Kinship Analysis in Immigration Cases Charles H Brenner

Kinship Analysis in Immigration Cases Charles H. Brenner, Ph. D. consulting in forensic mathematics Oakland, California http: //dna-view. com

Outline • I. Principles of analysis – Likelihood ratio comparing hypotheses – comparing more than 2 hypotheses • II. Computer demonstration – DNA·VIEW Kinship program • III. Genetic anomalies • IV. Role of the laboratory – attain requisite LR? – answer questions, or ask them?

I. Principles of analysis • Likelihood ratio comparing hypotheses – Paternity trio is the prototype – gotta be careful about treating it as archetype
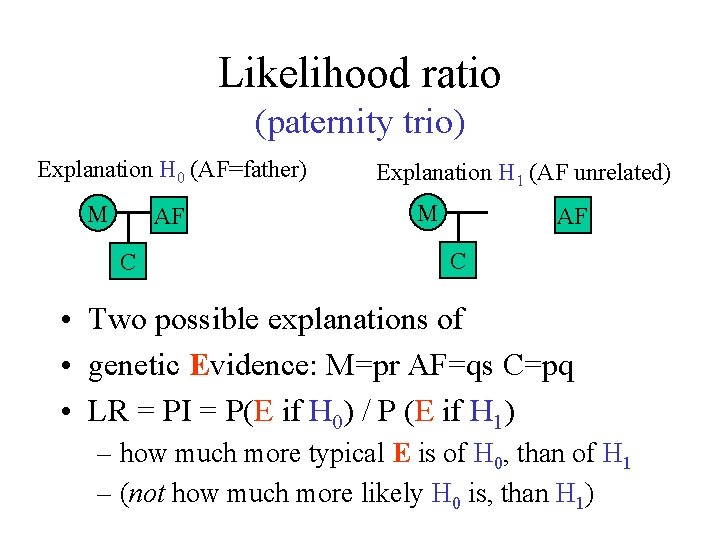
Likelihood ratio (paternity trio) Explanation H 0 (AF=father) M AF C Explanation H 1 (AF unrelated) M AF C • Two possible explanations of • genetic Evidence: M=pr AF=qs C=pq • LR = PI = P(E if H 0) / P (E if H 1) – how much more typical E is of H 0, than of H 1 – (not how much more likely H 0 is, than H 1)
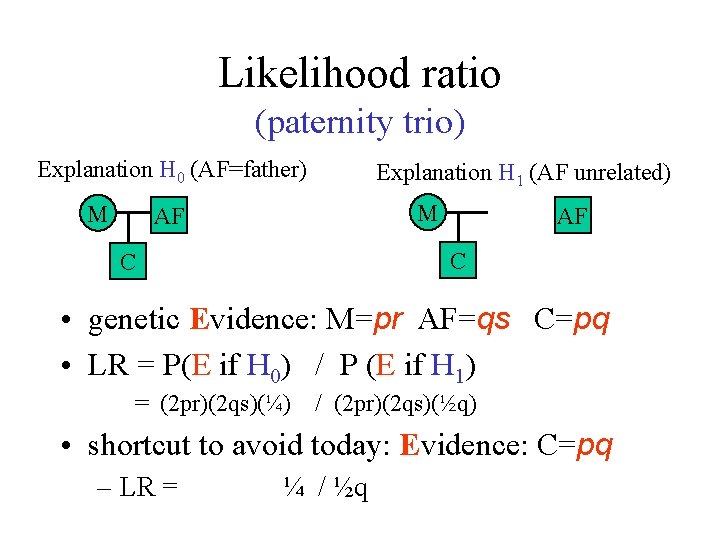
Likelihood ratio (paternity trio) Explanation H 0 (AF=father) M Explanation H 1 (AF unrelated) M AF AF C C • genetic Evidence: M=pr AF=qs C=pq • LR = P(E if H 0) / P (E if H 1) = (2 pr)(2 qs)(¼) / (2 pr)(2 qs)(½q) • shortcut to avoid today: Evidence: C=pq – LR = ¼ / ½q
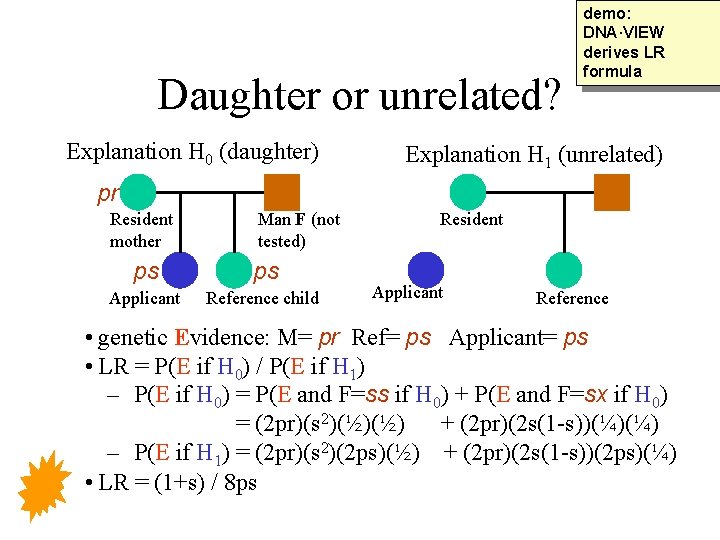
Daughter or unrelated? Explanation H 0 (daughter) demo: DNA·VIEW derives LR formula Explanation H 1 (unrelated) pr Resident mother Man F (not tested) ps ps Applicant Reference child Resident Applicant Reference • genetic Evidence: M= pr Ref= ps Applicant= ps • LR = P(E if H 0) / P(E if H 1) – P(E if H 0) = P(E and F=ss if H 0) + P(E and F=sx if H 0) = (2 pr)(s 2)(½)(½) + (2 pr)(2 s(1 -s))(¼)(¼) – P(E if H 1) = (2 pr)(s 2)(2 ps)(½) + (2 pr)(2 s(1 -s))(2 ps)(¼) • LR = (1+s) / 8 ps

DNA·VIEW computes LR KINSHIP File: W: meetingsAABB 20~1DAUGHTER. KIN Analysis mode: Co-dominant Autosomal ; AABB demo Nov 4, 2000 ; Is Applicant the daughter of Mother, or unrelated? Applicant/? : Mother + Fred Reference : Mother + Fred Applicant ps Reference ps Mother pr Likelihood ratio = 2. 71 (1+s) / 8 ps -- -Likelihood ratio: (1+s) / 8 ps Likelihood ratio = 2. 71 (using p=0. 2 s=0. 3)
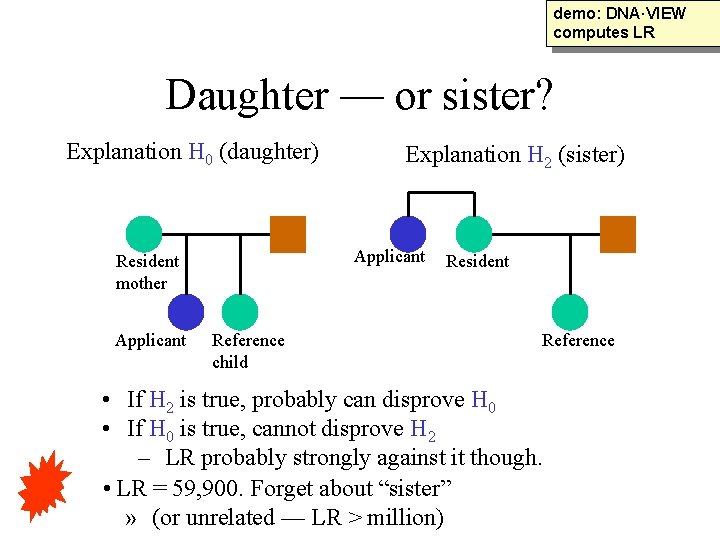
demo: DNA·VIEW computes LR Daughter — or sister? Explanation H 0 (daughter) Applicant Resident mother Applicant Explanation H 2 (sister) Reference child Resident Reference • If H 2 is true, probably can disprove H 0 • If H 0 is true, cannot disprove H 2 – LR probably strongly against it though. • LR = 59, 900. Forget about “sister” » (or unrelated — LR > million)

DNA·VIEW computation of “daughter or sister” LR Data for case 1204 2000/11/5 12: 31 M Mother 2000 -00001 c 2000/11/01 C Child #1 2000 -00003 - 2000/11/01 D Child #2 2000 -00004 - 2000/11/01 Computation per race(s): c Genotype patterns are: THO 1 TPOX CSF 1 PO M pq M p M q C pr C pq D pr D pq D 3 S 1358 M qr C pr D pq ; D=daughter or sister of M C, D/? : M + Fred M, ? /D : Granny + Gramps (Caucasian frequencies) -- D=daughter or sister of M THO 1 11 p 15. 5 PCR 2. 73 TPOX 2 p 25 -p 24 PCR 1. 31 CSF 1 PO 5 q 33 -34 PCR 3. 8 D 3 S 1358 3 p PCR 5. 89 VWA 12 p 13. 3 PCR 0. 982 FGA 4 q PCR 6. 28 D 8 S 1179 PCR 9. 81 D 21 S 11 PCR 4. 21 D 18 S 51 18 q 21. 33 PCR 0. 857 D 5 S 818 PCR 0. 869 D 13 S 317 PCR 0. 896 D 7 S 820 7 q 11 PCR 0. 764 D 16 S 539 16 q 24 PCR 5. 76 cumulative LR 59900 VWA M pq C p D pq FGA M pr C qr D pq D 8 S 1179 M qr C pr D pq (1+r) / (r+2 pr) 2 / (1+p) / (p+pq) (1+p) / (p+2 pq) (1+p+q) / (1+p+q+2 pq) (1+q) / (q+2 pq) (1+p) / (p+2 pq) (1+q) / (q+2 qr) 1 / (1+q) (1+p+q) / (1+p+q+2 pq) 1 / (1+2 p) (1+r) / (r+2 pr) D 21 S 11 M pr C qr D 18 S 51 M q C q D pq D 5 S 818 M pq C q D pq p=0. 237 r=0. 331 p=0. 528 p=0. 251 q=0. 309 p=0. 126 q=0. 256 p=0. 131 q=0. 0869 p=0. 171 q=0. 135 p=0. 0761 q=0. 221 q=0. 252 r=0. 0894 q=0. 167 p=0. 369 q=0. 35 p=0. 305 q=0. 307 p=0. 155 p=0. 107 r=0. 167 D 13 S 317 M pq C q D pq D 7 S 820 M pq C pq D pr D 16 S 539 M pq C qr D pr
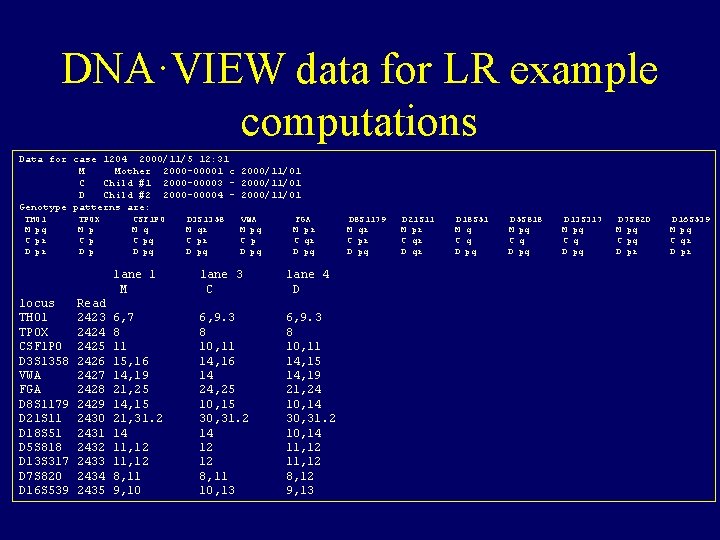
DNA·VIEW data for LR example computations Data for case 1204 2000/11/5 12: 31 M Mother 2000 -00001 c 2000/11/01 C Child #1 2000 -00003 - 2000/11/01 D Child #2 2000 -00004 - 2000/11/01 Genotype patterns are: THO 1 M pq C pr D pr locus THO 1 TPOX CSF 1 PO D 3 S 1358 VWA FGA D 8 S 1179 D 21 S 11 D 18 S 51 D 5 S 818 D 13 S 317 D 7 S 820 D 16 S 539 TPOX M p C p D p Read 2423 2424 2425 2426 2427 2428 2429 2430 2431 2432 2433 2434 2435 CSF 1 PO M q C pq D 3 S 1358 M qr C pr D pq VWA M pq C p D pq FGA M pr C qr D pq lane 1 M lane 3 C lane 4 D 6, 7 8 11 15, 16 14, 19 21, 25 14, 15 21, 31. 2 14 11, 12 8, 11 9, 10 6, 9. 3 8 10, 11 14, 16 14 24, 25 10, 15 30, 31. 2 14 12 12 8, 11 10, 13 6, 9. 3 8 10, 11 14, 15 14, 19 21, 24 10, 14 30, 31. 2 10, 14 11, 12 8, 12 9, 13 D 8 S 1179 M qr C pr D pq D 21 S 11 M pr C qr D 18 S 51 M q C q D pq D 5 S 818 M pq C q D pq D 13 S 317 M pq C q D pq D 7 S 820 M pq C pq D pr D 16 S 539 M pq C qr D pr
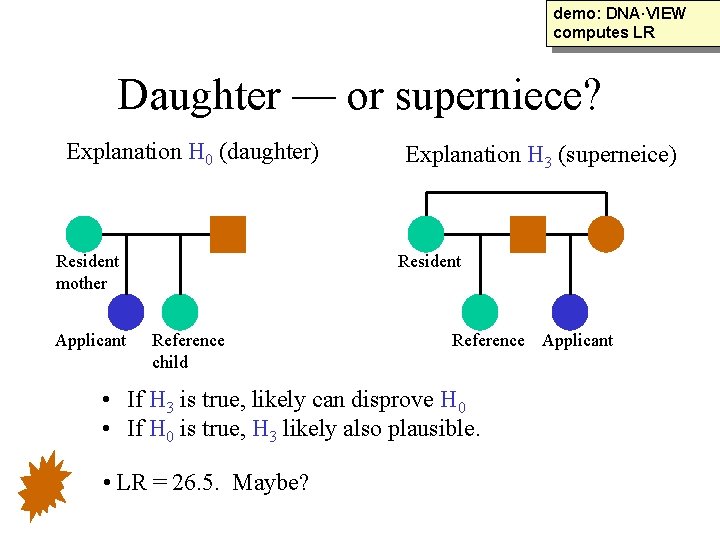
demo: DNA·VIEW computes LR Daughter — or superniece? Explanation H 0 (daughter) Resident mother Applicant Explanation H 3 (superneice) Resident Reference child Reference Applicant • If H 3 is true, likely can disprove H 0 • If H 0 is true, H 3 likely also plausible. • LR = 26. 5. Maybe?

DNA·VIEW computation of “daughter or superniece” LR ; D=daughter or superniece of M C : M + Fred D : M/Sophie + Fred M, Sophie: Granny + Gramps (Caucasian frequencies) -- D=daughter or superniece of M THO 1 11 p 15. 5 PCR 1. 26 (2+2 r) / (1+2 p+r+4 pr) TPOX 2 p 25 -p 24 PCR 1. 31 2 / (1+p) CSF 1 PO 5 q 33 -34 PCR 1. 46 (2+2 p) / (1+p+q+2 pq) D 3 S 1358 3 p PCR 1. 27 (2+2 p) / (1+p+2 q+4 pq) VWA 12 p 13. 3 PCR 1. 69 (2+2 p+2 q) / (1+p+3 q+4 pq) FGA 4 q PCR 1. 45 (2+2 q) / (1+2 p+q+4 pq) D 8 S 1179 PCR 1. 36 (2+2 p) / (1+p+2 q+4 pq) D 21 S 11 PCR 1. 65 (2+2 q) / (1+q+2 r+4 qr) D 18 S 51 18 q 21. 33 PCR 0. 857 1 / (1+q) D 5 S 818 PCR 1. 16 (2+2 p+2 q) / (1+3 p+q+4 pq) D 13 S 317 PCR 1. 24 (2+2 p+2 q) / (1+3 p+q+4 pq) D 7 S 820 7 q 11 PCR 0. 799 (2 p+2 q) / (3 p+q+4 pp+4 pq) D 16 S 539 16 q 24 PCR 1. 61 (2+2 r) / (1+2 p+r+4 pr) cumulative LR 26. 5 p=0. 237 r=0. 331 p=0. 528 p=0. 251 q=0. 309 p=0. 126 q=0. 256 p=0. 131 q=0. 0869 p=0. 171 q=0. 135 p=0. 0761 q=0. 221 q=0. 252 r=0. 0894 q=0. 167 p=0. 369 q=0. 35 p=0. 305 q=0. 307 p=0. 155 q=0. 195 p=0. 107 r=0. 167
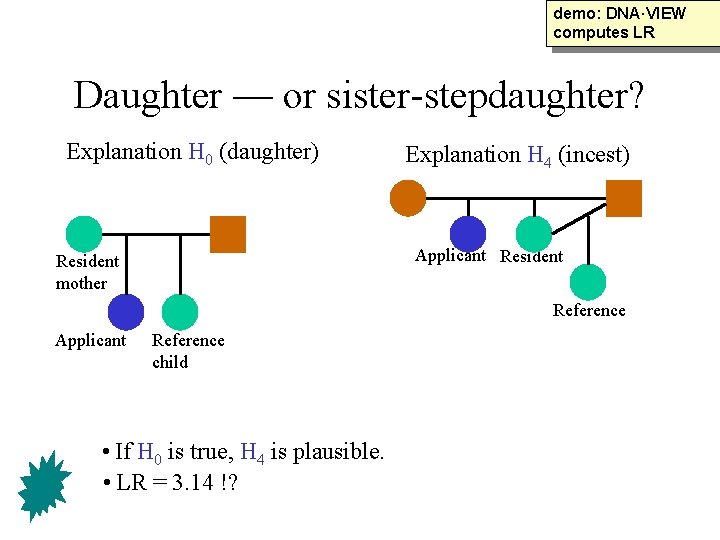
demo: DNA·VIEW computes LR Daughter — or sister-stepdaughter? Explanation H 0 (daughter) Explanation H 4 (incest) Applicant Resident mother Reference Applicant Reference child • If H 0 is true, H 4 is plausible. • LR = 3. 14 !?

DNA·VIEW computation of “daughter or incest” LR ; D=daughter or sister/stepdaughter of M? C : M + Fred M : Sophie + Fred D : M/Sophie + Fred (Caucasian frequencies) -- D=daughter or sister/stepdaughter of M? THO 1 11 p 15. 5 PCR 1. 11 2 / (1+2 p+r) TPOX 2 p 25 -p 24 PCR 1. 31 2 / (1+p) CSF 1 PO 5 q 33 -34 PCR 1. 28 2 / (1+p+q) D 3 S 1358 3 p PCR 1. 22 2 / (1+p+2 q) VWA 12 p 13. 3 PCR 1. 03 (1+5 p+q) / (1+4 p+q+ pp+5 pq) FGA 4 q PCR 1. 35 2 / (1+2 p+q) D 8 S 1179 PCR 1. 32 2 / (1+p+2 q) D 21 S 11 PCR 1. 4 2 / (1+q+2 r) D 18 S 51 18 q 21. 33 PCR 0. 75 1 / (1+2 q) D 5 S 818 PCR 0. 882 (1+p+5 q) / (1+p+4 q+5 pq+ qq) D 13 S 317 PCR 0. 918 (1+p+5 q) / (1+p+4 q+5 pq+ qq) D 7 S 820 7 q 11 PCR 0. 61 2 / (2+7 p+q) D 16 S 539 16 q 24 PCR 1. 45 2 / (1+2 p+r) cumulative LR 3. 14 p=0. 237 r=0. 331 p=0. 528 p=0. 251 q=0. 309 p=0. 126 q=0. 256 p=0. 131 q=0. 0869 p=0. 171 q=0. 135 p=0. 0761 q=0. 221 q=0. 252 r=0. 0894 q=0. 167 p=0. 369 q=0. 35 p=0. 305 q=0. 307 p=0. 155 q=0. 195 p=0. 107 r=0. 167

I. Principles of analysis • Comparing more than 2 hypotheses – – LR=10, 000 favoring “daughter” (H 0) over “unrelated” (H 1) LR=60, 000 favoring “daughter” over “sister” (H 2) LR=26 favoring “daughter” over “superniece” (H 3) LR=3. 14 favoring “daughter” over “incest” (H 4) • Hence comparing e. g. H 4 vs H 1 LR = 10, 000/3. 14 relative probability posterior probability Hypoth daughter L(E if H) 10, 000 prior 40% L·p 4, 000 L·p/sum(L·p)) 96% incest superniece sister unrelated 3, 000 400, 000 1 4% 10% 120, 000 40, 000 3% 1% 170 20% 26% 34 <1
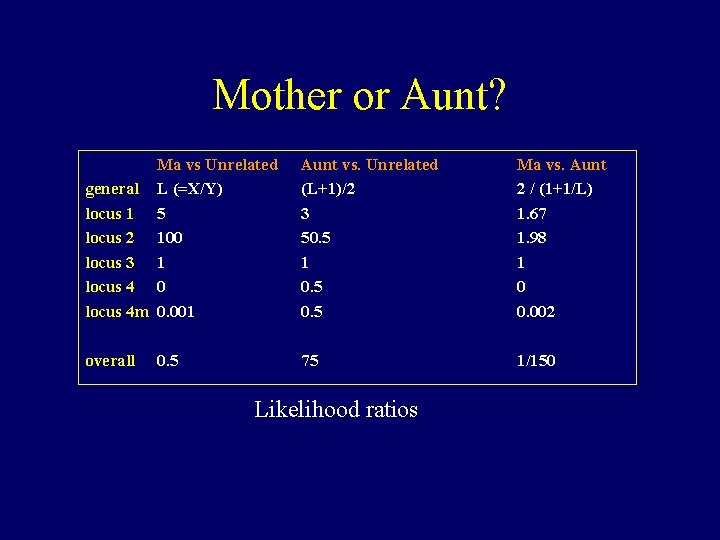
Mother or Aunt? Ma vs Unrelated general L (=X/Y) locus 1 5 locus 2 100 locus 3 1 locus 4 0 locus 4 m 0. 001 Aunt vs. Unrelated (L+1)/2 3 50. 5 1 0. 5 Ma vs. Aunt 2 / (1+1/L) 1. 67 1. 98 1 0 0. 002 overall 75 1/150 0. 5 Likelihood ratios

III. Genetic anomalies • Not co-dominant Mendelian inheritance • Incidence per meiosis (paternal) – 1/500 for VNTR (excluding p. H 30, PAC 425) – 1/400 for CODIS STR loci. • Incidence per case – 1/100 for VNTR (5 locus assay) – 1/30 for STR (13 locus assay)

STR mutation rates THO 1: 1/316 maternal

2 anomalous loci • 2 “exclusions” so to speak – RFLP-VNTR • 1 true paternity case per 25000 – STR • 1 true paternity case per 2500 • Immigration cases – may be more meioses/case

STR paternal inconsistencies • Inconsistency is like “exclusion” except maybe we don’t exclude • 1/400 inconsistent loci / true father – 1/500 one-step mutations – 1/20000 two-step mutations – 1/3000 null alleles (e. g. primer dropout)

Reasonable mutation model, STR’s Suggested model: s = /2 if s = 1 s = /20 if s = 2 etc. where =total mutation rate.

Mutation LR (new, for STR’s) (revised from http: //dna-view/gomera. htm#Mutation) • Data: Mother=PS, Child=PN, Man=RÑ – Explanation #1: Paternity; RÑ N • 50% chance transmit Ñ • = chance Ñ mutates • m = chance Ñ ends up as N, assuming mutation – m =0. 5 if N, Ñ are 1 step apart – m=0. 05 if 2 steps, etc – Explanation #2: Real father N • LR= /(4 q) (assuming single step) » q=allele frequency of the paternal allele N

IV. Role of the laboratory • attain requisite LR? – immigration rules • 99. 5%+ (but “contact lab if inconclusive”) » at 50% prior? LR>200 » at case-specific prior? (“possible fraud patterns”) LR> ? • “distinguish between family members” » need relationship-specific LR » cannot substitute larger LR threshold for wrong question • answer questions, or ask them? – advisory

LR target not a good criterion • Comparing more than 2 hypotheses – – LR=10, 000 favoring “daughter” (H 0) over “unrelated” (H 1) LR=60, 000 favoring “daughter” over “sister” (H 2) LR=26 favoring “daughter” over “superniece” (H 3) LR=3. 14 favoring “daughter” over “incest” (H 4) • Hence comparing e. g. H 4 vs H 1 LR = 10, 000/3. 14 relative probability posterior probability Hypoth daughter L(E if H) 10, 000 prior 40% L·p 4, 000 L·p/sum(L·p)) 96% incest superniece sister unrelated 3, 000 400, 000 1 4% 10% 120, 000 40, 000 3% 1% 170 20% 26% 34 <1

Summary • Principles of analysis – Likelihoods – calculate % if priors are given • Kinship program • Mutation up to 1/30 cases • Advisory role of the laboratory

• Thanks to Fairfax Identity Laboratory for the data from which STR mutation estimates were derived. • References: – Brenner CH (1997) Symbolic Kinship Program, Genetics 145: 535 -542 – Forensic mathematics — http: //dna-view. com The end
- Slides: 26