ISI Platinum Jubilee Jan 1 4 2008 Roys

ISI Platinum Jubilee, Jan 1 -4, 2008 Roy’s union-intersection principle, random fields, and brain imaging Keith Worsley, Mc. Gill Jonathan Taylor, Stanford and Université de Montréal
Sankhya (1937) 3: 65 -72

Deformation Based Morphometry (DBM) (Tomaiuolo et al. , 2004) l l l n 1 = 19 non-missile brain trauma patients, 3 -14 days in coma, n 2 = 17 age and gender matched controls Data: non-linear vector deformations needed to warp each MRI to an atlas standard l l Y 1 i(s) = deformation vector of trauma patient i at point s Y 2 i(s) = deformation vector of control patient i at point s l Locate damage: find regions where deformations are different l Test statistic: Hotelling’s T 2 (1931) l l T(s)2 = (Y 1. (s) – Y 2. (s))’Σ-1 (Y 1. (s) – Y 2. (s)) P. C. Mahalanobis with Harold Hotelling, at Gupta Nivas, 1939

Multivariate linear models for random field data £ 2 ½< Y(s)n£q is the observations variables data matrix at point s S At every point s we have a multivariate linear model: » Y(s)n£q = Xn£p B(s)n£q + E(s)n£q ; E(s)n£q N(0; I §): D. We detect sparse s where B(s) = 0 by testing H 0 : B(s) = 0 at every point s. Test statistics T (s): # regressors p=1 p>1 q=1 T F # variables q>1 Hotelling’s T 2 Need null distribution: P(maxs 2 S T (s) ¸ t) Wilks’ Λ Pillai’s trace Roy’s max root etc.

EC heuristic ¸ g t is either one At high thresholds t, the excursion set Xt = s : T (s) µ ¶ connected component (EC = 1) or empty (EC = 0), so f P max T (s) ¸ t ¼ E(EC(S s 2 S Theorem (1981, 1995) If T (s) is a smooth isotropic random ¯eld. X then E(EC(S Xt )) = Xt )): D ¹d (S)½d (t): d=0

E(EC(S D Xt )) = ¹d (S)½d (t) d=0 Intrinsic volume ¹d (S) For S with smooth boundary @S that has curvature matrix C(s), ¡ Z¡((D d)=2) = ¹d (S) 2¼ (D¡ d)=2 £ f g detr. D¡ 1¡ d C(s) ds; @S j j and ¹D (S) = S. For D = 3, ¹ 0 (S) = EC(S) ¹ 1 (S) = 2 Diameter(S) ¹ 2 (S) = 1 Surface area(S) 2 = ¹ (S) Volume(S): 3 EC density ½d (t) EC(S X Morse theory: 1 Xt ) = µ 1 f T (s)¸ ¶ tg f @T (s)=@s=0 g £ s ¡ @2 T sign µ µ + boundary ¶ @s@s 0 ¡ @2 T ¯ =E ¶ µ ¶ ¯ ½D (t) 1 f T ¸ tg det ¯ @s@s 0 ¯ @T @T =0 P =0 @s @s For a Gaussianµrandom ¶ ¯eld, ¸ = Sd( @Z ), @s ¡ p¸ @ d ¸ P(Z t) ½d (t) = 2¼ @t


Last case # regressors p=1 p>1 q=1 T✔ F✔ # variables q>1 Hotelling’s T 2✔ Wilks’ Λ Pillai’s trace Roy’s max root etc. ? We shall now ¯nd a P-value approximation (but not quite the EC density) for the maximum Roy 0 s maximum root, T (s) =maximum eigenvalue of Y(s)0 X(X 0 X)¡ 1 X 0 Y(s)(Y(s)0(I ¡ X(X 0 X)¡ 1 X 0)Y(s))¡ 1 : The above messy algebra is just too complicated. Instead. . .

Roy’s union-intersection principle Make it into a univariate linear model by multiplying by vector uq£ 1 : (Y(s)u)n£ 1 = Xn£p (B(s)u)q£ 1 + (E(s)u)n£ 1 ; H 0 u : (B(s)u)q£ 1 = 0 ° ½< q be the unit Let F (s; u) be the usual F-statistic for testing H 0 u. Let q q-sphere. Roy 0 s maximum root is T (s) = maxu 2 ° F (s; u): q ° Now F (s; u) is an F-¯eld in ¸ search region S and we already know the EC q µ ¶ density of the F-¯eld, ½d (F t) so P max T (s) s 2 S ¸ t = P X max s 2 S; u 2 ° ¼ D+q 1 2 X ¹d (S d=0 Why ½? F(s, u)=F(s, -u) = d=0 ¹d (S) t q ° Xq q D 1 2 F (s; u) ¸ k=0 )½d✔(F ¸ t) ° ¹k ( q )½d+k ✔ (F Almost the EC density of the Roy’s maximum ¸ root field t)

Non-negative least squares random field theory
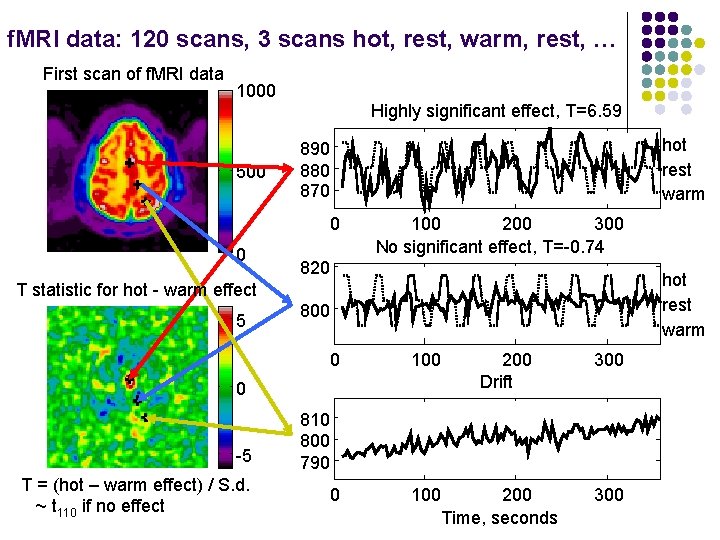
f. MRI data: 120 scans, 3 scans hot, rest, warm, rest, … First scan of f. MRI data 1000 500 Highly significant effect, T=6. 59 hot rest warm 890 880 870 0 0 100 200 300 No significant effect, T=-0. 74 820 hot rest warm T statistic for hot - warm effect 5 800 0 100 0 -5 T = (hot – warm effect) / S. d. ~ t 110 if no effect 200 Drift 300 810 800 790 200 Time, seconds 300
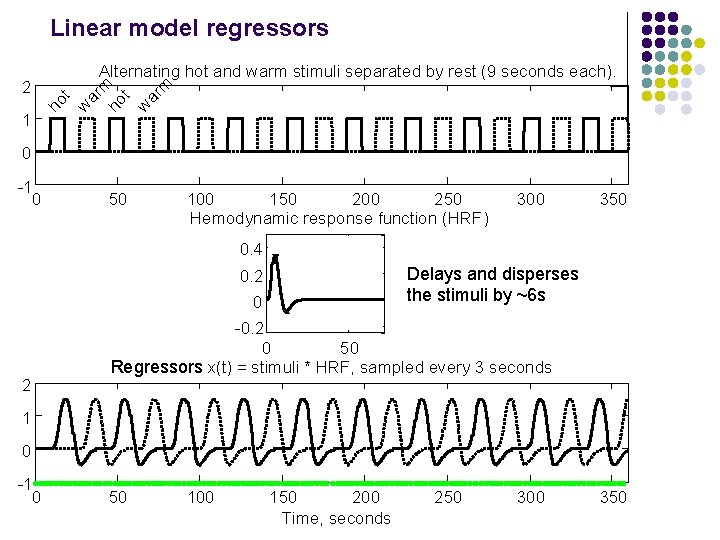
Linear model regressors 1 ho t w ar m 2 Alternating hot and warm stimuli separated by rest (9 seconds each). 0 -1 0 50 100 150 200 250 Hemodynamic response function (HRF) 300 350 0. 4 Delays and disperses the stimuli by ~6 s 0. 2 0 2 -0. 2 0 50 Regressors x(t) = stimuli * HRF, sampled every 3 seconds 1 0 -1 0 50 100 150 200 Time, seconds 250 300 350

Linear model for f. MRI time series with AR(p) errors + z(t)°? ¢ ¢+¢ ²(t) Y (t) = x(t)¯? ¡ ¡ + ap ²(t p) + ¾W N (t) ²(t) = a 1 ²(t 1) + ? ? ? t = time Y (t) = f. MRI data x(t) = stimuli convolved with HRF z(t) = drift etc: ²(t) = » error W N (t) N(0; 1) independently unknown parameters
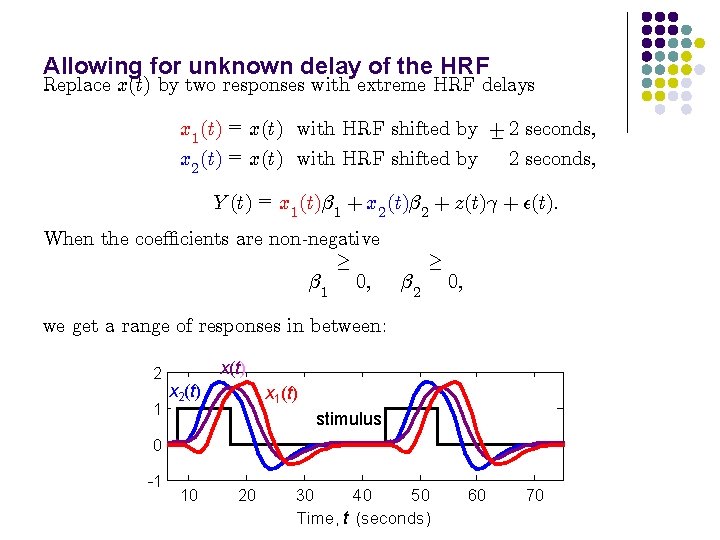
Allowing for unknown delay of the HRF Replace x(t) by two responses with extreme HRF delays x 1 (t) = x(t) with HRF shifted by + ¡ 2 seconds; x 2 (t) = x(t) with HRF shifted by Y (t) = x 1 (t)¯ 1 + x 2 (t)¯ 2 + z(t)° + ²(t): When the coe±cients are non-negative ¸ ¯ 1 0; ¯ 2 ¸ 0; we get a range of responses in between: 2 1 x(t) x 2(t) x 1(t) stimulus 0 -1 10 20 30 40 50 Time, t (seconds) 60 70
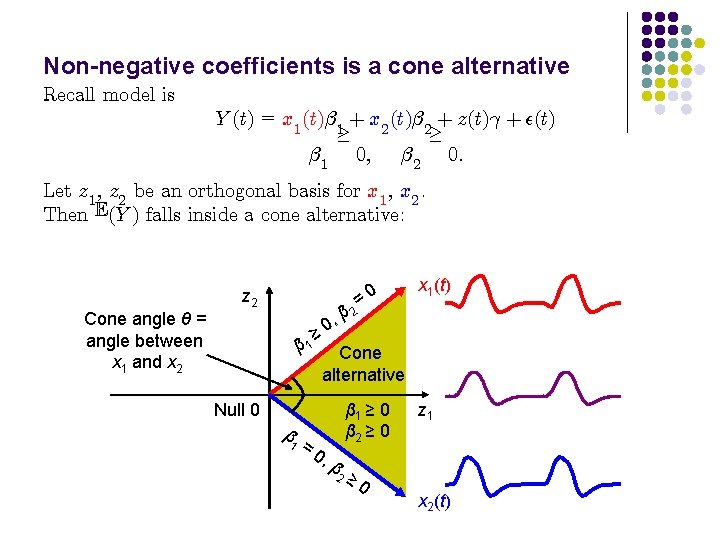
Non-negative coefficients is a cone alternative Recall model is Y (t) = x 1 (t)¯ 1 + x 2 (t)¯ 2 + z(t)° + ²(t) ¸ ¸ ¯ 1 0; ¯ 2 0: Let z 1 , z 2 be an orthogonal basis for x 1 , x 2. Then E(Y ) falls inside a cone alternative: Cone angle θ = angle between x 1 and x 2 z 2 = 2 β , 0 x 1(t) ≥ 0 β 1 Cone alternative Null 0 β 1 =0 , β 2 β 1 ≥ 0 β 2 ≥ 0 z 1 x 2(t)
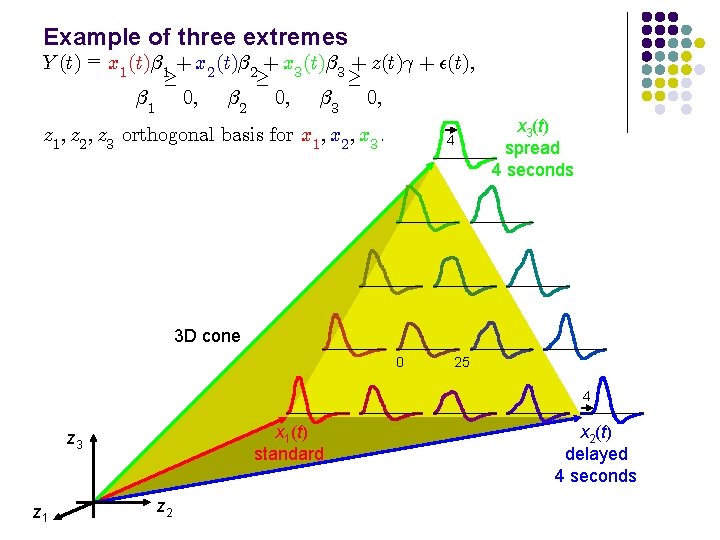
Example of three extremes Y (t) = x 1 (t)¯ 1 + x 2 (t)¯ 2 + x 3 (t)¯ 3 + z(t)° + ²(t); ¸ ¸ ¸ ¯ 1 0; ¯ 2 0; ¯ 3 0; z 1 ; z 2 ; z 3 orthogonal basis for x 1 ; x 2 ; x 3. 4 x 3(t) spread 4 seconds 3 D cone 0 25 4 x 1(t) standard z 3 z 1 z 2 x 2(t) delayed 4 seconds

General non-negative least squares problem In general we may have p constrained regressors and q unconstrained regressors. The constrained regressors could be the extremes of the HRF, e. g. min/max latency shift, min/max spread, etc. The model is then all HRFs in between. In vector form: Yn£ 1 ¯p£ 1 °q£ 1 ²n£ 1 =X + ²n£ 1 ° ¸ n£p ¯p£ 1 + Z¡ n£q q£ 1 0 (component wise) » unconstrained N(0 n£ 1 ; In£n ¾ 2 ) (without loss of generality) We might also have far more regressors than actual observations(!), i. e. p >> n. Footnote: Woolrich et al. (2004) replace “hard” constraints by “soft” constraints through a prior distribution on β, taking a Bayesian approach.


Pick a range of say p = 150 plausible values of the non-linear parameter ν: ν 1, …, νp

Fitting the NNLS model Simple: • Do “all subsets” regression • Throw out any model that does not satisfy the non-negativity constraints • Among those left, pick the model with smallest error sum of squares For larger models there are more efficient methods e. g. Lawson & Hanson (1974). The non-negativity constraints tend to enforce sparsity, even if regressors are highly correlated (e. g. PET). Why? Highly correlated regressors have huge positive and negative unconstrained coefficients – non-negativity suppresses the negative ones.
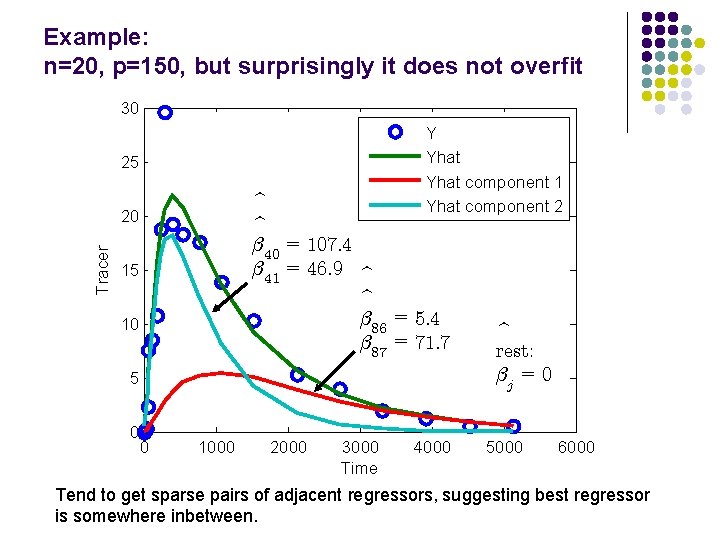
Example: n=20, p=150, but surprisingly it does not overfit 30 Y Yhat component 1 Yhat component 2 25 b b ¯ 40 = 107: 4 ¯ 41 = 46: 9 b b ¯ 86 = 5: 4 ¯ 87 = 71: 7 Tracer 20 15 10 5 0 0 1000 2000 3000 Time 4000 b rest: ¯j = 0 5000 6000 Tend to get sparse pairs of adjacent regressors, suggesting best regressor is somewhere inbetween.

! P-values for testing the cone alternative? H 0 : ¯p£ 1 = ! error sum of squares = SSE 0 ¸ 0 p£ 1 ¡ H 1 : ¯p£ 1 0 p£ 1 (component wise) error sum of squares = SSE 1 Likelihood ratio test statistic is the Beta-bar statistic, equal to the coe±cient of determination or multiple correlation R 2 : ¡ ¹ = SSE 0 SSE 1 B SSE 0 ¡ ¡ ¢ ¢ Null distribution if there are no constraints, i. e. H 1 : ¯p£ 1 = 0 p£ 1 : ¸ ¸ ¡ ¹ t) = P Beta p ; º¡ p P(B t ; º=n q 2 2 Null distribution with constraints is a weighted average of Beta distributions, hence the name Beta-bar (Lin. X & Lindsay, Takemura 1997): ¡ ¡ ¢ & Kuriki, ¢ ¹ P(B ¸ t) = º j =1 wj P Beta j ; º¡ j 2 2 b ¸ t =1 when j=ν f g wj = P(# unconstrained ¯ 0 s > 0 = j)

P-values¹ for = PET data at a is t 0: 9524, º = 20. Observed B single voxel p = 2: Cone weights: w 1 = ½, w 2 = θ/2π ¹ weights w Easiest way to ¯nd the B j is by simulation (10000 iterations): ¡ P Beta wj ¹ P(B ¸ X j t) = º j =1 ¡ wj P Beta ¡ j ; º¡ j 2 2 ¢ ¸ ¢ ¡ j ; º¡ j 2 2 t = 2: 35 £ ¢ ¸ j 10¡ 12 t ¢

The Beta-bar random field Recall that at a single point (voxel): X ¹ P(B ¸ t) = º ¡ wj P Beta ¡ j ; º¡ j 2 2 ¢ ¸ ¢ t j =1 2 ½< D , is an isotropic random ¯eld: that if T (s); s ¶ S Recall µ X P max T (s) ¸ t ¼ ¸ g f E(EC(S s : T (s) t )) = s 2 S with ½ 0 ´ D ¹d (S)½d (T ¸ t) d=0 P. Taylor & Worsley X (2007): ¡ µ ¹ ½d (B ¸ t) = wj ½d Beta ¶ X µ = j 1 From simulations at a single¸ voxel ¼ ¹ P max B(s) s 2 S º ¡ j ; º¡ j 2 2 ¡ ¢ ¸ ºSame linear combination! wj P max Beta t j =1 s 2 S ¢ t ¢ j ; º¡ j 2 2 =1 when j=ν ¶ From well-known EC ¸ densities of F field (s) t
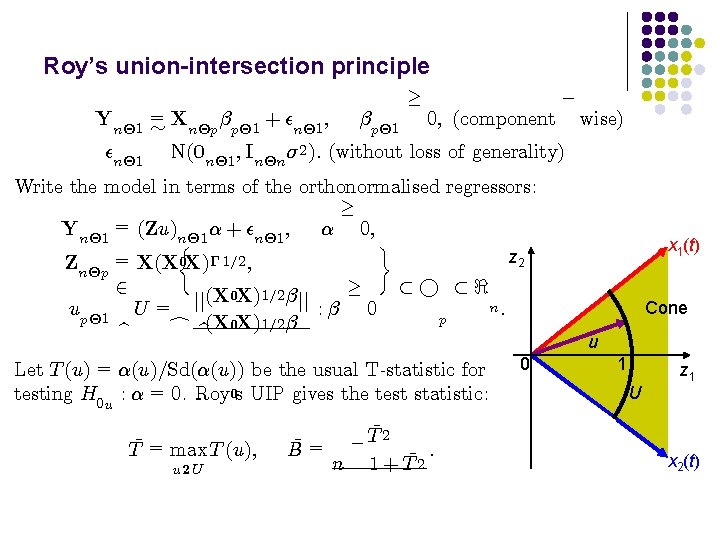
Roy’s union-intersection principle ¡ ¸ Yn£ 1 = » Xn£p ¯p£ 1 + ²n£ 1 ; ¯p£ 1 0; (component wise) ²n£ 1 N(0 n£ 1 ; In£n ¾ 2 ): (without loss of generality) Write the model in terms of the orthonormalised regressors: ¸ Yn£ 1 = (Zu)n£ 1 ® + ²n£ 1 ; ® 0; ½ ¾ z 2 Zn£p = X(X 0 X)¡ 1=2 ; ¸ 2 ½° ½< (X 0 X)1=2 ¯ jj jj n: up£ 1 U = : ¯ 0 p c b(X 0 X)1=2 ¯ b Let T (u) = ®(u)=Sd(®(u)) be the usual T-statistic for testing H 0 u : ® = 0. Roy 0 s UIP gives the test statistic: T¹ = max T (u); u 2 U ¹= B ¹ ¡T 2 : ¹ n 1 + T 2 0 x 1(t) Cone u 1 U z 1 x 2(t)

Roy’s union-intersection principle Can we use the same trick of adding U to the search space? Let 0 s put back the parameter s. The T-bar statistic is » µ ¶ ¹ = T (s) max T (s; u); T (s; u) H tn¡ 1 ; u 2 U P max T¹(s) s 2 S 0 ¸ t = PX max T (s; u) ¸ t s 2 S; u 2 U ¼ D+q X ¹d (S d=0 = U )½✔(T X d ¸ t) q D d=0 ¹d (S) ¹k (U )½d+k ✔ (T k=0 ¸ t): b Sadly this doesn 0 t work: although c. T b(s; u) is a T-¯eld in s for each u, it is not a T-¯eld in (s; u), despite the ¡ fact that ®(s; u) is a Gaussian random ¯eld in (s; u). The reason is that (n 1)Sd(®(s; u))2 is not a 2 ¯eld in (s; u). n¡ 1
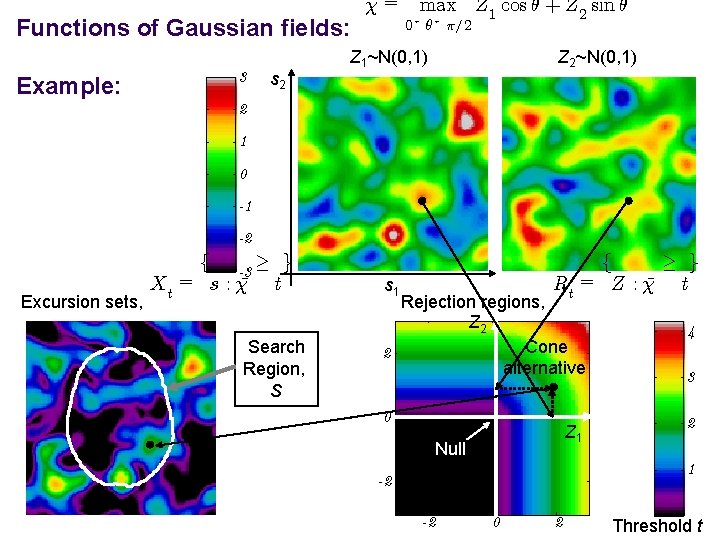
Functions of Gaussian fields: Example: 3 s 2 ¹= max Z 1 cos µ + Z 2 sin µ 0· µ· ¼=2 Z 1~N(0, 1) Z 2~N(0, 1) 2 1 0 -1 -2 Excursion sets, f -3 ¸ g Xt = s : ¹ t Search Region, S ¸ g f Rt = Z : ¹ t s 1 Rejection regions, Z 2 Cone 2 alternative 4 0 2 Z 1 Null 3 1 -2 -2 0 2 Threshold t
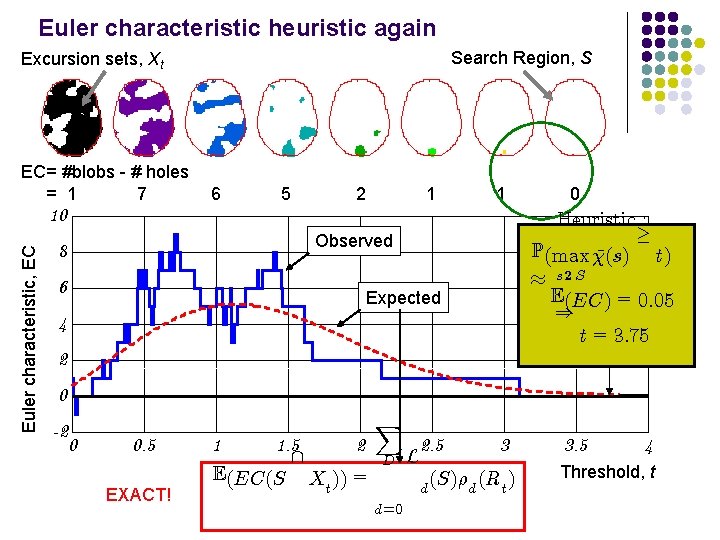
Euler characteristic heuristic again Search Region, S Excursion sets, Xt EC= #blobs - # holes = 1 7 6 5 2 1 1 Euler characteristic, EC 10 Heuristic : ¸ P(max (s) ¹ t) ¼ s 2 S E(EC) = 0: 05 ) t = 3: 75 Observed 8 6 0 Expected 4 2 0 -2 0 0. 5 EXACT! 1 1. 5 E(EC(S 2 Xt )) = X D d=0 L 2. 5 3 (S)½d (Rt ) d 3. 5 4 Threshold, t

E(EC(S Beautiful symmetry: Lipschitz-Killing curvature L d D Xt )) = L d (S)½d (Rt ) ¸ = Sd d=0 (S) Steiner-Weyl Tube Formula (1930) @Z @s EC density ½d (Rt ) Taylor Gaussian Tube Formula (2003) • Put a tube of radius r about the search region λS and rejection region Rt: Z 2~N(0, 1) r Rt Tube(λS, r) Tube(Rt, r) r λS t-r t X X d= 0 d=0 Z 1~N(0, 1) • Find volume or probability, expand as a power series in r, pull off coefficients: 1 (2¼)d=2 j j L D ¼d ½d (Rt )rd d P(Tube(R ; r)) = (S)r Tube(¸S; r) = t d D¡ d! ¡(d=2 + 1)

Lipschitz-Killing L curvature d (S) of a triangle r Tube(λS, r) λS ¸ = Sd µ @Z @s ¶ X Steiner-Weyl Volume of Tubes Formula (1930) L ¼ d=2 Area(Tube(¸S; r)) = (S)rd d D¡ ¡(d=2 + 1) L L L d=0 = (S) + 2 (S)r + ¼ (S)r 2 2 1 0 = Area(¸S) + Perimeter(¸S)r + EC(¸S)¼r 2 L (S) = EC(¸S) L 0 =1 L 1 (S) 2 Perimeter(¸S) (S) = Area(¸S) 2 D Lipschitz-Killing curvatures are just “intrinisic volumes” or “Minkowski functionals” in the (Riemannian) metric of the variance of the derivative of the process
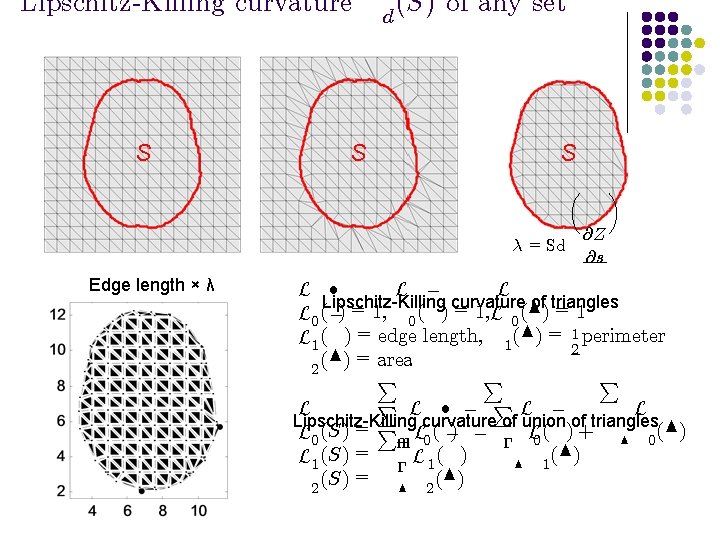
Lipschitz-Killing curvature S S d (S) of any set S ¸ = Sd Edge length × λ µ @Z @s ¶ L ² L ¡ L Lipschitz-Killing curvature of triangles L 0 (¡) = 1, 0 ( ) = 1, L 0 (N) = 1 L 1 ( ) = edge length, 1 (N) = 12 perimeter (N) = area 2 P P L ² ¡ PL ¡ L L Lipschitz-Killing curvature of union of triangles L 0 (S) = P² L 0 ( ¡) ¡ ¡ L 0 ( ) + N 0 (N) L 1 (S) = ¡ L 1 ( ) N 1 (S) = N 2 (N) 2
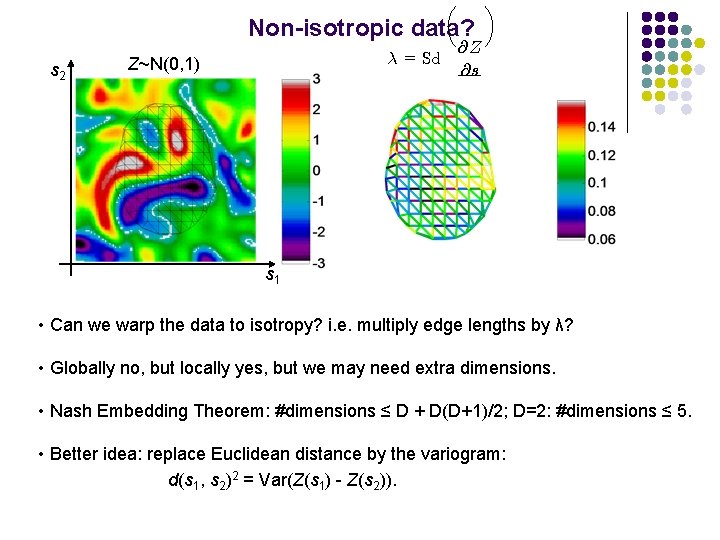
µ Non-isotropic data? s 2 ¸ = Sd Z~N(0, 1) @Z @s ¶ s 1 • Can we warp the data to isotropy? i. e. multiply edge lengths by λ? • Globally no, but locally yes, but we may need extra dimensions. • Nash Embedding Theorem: #dimensions ≤ D + D(D+1)/2; D=2: #dimensions ≤ 5. • Better idea: replace Euclidean distance by the variogram: d(s 1, s 2)2 = Var(Z(s 1) - Z(s 2)).

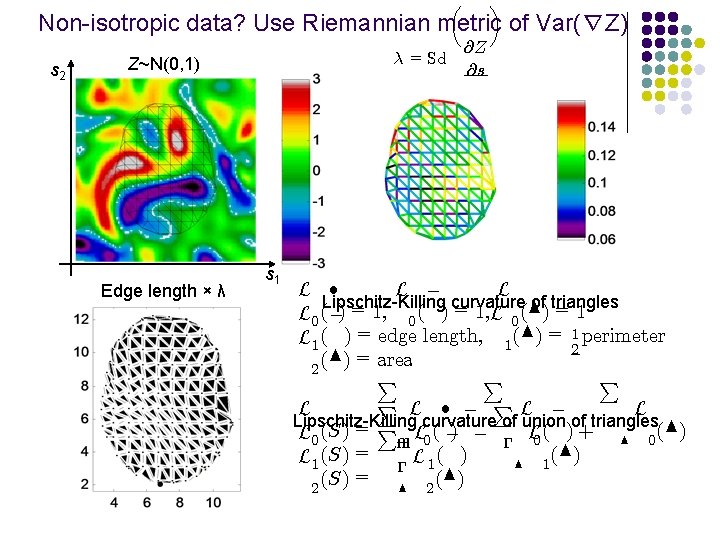
µ ¶ Non-isotropic data? Use Riemannian metric of Var(∇Z) s 2 ¸ = Sd Z~N(0, 1) Edge length × λ s 1 @Z @s L ² L ¡ L Lipschitz-Killing curvature of triangles L 0 (¡) = 1, 0 ( ) = 1, L 0 (N) = 1 L 1 ( ) = edge length, 1 (N) = 12 perimeter (N) = area 2 P P L ² ¡ PL ¡ L L Lipschitz-Killing curvature of union of triangles L 0 (S) = P² L 0 ( ¡) ¡ ¡ L 0 ( ) + N 0 (N) L 1 (S) = ¡ L 1 ( ) N 1 (S) = N 2 (N) 2
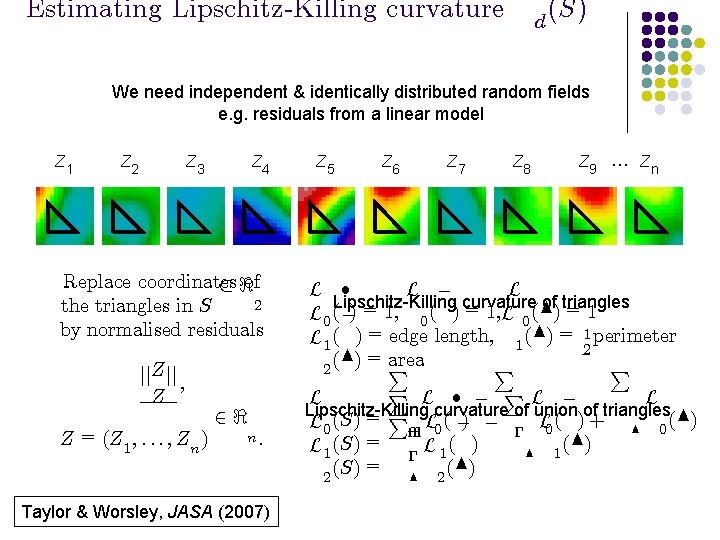
Estimating Lipschitz-Killing curvature d (S) We need independent & identically distributed random fields e. g. residuals from a linear model Z 1 Z 2 Z 3 Z 4 Replace coordinates ½ <of 2 the triangles in S by normalised residuals jj. Z jj ; Z Z = (Z 1 ; : : : ; Zn ) 2< n: Taylor & Worsley, JASA (2007) Z 5 Z 6 Z 7 Z 8 Z 9 … Zn L ² L ¡ L Lipschitz-Killing curvature of triangles L 0 (¡) = 1, 0 ( ) = 1, L 0 (N) = 1 L 1 ( ) = edge length, 1 (N) = 12 perimeter area (N) = P P P 2 P L ² ¡ PL ¡ L L Lipschitz-Killing curvature of union of triangles L 0 (S) = P² L 0 ( ¡) ¡ ¡ L 0 ( ) + N 0 (N) L 1 (S) = ¡ L 1 ( ) N 1 (S) = N 2 (N) 2

E(EC(S Beautiful symmetry: Lipschitz-Killing curvature L d D Xt )) = L d (S)½d (Rt ) ¸ = Sd d=0 (S) Steiner-Weyl Tube Formula (1930) @Z @s EC density ½d (Rt ) Taylor Gaussian Tube Formula (2003) • Put a tube of radius r about the search region λS and rejection region Rt: Z 2~N(0, 1) r Rt Tube(λS, r) Tube(Rt, r) r λS t-r t X X d= 0 d=0 Z 1~N(0, 1) • Find volume or probability, expand as a power series in r, pull off coefficients: 1 (2¼)d=2 j j L D ¼d ½d (Rt )rd d P(Tube(R ; r)) = (S)r Tube(¸S; r) = t d D¡ d! ¡(d=2 + 1)

EC density ½d ( ¹ of the ¹ statistic t) Z 2~N(0, 1) Tube(Rt, r) r t-r Rejection region Rt Z 1~N(0, 1) t X Taylor’s Gaussian Tube Formula (2003) ¸ d=2 (2¼) P (Z ; Z ¹ t)r d Tube(Rt ; r) = ½d ( 1 2 d! d=0 Z ¢¢¢ ¸ ¸ ¸ = ½ ( ¹ t) + (2¼)1=2 ½ 1 ( ¹ t)r + (2¼)½ 2 ( ¹ t)r 2 =2 + 0 2 1 1 = t¡ r ½ 0 ( ¹ ½ 1 ( ¹ ½ 2 ( ¹ ¸ ¸ ¸ (2¼)Z¡ 1=2 e¡ z 2 =2 dz + e¡ (t¡ r)2 =2 =4 1 t) = (2¼)¡ 1=2 e¡ z 2 =2 dz + e¡ t 2 =2 =4 t t) = (2¼)¡ 1 e¡ t 2 =2 + (2¼)¡ 1=2 e¡ t 2 =2 t=4 t) = (2¼)¡ 3=2 e¡ t 2 =2 t + (2¼)¡ 1 e¡ t 2 =2 (t 2. . . ¡ 1)=8

¹ EC density ½d (B ¹ statistic t) of the B Recall that at a single point (voxel): X ¹ P(B ¸ t) = n ¡ wj P Beta ¡ j ; º¡ j 2 2 ¢ ¸ ¢ t j =1 2 ½< D , is an isotropic random ¯eld: Recallµthat if F (s); s ¶ S X P max Z(s) ¸ t ¼ E(EC(S ¸ g s : Z(s) t )) = f s 2 S with ½ 0 ´ L D d (S)½d (F ¸ t) d=0 P. Taylor & Worsley X (2007): ¡ µ ¹ ½d (B ¸ n t) = wj ½d Beta ¶ X µ = j 1 ¡ j ; º¡ j 2 2 ¡ ¢ ¸ From simulations linear combination! n at a single¸ voxel ¼ Same ¹ P max B(s) s 2 S wj P max Beta t j =1 s 2 S ¢ t ¢ j ; º¡ j 2 2 ¶ From well-known EC ¸ densities of F field (s) t
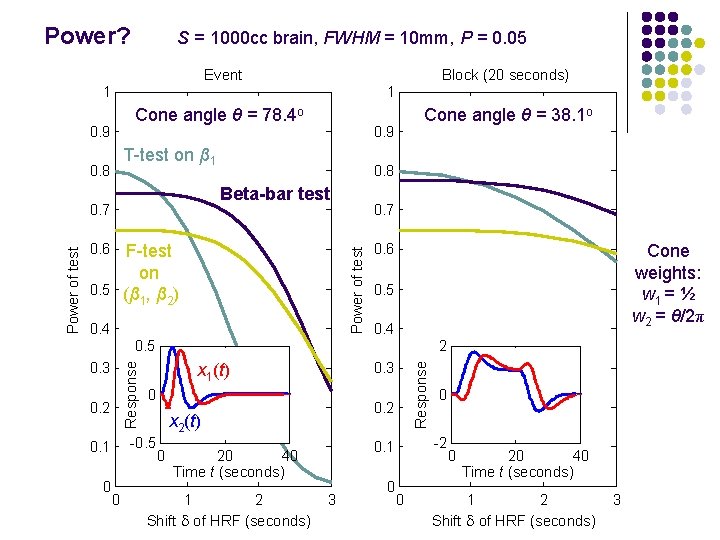
Power? S = 1000 cc brain, FWHM = 10 mm, P = 0. 05 Event Block (20 seconds) 1 1 Cone angle θ = 78. 4 o 0. 9 0. 8 Beta-bar test 0. 7 0. 6 0. 7 Power of test F-test on 0. 5 (β , β ) 1 2 0. 4 0. 6 Cone weights: w 1 = ½ w 2 = θ/2π 0. 5 0. 4 0. 5 0. 2 0 0 0. 3 x 1(t) 0 0. 2 x 2(t) -0. 5 0. 1 2 0 2 Shift d of HRF (seconds) 3 0 0 -2 0. 1 20 40 Time t (seconds) 1 Response 0. 3 Response Power of test 0. 9 T-test on β 1 0. 8 Cone angle θ = 38. 1 o 0 0 20 40 Time t (seconds) 1 2 Shift d of HRF (seconds) 3

Cross correlation random field 2 ½< £ M , and Y (t), t N be n Let X(s), s S T 1 vectors of Gaussian random ¯elds. De¯ne the cross correlation random ¯eld as P µ C(s; t) = max ¶ u; v ¸ ¼ : ¸ g 2 f 2 E(EC s S; t T : C(s; t) c ) ¸ L (S) j (T )½ij (C c) i dim(S) dim(T ) L i=0 ½ij (C (X(s)0 X(s) Y (t)0 Y (t))1=2 max C(s; t) c X X s 2 S; t 2 T = ¸ X(s)0 Y (t) ¡ j =0 X ¡ 2 n¡ 2¡ h (i 1)!j! b(h¡ 1)=2 c ¡ c) = ( 1)k ch¡ 1¡ 2 k (1 c 2 )(n¡ 1¡ h)=2+k ¼ h=2+1 k=0 XX k k l=0 m=0 n¡ j ¡ ¡ ¡( n¡ 2 i + l)¡( 2 ¡+ m) ¡ ¡ : l!m!(k l m)!(n 1 h + l + m + k)!(i 1 k l + m)!(j k m + l)!

Maximum canonical cross correlation random field 2 ½< M , and Y(t) N be matrices of Gaussian , t T Let X(s)n£p , s S n£q random ¯elds. De¯ne the maximum canonical cross correlation random ¯eld as C(s; t) = max u; v u 0 X(s)0 Y(t)v (u 0 X(s)u v 0 Y(t)v)1=2 ; the maximum of the canonical correlations between X and Y, de¯ned as the µ singular values of (X 0¶ X)¡ 1=2 X X 0 Y(Y 0 Y) X¡ 1=2. X X P max C(s; t) s 2 S; t 2 T where ¸ c ¼ M 1 2 L N i (S) i=0 L ° ( p) = k L p j (T ) j =0 ³ 2 k+1 ¼ k 2 ¡ k! L ° ( p) k ¡ k=0 ¢ ´ p+1 2 p¡ 1¡ k ! 2 ¡ ¡ ¡ if p 1 k is even, and zero otherwise, k = 0; : : : ; p 1. q l=0 L ° ¸ ( q )½i+k; j+l (C c) l
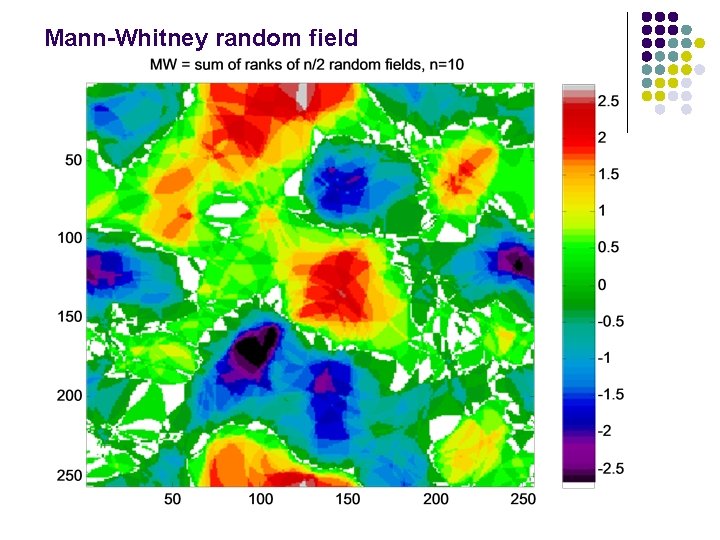
Mann-Whitney random field
- Slides: 43