IPAM 3 Childhood Sarcoma Classification by Gene Expression

IPAM #3: Childhood Sarcoma Classification by Gene Expression Profiles Timothy J. Triche CHLA/USC

Clinical Classification of Childhood Cancer • Historical: Morphologic diagnosis + clinical data => risk group, protocol eligibility, treatment (eg, group-based treatment) • Current: Combined (morphology, immunophenotype, genomic defect) => patient-specific group-based treatment • Future: Patient-specific therapy, based on multi-genic phenotype?

Osteosarcoma • Five histologic types, no prognostic value • Weak prognostic features: site, size, age • No specific, predictive genetic abnormality (RB, p 53) • Clinical stage only significant prognostic indicator at presentation

Osteosarcoma Prognosis • Pre-resection chemotherapy => major increase in survival • Improved survival limited to patients with ≥ 95% tumor kill • Patients w/ metastases can be salvaged But, many exceptions occur: – Responders who metastasize & die – Non-responders who survive – Metastatic patients who survive after resection of mets Thus, predicting outcome & tailoring therapy remains a major problem
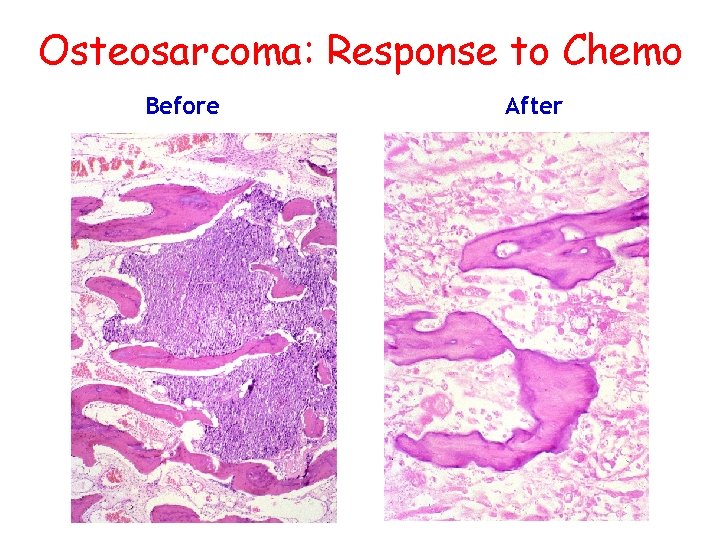
Osteosarcoma: Response to Chemo Before After

Osteosarcoma Survival • Surgery only: <10% • Metastases, no surgery: 0% • Metastases, surgery: ~20% • Single-agent chemotherapy: <20% • Conventional chemotherapy: ~44% • Up-front chemotherapy: ~65% • Responders: ~80% • Non-responders: <40%

Multi-gene Analysis by Microarrays • Single gene abnormalities, even when present, are inadequate alone to: – – – Establish a diagnosis Identify individual patients risk profile Predict clinical course Predict response to therapy Predict outcome • Increasing evidence suggests gene expression profiles may favorably address these issues

Gene Expression Analyses • Scatter Analyses – 1 X 1 – Groups • Outlier Gene Analyses – Up & down regulated from mean – Identity • Cluster Analyses – All genes – Various methods
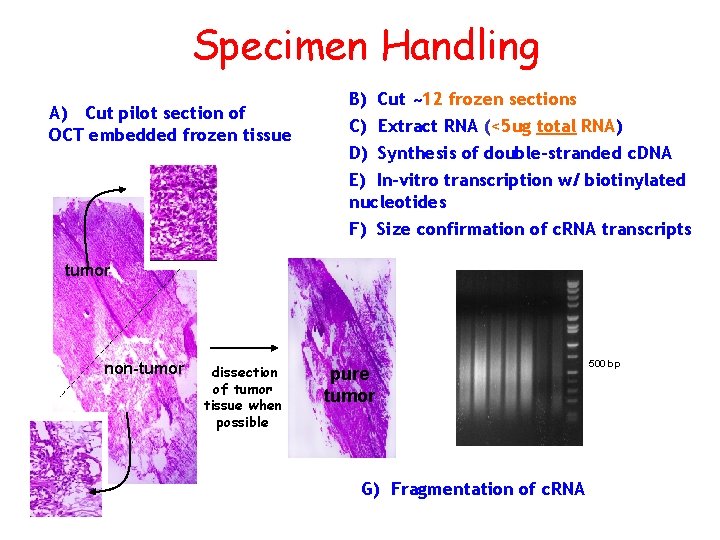
Specimen Handling A) Cut pilot section of OCT embedded frozen tissue B) Cut ~12 frozen sections C) Extract RNA (<5 ug total RNA) D) Synthesis of double-stranded c. DNA E) In-vitro transcription w/ biotinylated nucleotides F) Size confirmation of c. RNA transcripts tumor non-tumor dissection of tumor tissue when possible pure tumor G) Fragmentation of c. RNA 500 bp
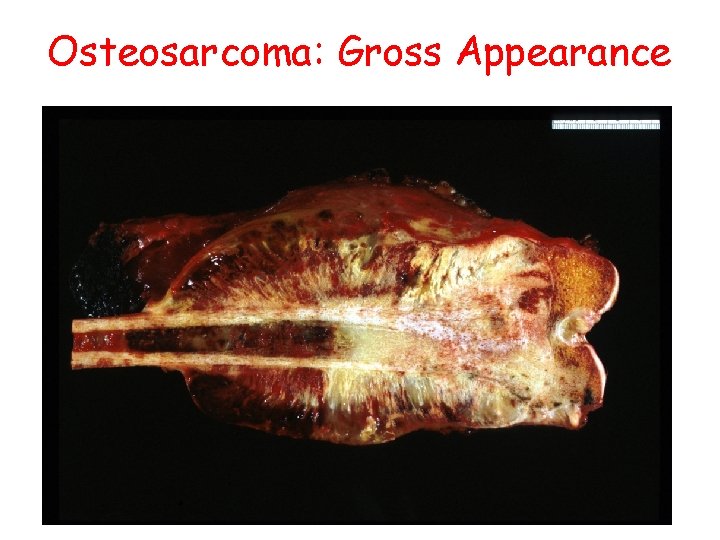
Osteosarcoma: Gross Appearance
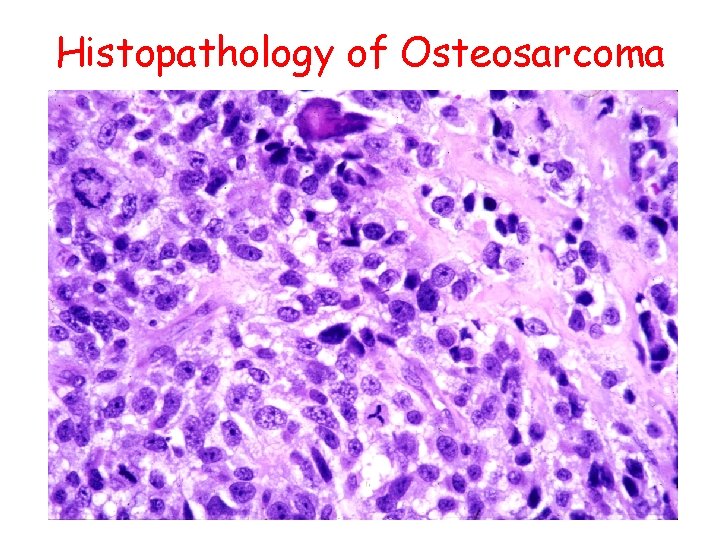
Histopathology of Osteosarcoma
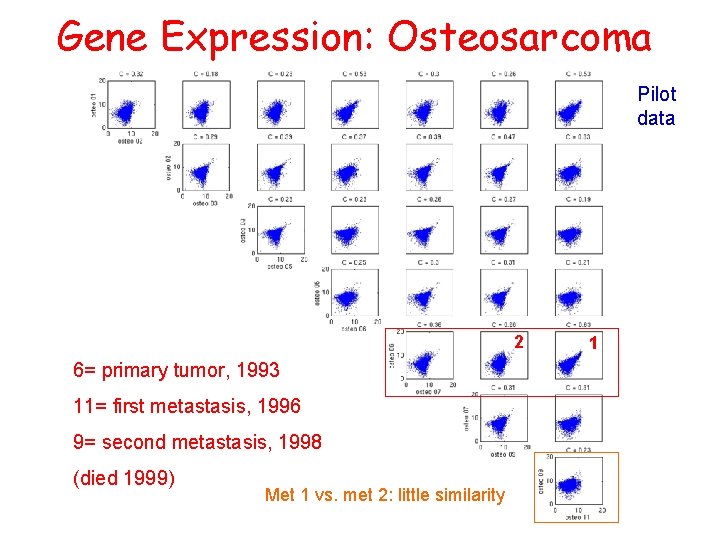
Gene Expression: Osteosarcoma Pilot data 2 6= primary tumor, 1993 11= first metastasis, 1996 9= second metastasis, 1998 (died 1999) Met 1 vs. met 2: little similarity 1
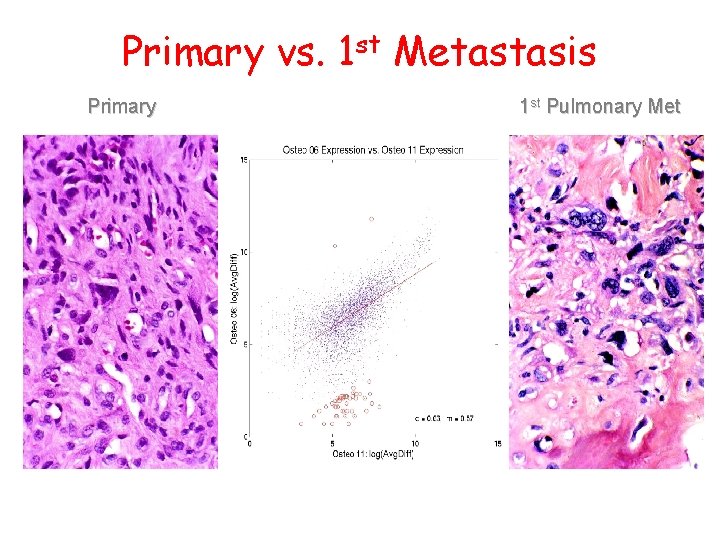
Primary vs. 1 st Metastasis Primary 1 st Pulmonary Met
Differential Gene Expression: Primary vs. Metastatic Osteosarcoma Osteonectin lost in metastasis
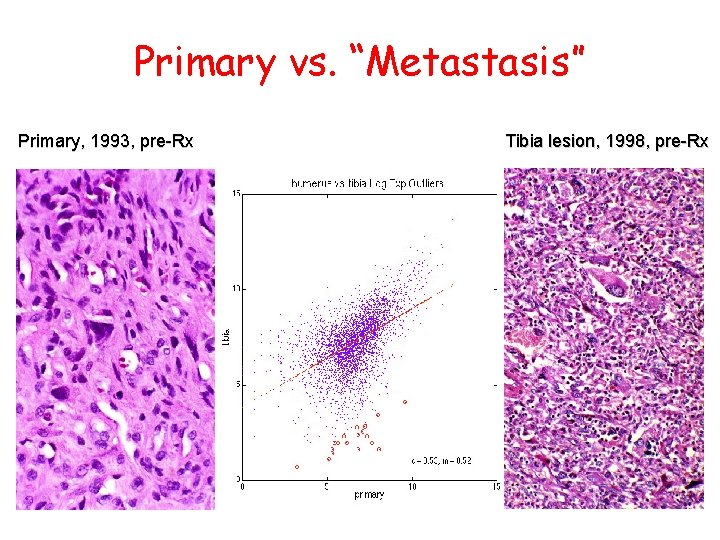
Primary vs. “Metastasis” Primary, 1993, pre-Rx Tibia lesion, 1998, pre-Rx
Gene Expression Data Clustering Multiple methods work
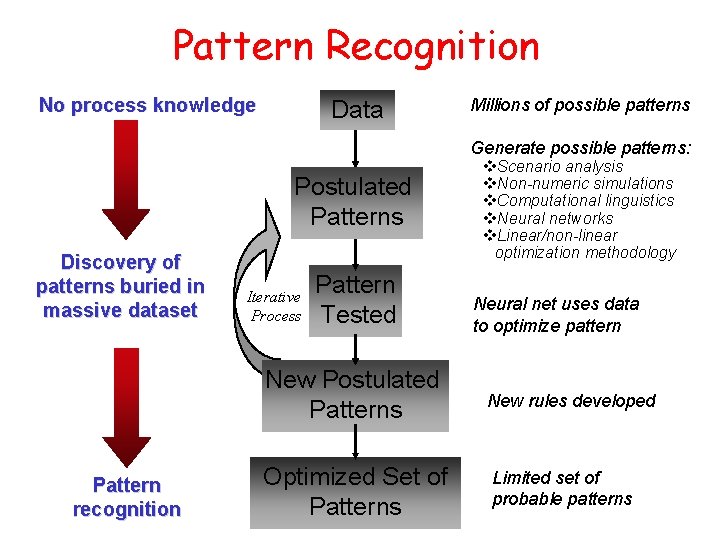
Pattern Recognition No process knowledge Data Millions of possible patterns Generate possible patterns: Postulated Patterns Discovery of patterns buried in massive dataset Iterative Process Pattern Tested New Postulated Patterns Pattern recognition Optimized Set of Patterns v. Scenario analysis v. Non-numeric simulations v. Computational linguistics v. Neural networks v. Linear/non-linear optimization methodology Neural net uses data to optimize pattern New rules developed Limited set of probable patterns

Agglomerative vs. Optimizing Hierarchical Clustering • Both build a tree of clusters, with data points as leaves, & “nearby” data points as siblings. • Agglomerative method repeatedly finds closest pair and irreversibly groups them. Bottom-up. Binary tree. • Optimization methods reconsider assignments based on other assignments and their effects on cluster means & variances. • Minimize sum of squared distances. – Distance measure matters. – Relate to statistical noise models, co-regulation models & likelihood of fit.

Agglomerative vs. Optimizing Hierarchical Clustering, cont. • Optimize means, variances, and cluster memberships. • Currently we optimize top-down, by levels • Expectation Maximization: soft memberships. K-means: hard. • Optimize tree topology (fanout) by CV • SOM also optimizes at one level, and requires low-dimensional grid embedding of cluster means. • Alternative to data-cluster distances: cliques of low data-data distances. Also has EM-like stat mech algorithms.

Mimir User Interface Courtesy of Eric Mjolsness, JPL
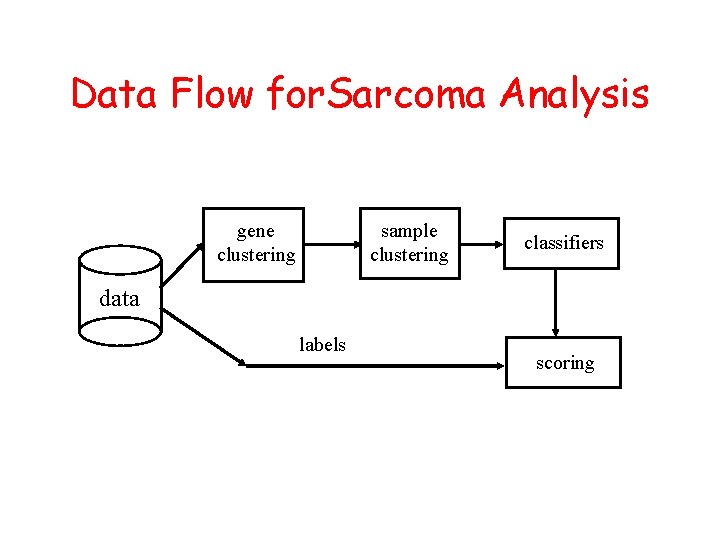
Data Flow for. Sarcoma Analysis gene clustering sample clustering classifiers data labels scoring

Pilot Study of Sarcomas 17 cases of osteosarcoma and rhabdomyosarcoma 6800 Gene. Chip analysis 6800 genes yield 14 gene clusters Reduced mean space yields 4 sample clusters OS OS OS, OS ERMS OS, ARMS 1ª ERMS X 4 OS x 3 ARMS met X 3
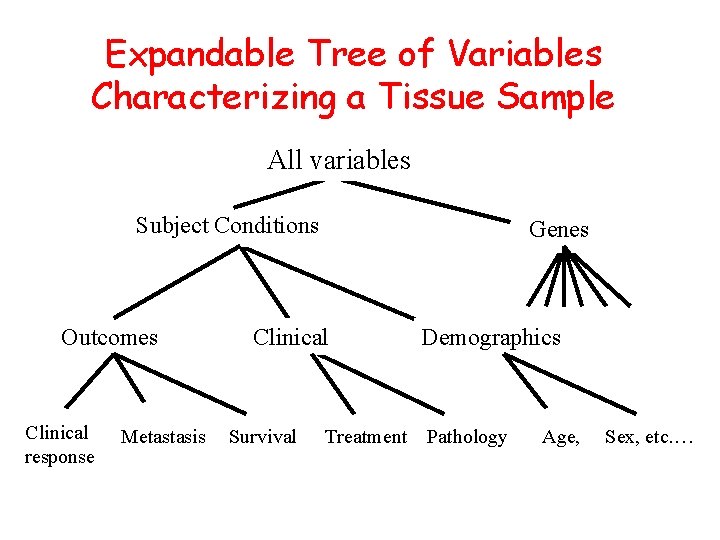
Expandable Tree of Variables Characterizing a Tissue Sample All variables Subject Conditions Outcomes Clinical response Metastasis Genes Clinical Survival Demographics Treatment Pathology Age, Sex, etc. …
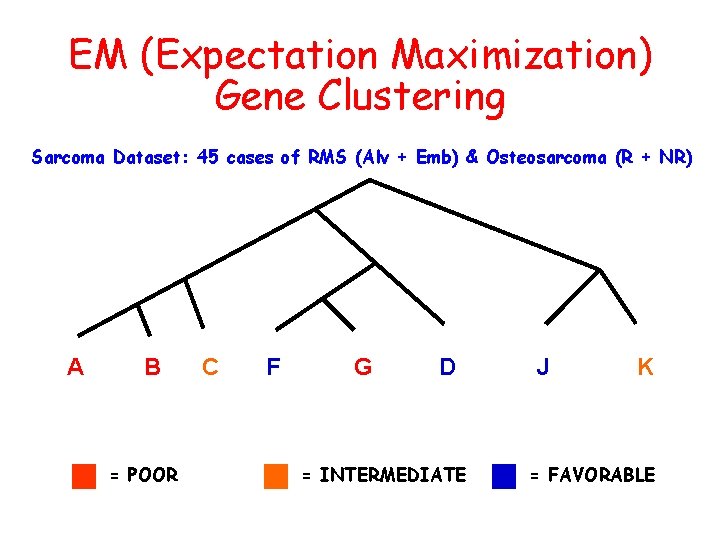
EM (Expectation Maximization) Gene Clustering Sarcoma Dataset: 45 cases of RMS (Alv + Emb) & Osteosarcoma (R + NR) A B = POOR C F G D = INTERMEDIATE J K = FAVORABLE

Working hypothesis: Gene expression profiling can detect prognostic distinctions among sarcomas independently of conventional clinical or diagnostic criteria

Future Directions • Analyze larger data set (institutional, COG) to test hypothesis • Expand to all sarcomas (RMS, non-RMS, OS, ESFTs) • Identify biologically important genes • Creation of custom “sarcoma” arrays using oligomers representing these genes • Long term studies of COG sarcoma patients using arrays in context with current clinical & biology studies
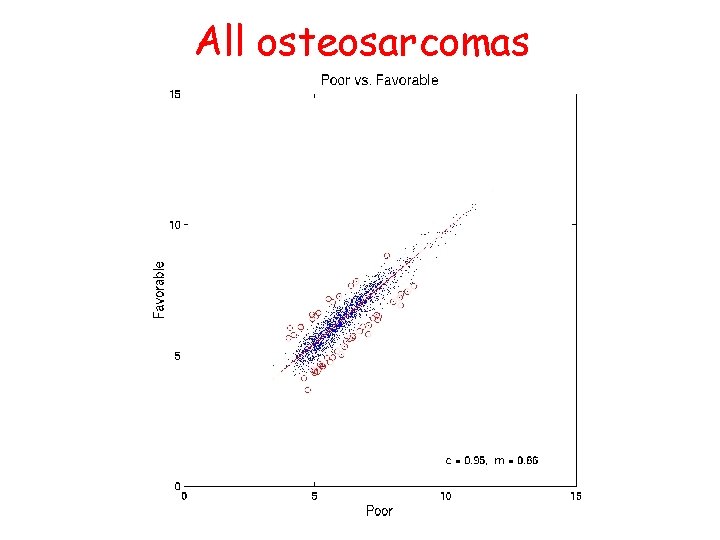
All osteosarcomas

Osteosarcoma vs RMS Genes
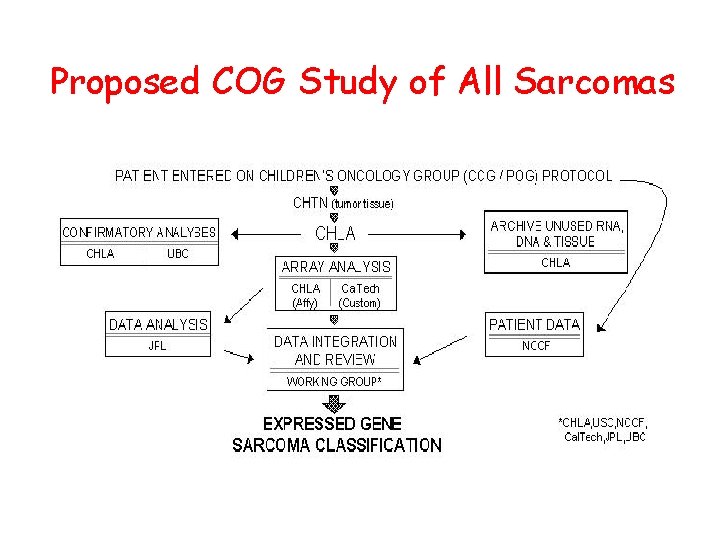
Proposed COG Study of All Sarcomas

Acknowledgements • CHLA: – Deb Schofield – Jingsong Zhang • USC: – Jonathan Buckley – Kim Siegmund • NCCF: – Mark Krailo • Caltech: – Barbara Wold – Chris Hart • JPL: – – Eric Mjolsness Tobias Mann Joe Roden Bornstein • UBC: – Poul Sorensen
- Slides: 30