Investigation of Bradi 4 g 40240 in Brachypodium

Investigation of Bradi 4 g 40240 in Brachypodium distachyon Young Choi and David Abraham

What is the function of Bradi 4 g 40240? How does a high impact mutation affect the gene?

Brachypodium distachyon as a model organism ● Common model organism ○ Easily grown ○ Self fertilizes ● Lots of info on B. distachyon https: //ceb. bio. uci. edu/data/invasive-species-handbook/brachypodiumdistachyon/

Anticipated Outcomes ● Better understanding of a gene in B. distachyon ● More data on plant mutations ○ Implications on GMOs

Forming a Gene Plot
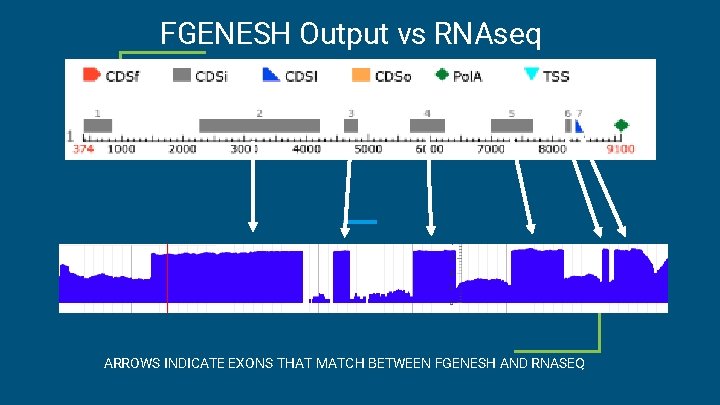
FGENESH Output vs RNAseq ARROWS INDICATE EXONS THAT MATCH BETWEEN FGENESH AND RNASEQ
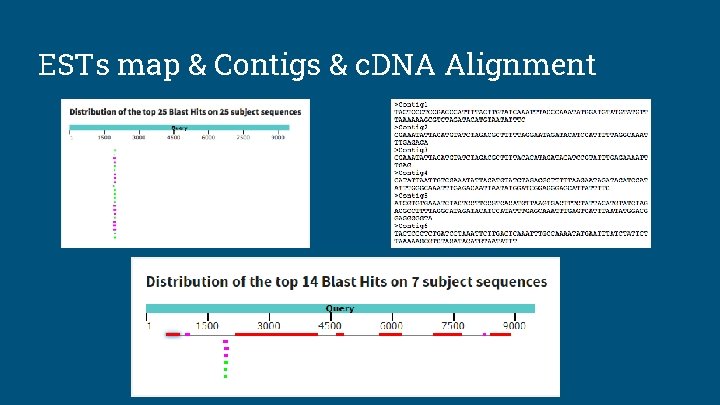
ESTs map & Contigs & c. DNA Alignment
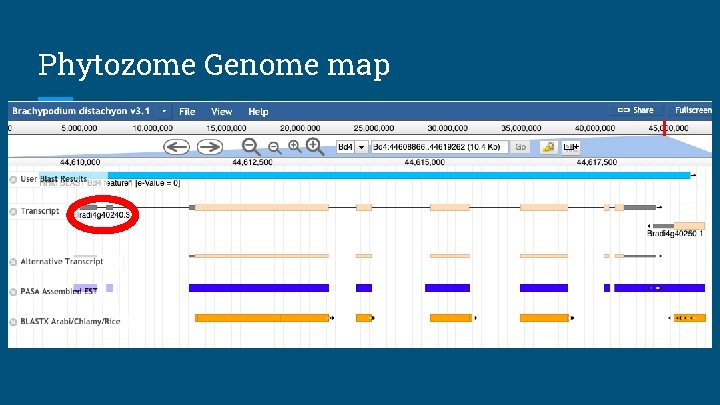
Phytozome Genome map
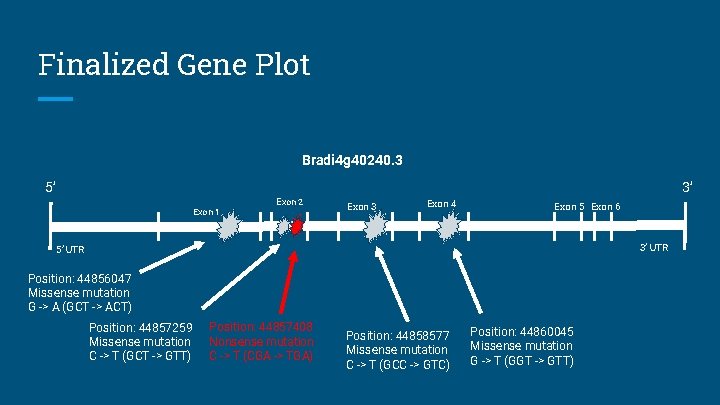
Finalized Gene Plot Bradi 4 g 40240. 3 5’ 3’ Exon 1 Exon 2 Exon 3 Exon 4 Exon 5 Exon 6 3’ UTR 5’ UTR Position: 44856047 Missense mutation G -> A (GCT -> ACT) Position: 44857259 Missense mutation C -> T (GCT -> GTT) Position: 44857408 Nonsense mutation C -> T (CGA -> TGA) Position: 44858577 Missense mutation C -> T (GCC -> GTC) Position: 44860045 Missense mutation G -> T (GGT -> GTT)

Finding a Function for the Bradi 4 g 40240 Gene: BLAST and Phytozome Evidence

BLASTN Results Zea mays STICHEL Elaeis guineensis STICHEL Oryza sativa STICHEL Other plants that are not shown on this figure: Nicotiana tabacum, Coffea eugenioides
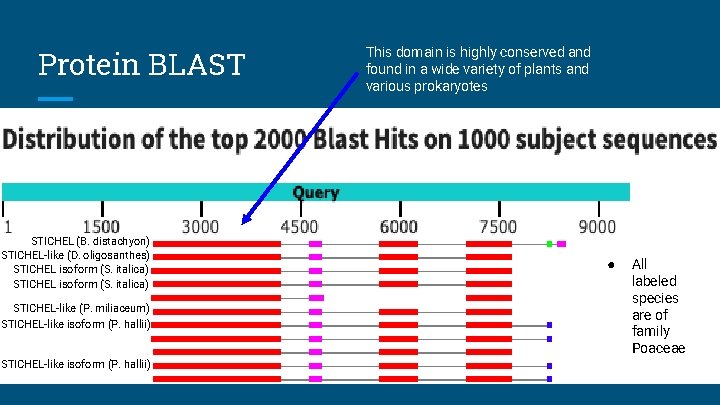
Protein BLAST STICHEL (B. distachyon) STICHEL-like (D. oligosanthes) STICHEL isoform (S. italica) STICHEL-like (P. miliaceum) STICHEL-like isoform (P. hallii) This domain is highly conserved and found in a wide variety of plants and various prokaryotes ● All labeled species are of family Poaceae
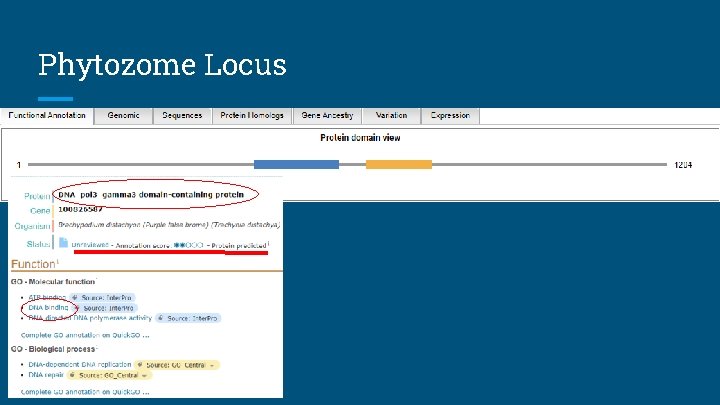
Phytozome Locus
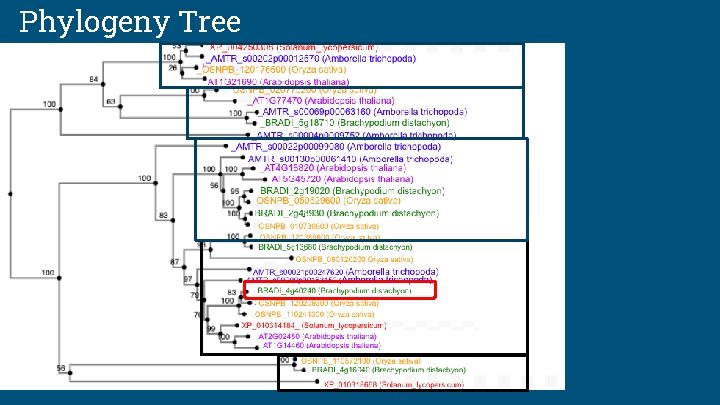
Phylogeny Tree

Finding a Function for the Bradi 4 g 40240 Gene: Sanger Sequencing
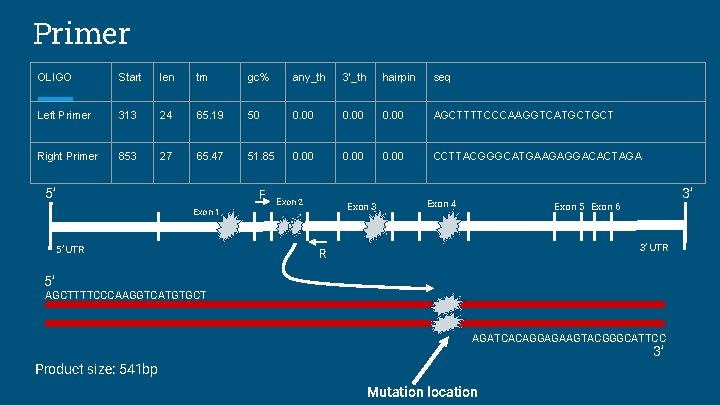
Primer OLIGO Start len tm gc% any_th 3’_th hairpin seq Left Primer 313 24 65. 19 50 0. 00 AGCTTTTCCCAAGGTCATGCTGCT Right Primer 853 27 65. 47 51. 85 0. 00 CCTTACGGGCATGAAGAGGACACTAGA 5’ F Exon 1 5’ UTR Exon 2 Exon 3 Exon 4 3’ Exon 5 Exon 6 3’ UTR R 5’ AGCTTTTCCCAAGGTCATGTGCT AGATCACAGGAGAAGTACGGGCATTCC 3’ Product size: 541 bp Mutation location
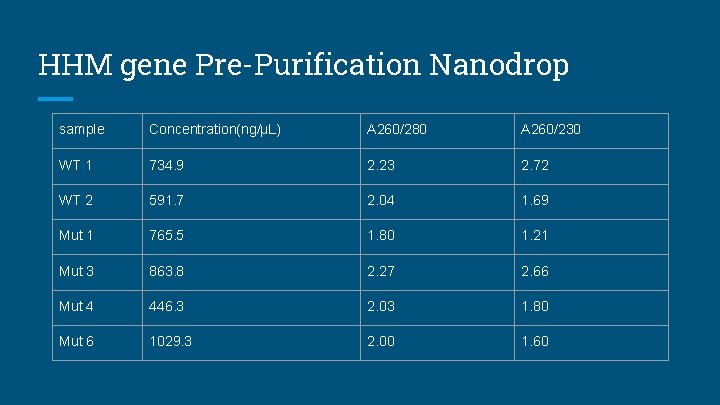
HHM gene Pre-Purification Nanodrop sample Concentration(ng/µL) A 260/280 A 260/230 WT 1 734. 9 2. 23 2. 72 WT 2 591. 7 2. 04 1. 69 Mut 1 765. 5 1. 80 1. 21 Mut 3 863. 8 2. 27 2. 66 Mut 4 446. 3 2. 03 1. 80 Mut 6 1029. 3 2. 00 1. 60
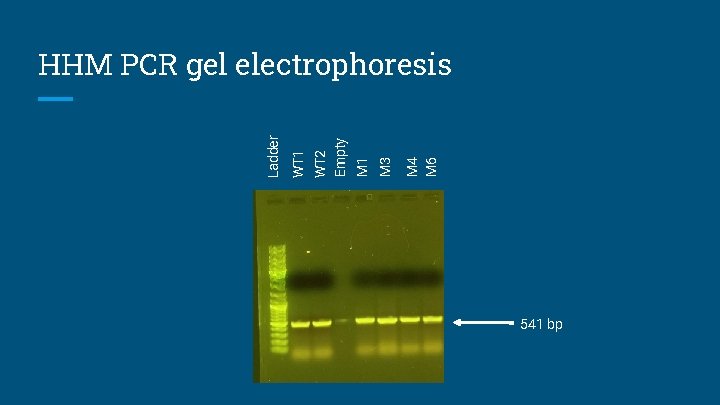
M 4 M 6 M 3 M 1 WT 2 Empty WT 1 Ladder HHM PCR gel electrophoresis 541 bp
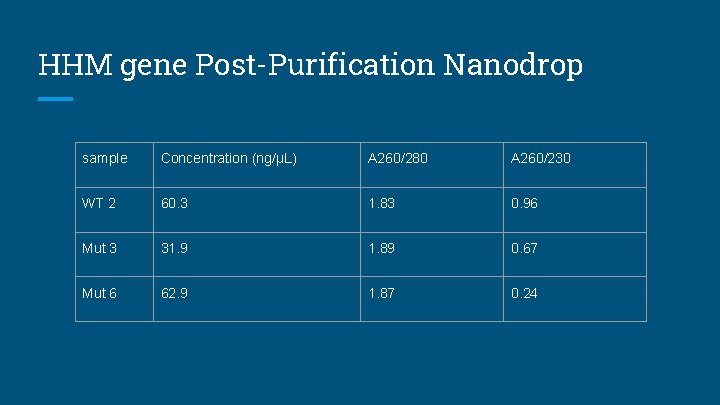
HHM gene Post-Purification Nanodrop sample Concentration (ng/µL) A 260/280 A 260/230 WT 2 60. 3 1. 83 0. 96 Mut 3 31. 9 1. 89 0. 67 Mut 6 62. 9 1. 87 0. 24

Sanger Sequencing Results Section of M 3 Sequencing

Differential Gene Expression in Bradi 4 g 40240
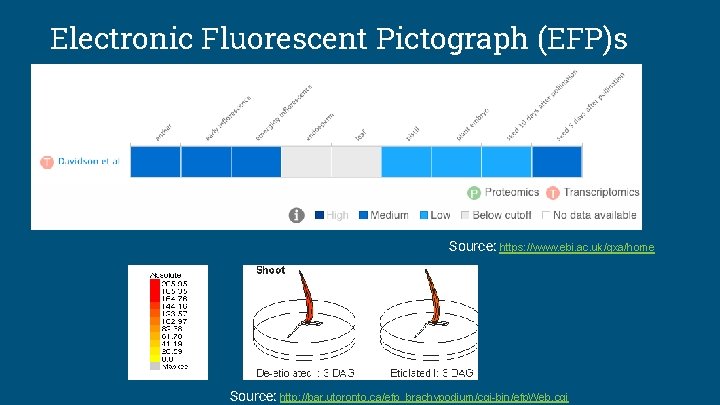
Electronic Fluorescent Pictograph (EFP)s Source: https: //www. ebi. ac. uk/gxa/home Source: http: //bar. utoronto. ca/efp_brachypodium/cgi-bin/efp. Web. cgi

HHM Gene Expression from Data Mining Source: https: //phytozome. jgi. doe. gov/
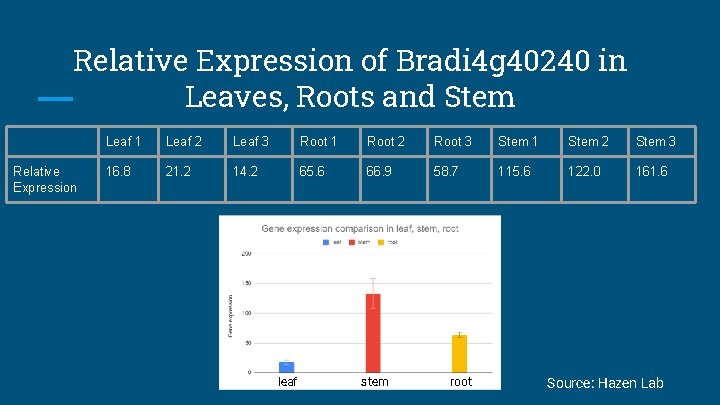
Relative Expression of Bradi 4 g 40240 in Leaves, Roots and Stem Relative Expression Leaf 1 Leaf 2 Leaf 3 Root 1 Root 2 Root 3 Stem 1 Stem 2 Stem 3 16. 8 21. 2 14. 2 65. 6 66. 9 58. 7 115. 6 122. 0 161. 6 leaf stem root Source: Hazen Lab
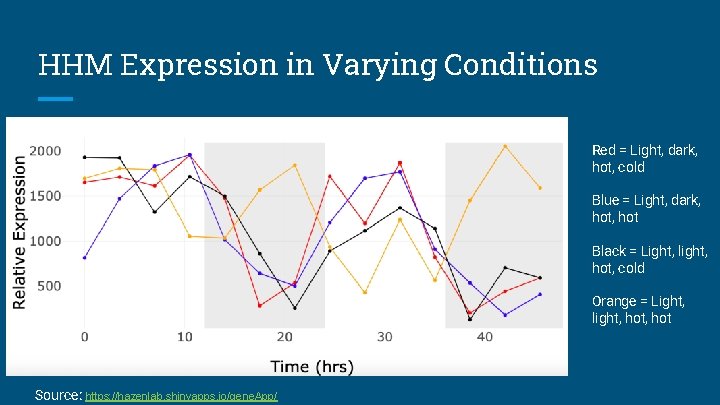
HHM Expression in Varying Conditions Red = Light, dark, hot, cold Blue = Light, dark, hot Black = Light, light, hot, cold Orange = Light, light, hot Source: https: //hazenlab. shinyapps. io/gene. App/

Statistical Analysis of Gene Expression Two-tailed distribution ttest is used. p=0. 991 p=0. 302 p=0. 304 p=0. 00019* p=0. 0370* p=0. 0137*

Wild Type vs. Mutant: Cell Wall Thickness
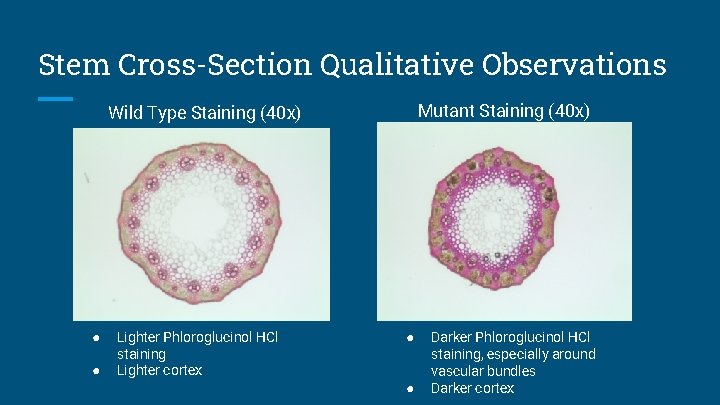
Stem Cross-Section Qualitative Observations Mutant Staining (40 x) Wild Type Staining (40 x) ● ● Lighter Phloroglucinol HCl staining Lighter cortex ● ● Darker Phloroglucinol HCl staining, especially around vascular bundles Darker cortex
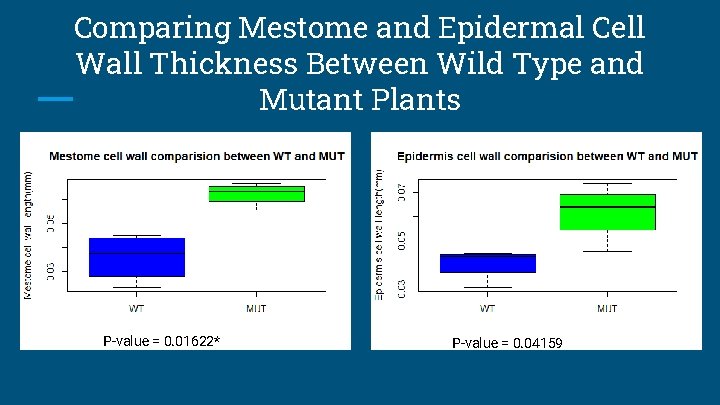
Comparing Mestome and Epidermal Cell Wall Thickness Between Wild Type and Mutant Plants P-value = 0. 01622* P-value = 0. 04159

Conclusions on Gene Function

Postulated WT STICHEL Bradi 4 g 40240 Gene Function 5’ Postulated Mutant STICHEL 3’ d pro 3’ UTR CELL ion ss in e r te p ex Pro e L n Ge ICHE ST HOM E L STICHE es uc 5’ UTR STICHEL protein stabilizes microtubules, leading to branching as a result of a branched microtubule conformation GET TAR HE tric L ent ho ers me t cel arget l TRIC ST IC Microtubule Organizing Center (MTOC)

Further Research
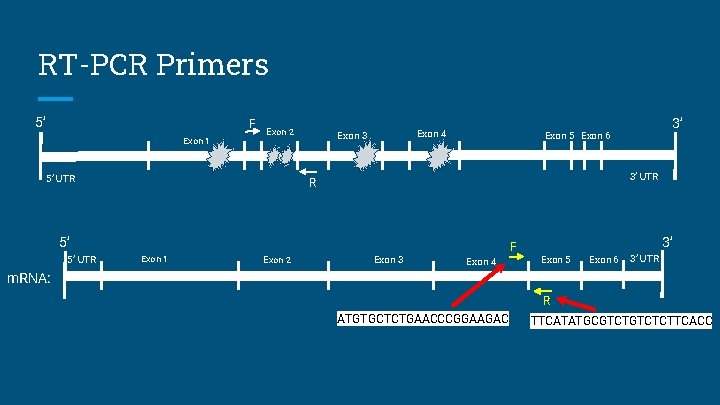
RT-PCR Primers 5’ F Exon 1 Exon 2 5’ UTR Exon 4 Exon 3 3’ UTR R 5’ 5’ UTR Exon 1 Exon 2 3’ Exon 5 Exon 6 Exon 3 F Exon 4 3’ Exon 5 Exon 6 3’ UTR m. RNA: R ATGTGCTCTGAACCCGGAAGAC TTCATATGCGTCTCTTCACC
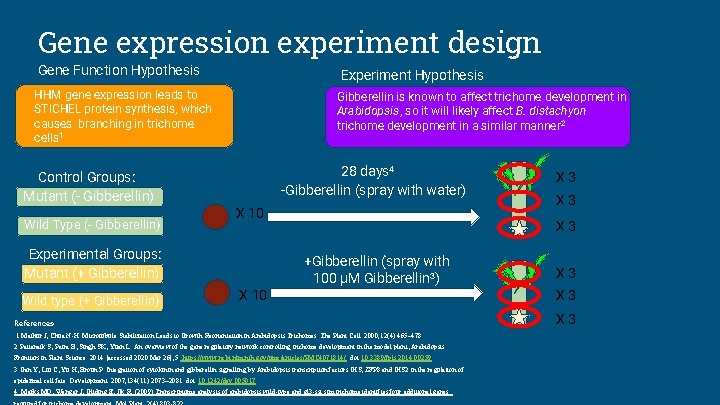
Gene expression experiment design Gene Function Hypothesis Experiment Hypothesis HHM gene expression leads to STICHEL protein synthesis, which causes branching in trichome cells 1 Control Groups: Mutant (- Gibberellin) Wild Type (- Gibberellin) Gibberellin is known to affect trichome development in Arabidopsis, so it will likely affect B. distachyon trichome development in a similar manner 2 28 days 4 -Gibberellin (spray with water) X 10 Experimental Groups: Mutant (+ Gibberellin) Wild type (+ Gibberellin) X 3 X 3 +Gibberellin (spray with 100 µM Gibberellin 3) X 10 References 1. Mathur J, Chua N-H. Microtubule Stabilization Leads to Growth Reorientation in Arabidopsis Trichomes. The Plant Cell. 2000; 12(4): 465– 478. 2. Pattanaik S, Patra B, Singh SK, Yuan L. An overview of the gene regulatory network controlling trichome development in the model plant, Arabidopsis. Frontiers in Plant Science. 2014 [accessed 2020 Mar 26]; 5. https: //www. ncbi. nlm. nih. gov/pmc/articles/PMC 4071814/. doi: 10. 3389/fpls. 2014. 00259 3. Gan Y, Liu C, Yu H, Broun P. Integration of cytokinin and gibberellin signalling by Arabidopsis transcription factors GIS, ZFP 8 and GIS 2 in the regulation of epidermal cell fate. Development. 2007; 134(11): 2073– 2081. doi: 10. 1242/dev. 005017 4. Marks MD, Wenger J, Gliding E, Jik R. (2009) Transcriptome analysis of arabidopsis wild-type and gl 3 -sst sim trichome identifies four additional genes X 3 X 3

Summary ● Bradi 4 g 40240 contains 6 exons ● Nonsense mutation located in Exon 6 ● Bradi 4 g 40240 gene is conserved in many other plants including O. sativa, A. thaliana ● High gene expression is observed in stems and developing young B. distachyon.

References 1. Draper J, Mur LAJ, Jenkins G, Ghosh-Biswas GC, Bablak P, Hasterok R, Routledge APM. Brachypodium distachyon. A New Model System for Functional Genomics in Grasses. Plant Physiology. 2001; 127(4): 1539– 1555. 2. Mathur J, Chua N-H. Microtubule Stabilization Leads to Growth Reorientation in Arabidopsis Trichomes. The Plant Cell. 2000; 12(4): 465– 478. 3. Manzaneda AJ, Rey PJ, Bastida JM, Weiss-Lehman C, Raskin E, Mitchell-Olds T. Environmental aridity is associated with cytotype segregation and polyploidy occurrence in Brachypodium distachyon (Poaceae). The New Phytologist. 2012; 193(3): 797– 805. doi: 10. 1111/j. 1469 -8137. 2011. 03988. x 4. Xi A, Yang X, Deng M, Chen Y, Shao J, Zhao J, An L. Isolation and identification of two new alleles of STICHEL in Arabidopsis. Biochemical and Biophysical Research Communications. 2018; 499(3): 605– 610. doi: 10. 1016/j. bbrc. 2018. 03. 197
- Slides: 36