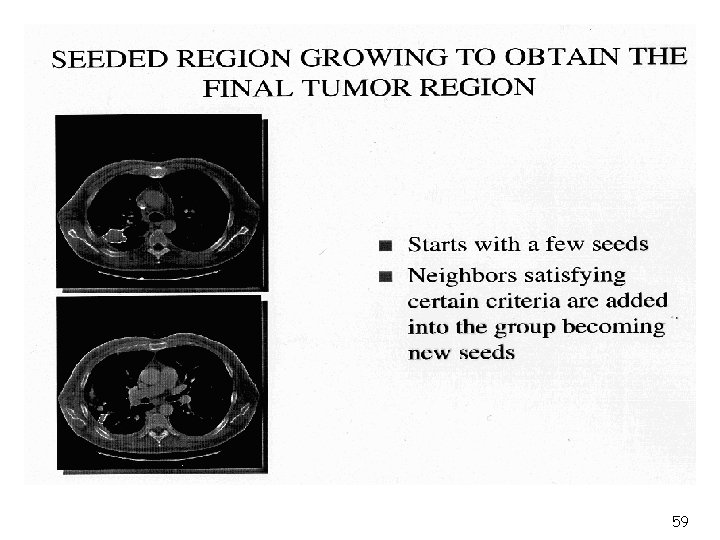
59
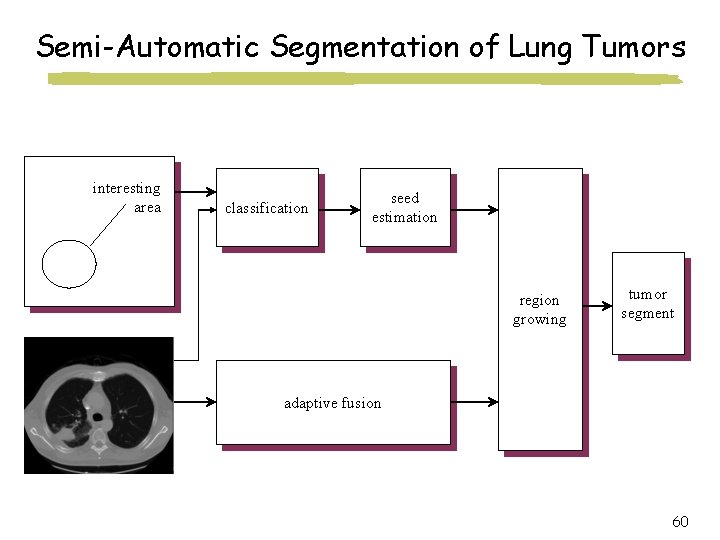
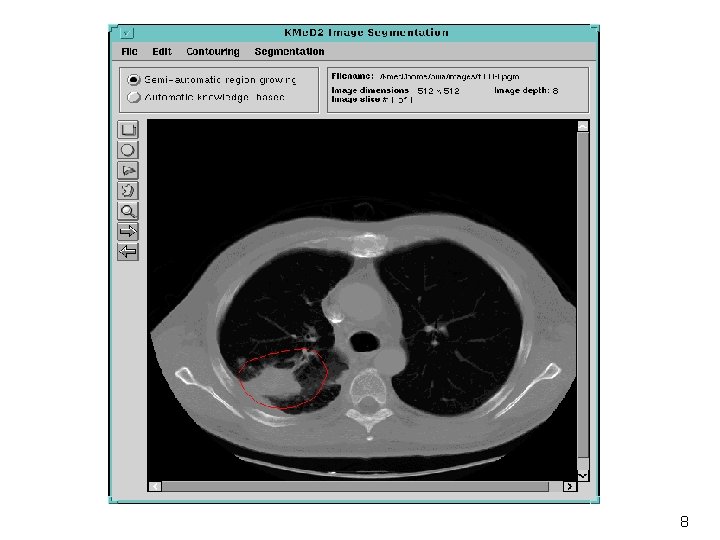
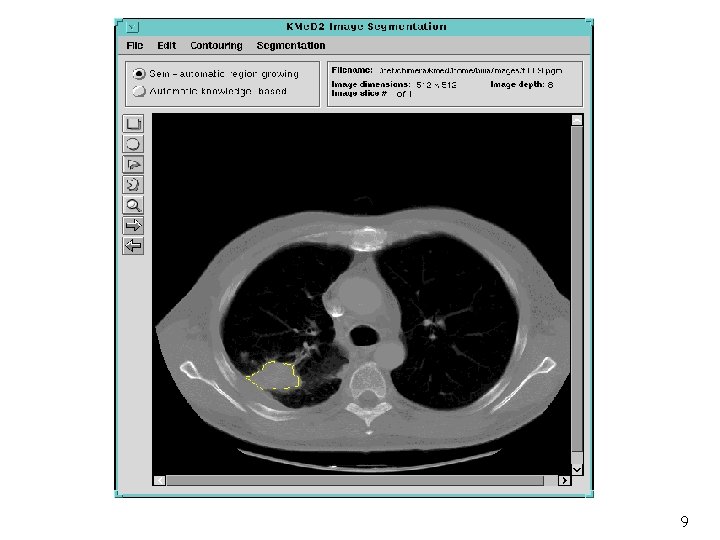
Semi-Automatic Segmentation of Lung Tumors interesting area classification seed estimation region growing tumor segment adaptive fusion 60
61

62
- Slides: 62