Investigating Sequences Stephen Everse University of Vermont Biochemical

Investigating Sequences Stephen Everse University of Vermont
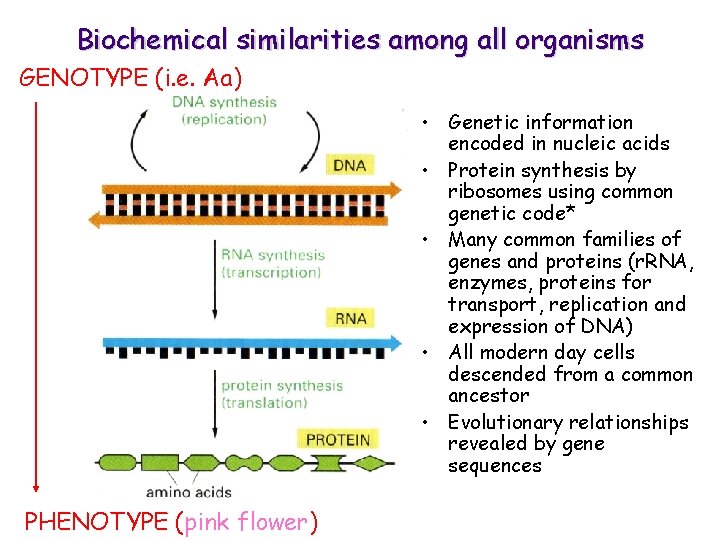
Biochemical similarities among all organisms GENOTYPE (i. e. Aa) • Genetic information encoded in nucleic acids • Protein synthesis by ribosomes using common genetic code* • Many common families of genes and proteins (r. RNA, enzymes, proteins for transport, replication and expression of DNA) • All modern day cells descended from a common ancestor • Evolutionary relationships revealed by gene sequences PHENOTYPE (pink flower)
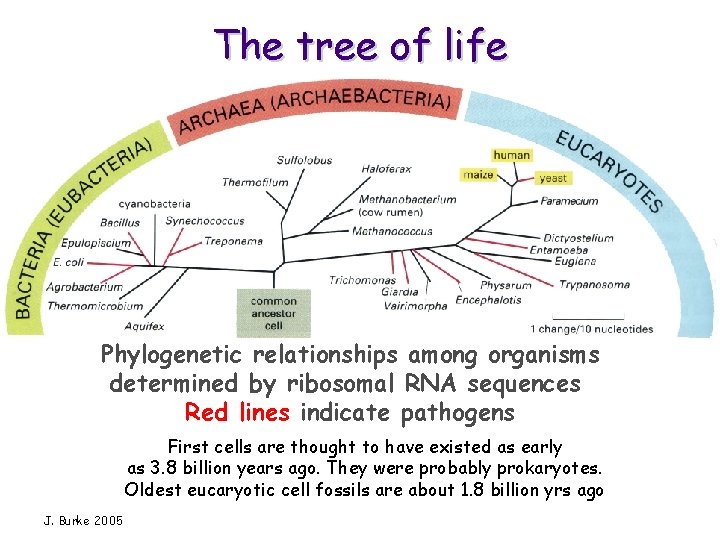
The tree of life Phylogenetic relationships among organisms determined by ribosomal RNA sequences Red lines indicate pathogens First cells are thought to have existed as early as 3. 8 billion years ago. They were probably prokaryotes. Oldest eucaryotic cell fossils are about 1. 8 billion yrs ago J. Burke 2005
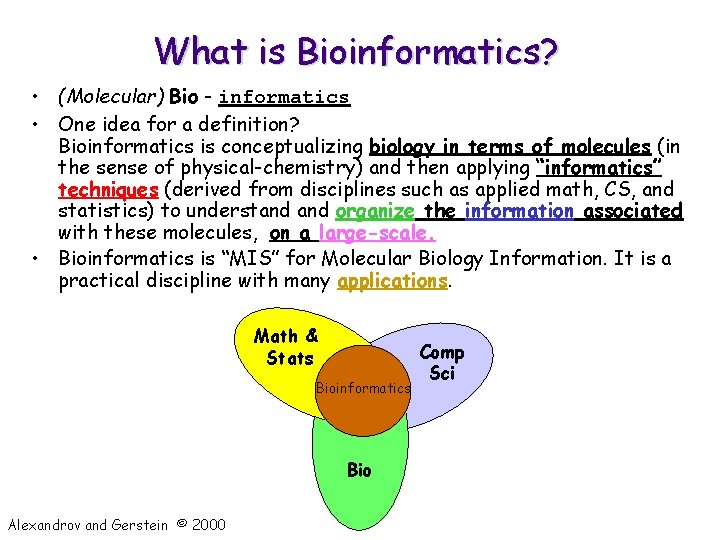
What is Bioinformatics? • (Molecular) Bio - informatics • One idea for a definition? Bioinformatics is conceptualizing biology in terms of molecules (in the sense of physical-chemistry) and then applying “informatics” techniques (derived from disciplines such as applied math, CS, and statistics) to understand organize the information associated with these molecules, on a large-scale. • Bioinformatics is “MIS” for Molecular Biology Information. It is a practical discipline with many applications. Math & Stats Bioinformatics Bio Alexandrov and Gerstein © 2000 Comp Sci

Areas of current and future development of bioinformatics • Molecular biology and genetics • Phylogenetic and evolutionary sciences • Different aspects of biotechnology including pharmaceutical and microbiological industries • Medicine • Agriculture • Eco-management
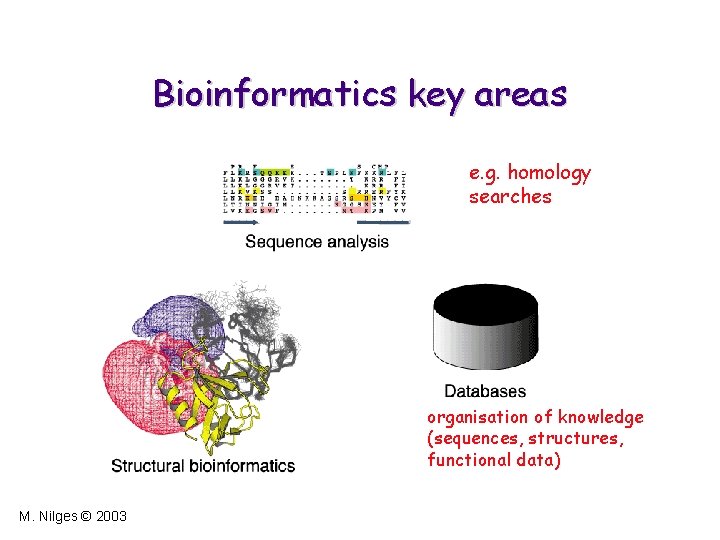
Bioinformatics key areas e. g. homology searches organisation of knowledge (sequences, structures, functional data) M. Nilges © 2003

Why do we want to compare sequences? • Relationships – Phylogenetic trees can be constructed based on comparison of the sequences of a molecule (example: 16 S r. RNA) taken from different species – Residues conserved during evolution play an important role • Prediction of protein structure and function – Proteins which are very similar in sequence generally have similar 3 D structure and function as well – By searching a sequence of unknown structure against a database of known proteins the structure and/or function can in many cases be predicted • Center for Biological Sequence Analysis © 2001
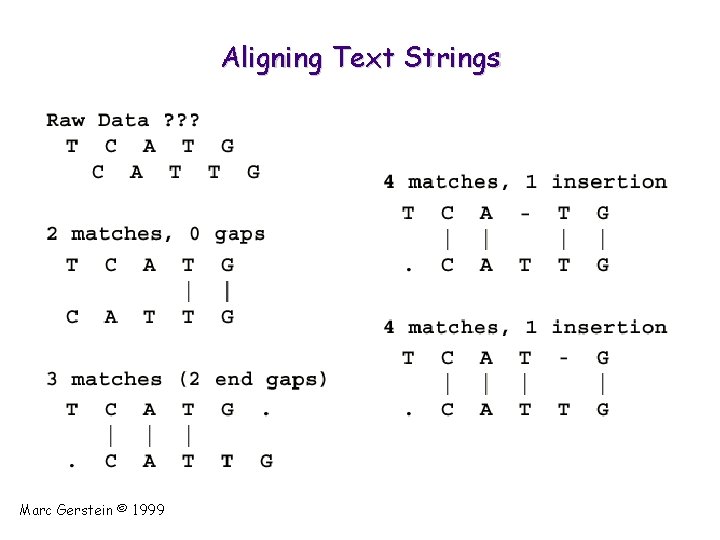
Aligning Text Strings Marc Gerstein © 1999

Mol Bio Information - Protein Marc Gerstein © 1999

Summary • Central dogma of biology generates material appropriate for bioinformatical study (DNA, RNA, proteins, phenotype, etc) • One form of bioinformatics is the comparison of sequences • BUT, How do we bring this to the classroom?
Creating Inquiry Opportunities Domain Principles Analysis Tools Data Sets
Establishing a Problem Space Domain Principles Problem Space Analysis Data Tools Sets Creating problem spaces that provide a rich context for using bioinformatics data and tools allows students to focus on using their understanding of biology to investigate meaningful questions.

Problem Spaces • Foundation – – – Introduction Background Data Tools Bibliography Curricular Resources • Starting Points

Malaria is caused by one of four species of Plasmodium (falciparum, vivax, malariae and ovale). Of these P. falciparum is the most lethal being estimated to cause 200 million clinical cases, and 1 -3 million deaths (including many children) every year world-wide.
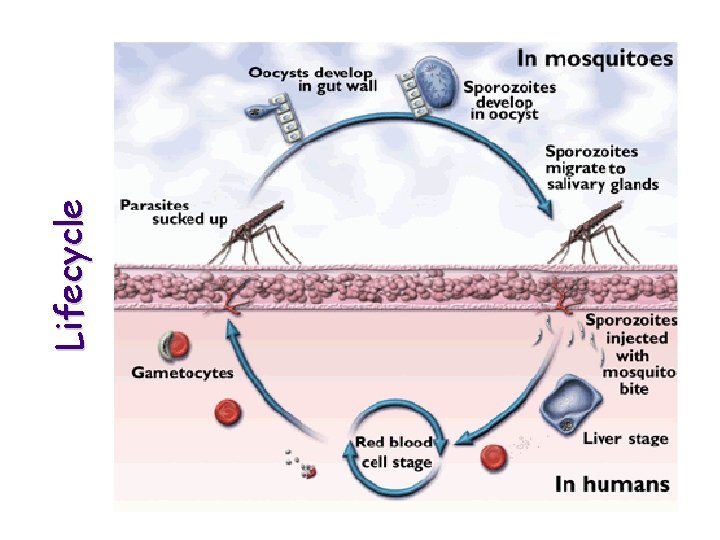
Lifecycle

The Plasmodium falciparum Genome- A Consortium Project (chromosomes 1, 3 -9, 13) (chromosomes 2, 10, 11 and 14) (chromosome 12)

Plasmodium genomics special issue Nature 3 rd October 2002

Plasmodium falciparum Genome Project Curation To maximise the benefits to the scientific community of Plasmodium genome sequencing, the Pathogen Genomics group is committed to the curation of Plasmodium spp. This will ensure that annotation is updated and maintained, and will form a framework that underpins global efforts to understand the parasite and the disease it causes. If you would like to contribute to the curation of any gene(s) please contact the curator ucb@sanger. ac. uk and visit Gene. DB. http: //www. sanger. ac. uk/Projects/P_falciparum/ See how Brad Goodner at Hiram College involves his students in curation: http: //www. hiram. edu/biology/faculty/goodner. html

Drug search … International computing grid searches for malaria drugs 2/2/07. Using an international computing grid spanning 27 countries, scientists on the WISDOM project analysed an average of 80 000 possible drug compounds against malaria every hour. In total, the challenge processed over 140 million compounds, with a UK physics grid providing nearly half of the computing hours used. http: //malaria. wellcome. ac. uk/doc_WTX 037265. html Enabling Grids for E-scienc. E (EGEE) is the largest multidisciplinary grid infrastructure in the world, which brings together more than 120 organizations to produce a reliable and scalable computing resource available to the global research community. At present, it consists of 250 sites in 48 countries and more than 68, 000 CPUs available to some 8, 000 users 24 hours a day, 7 days a week http: //www. eu-egee. org/

The data Genetic structure of Plasmodium falciparum field isolates in eastern and north-eastern India H. Joshi, N. Valecha, A. Verma, A. Kaul, P. K. Mallick, S. Shalini, S. K. Prajapati, S. K. Sharma, V. Dev, S. Biswas, N. Nanda, M. S. Malhotra, S. K. Subbarao & A. P. Dash. Malaria Journal 2007 Vol 6 Page 60 http: //www. malariajournal. com/content/6/1/60

The study… • Isolates were collected from microscopically diagnosed P. falciparum positive subjects in three Indian states with varied malaria epidemiology; • Merozoite surface protein-1 (MSP-1, 17 k. Da) & protein -2 (MSP-2, 46 -53 k. Da) of P. falciparum is a target of the host's humoral immunity and a malaria vaccine candidate; and • 131 P. falciparum isolates of msp-1 (block 2) and msp-2 (central repeat region, block 3) were obtained as well as others from Genbank. msp-1 blocks Hoffmann et al. (2003) Malaria J. 2: 24

Block 2 of msp-1 Nucleotide Sequence aatgaagaag gtggtgcaagtgct agtgctcaaa gtacaagtcc aaatacttca aaattactac tgctcaaagtggtg gtggtacaag atcatctcgt tctggtgcaa aaaaggtgcaagtg caagtgctca tggtccaagt tcaaacactt gccctccagc agtgctcaaagtgg aagtggtgca ggtccaagtg tacctcgttc tgatgcaagc Amino Acid Sequence NEEEITTKGASAQSGASAQSGASAQSGTSGPSGPSGTSPSSRSNTLPRSNTSSGASPP ADAS
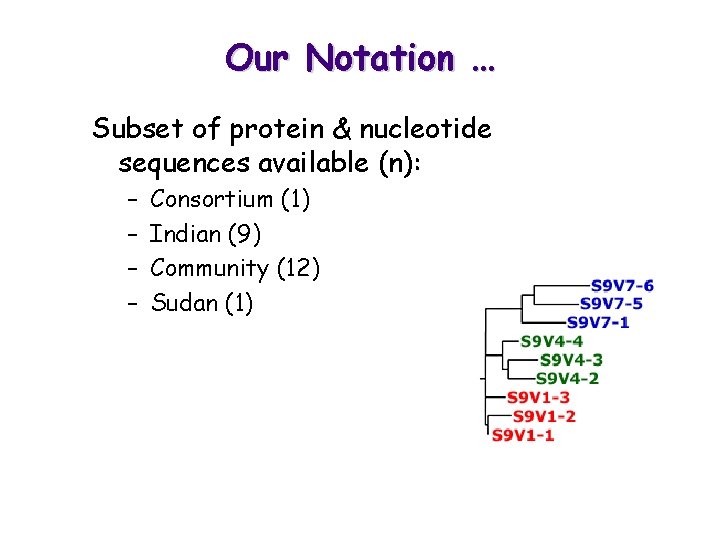
Our Notation … Subset of protein & nucleotide sequences available (n): – – Consortium (1) Indian (9) Community (12) Sudan (1)

Our workspace … • National Center Biological Information (http: //ncbi. nlm. nih. gov) • Biology Workbench (http: //workbench. sdsc. edu)

Malaria Triad: Genetics & Genomics This web resource provides data and information relevant to malaria genetics and genomics. These resources include organism specific sequence BLAST databases (Plasmodium falciparum only, all Plasmodium ), genome maps, linkage markers, and information about genetic studies. Links are provided for other malaria web sites and genetic data on related apicomplexan parasites. http: //www. ncbi. nlm. nih. gov/projects/Malaria/

The tools … • Session Tools ~ file folders • Protein Tools/Nucleic Tools – Find sequences (Ndjinn) – Upload sequences (Add) – Align sequences (CLUSTALW) • Alignment Tools – Display options (BOXSHADE, DRAWGRAM)

Let’s explore … > Sequence 1 GAGGTAGTAATTAGATCCGAAA… > Sequence 2 GAGGTAGTAATTAGATCTGAAA… > Sequence 3 GAGGTAGTAATTAGATCTGTCA… • Form groups of ~3/computer • Look at the data (http: //bioquest. org/oakwood_2008/malaria-problem-space) • Choose a problem/question to explore

Favorite movie of the week … Inside the Cell Harvard Bio. Visions Video What you are seeing is discussed here

Homology • Homologous sequences can be divided into two groups – orthologous sequences: sequences that differ because they are found in different species (e. g. human a-globin and mouse a-globin) – paralogous sequences: sequences that differ because of a gene duplication event (e. g. human a-globin and human bglobin, various versions of both ) M. Craven @ 2002
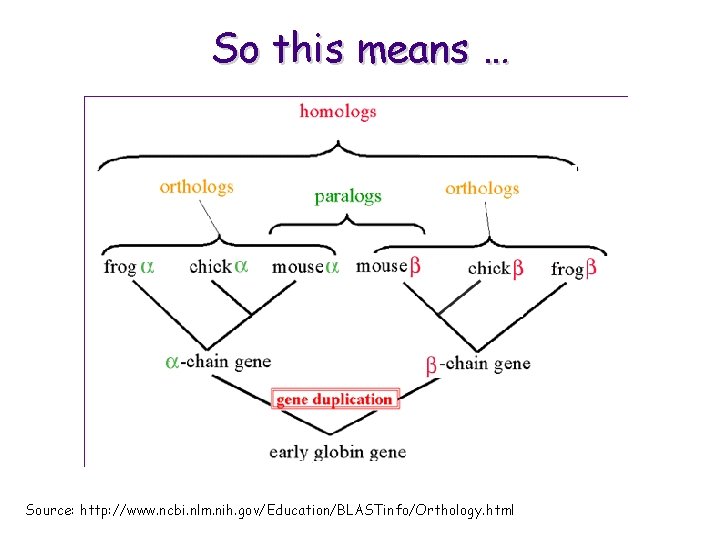
So this means … Source: http: //www. ncbi. nlm. nih. gov/Education/BLASTinfo/Orthology. html

Search algorithms • Smith-Waterman (1981) • FASTA (Pearson 1995) • BLAST (Altschul 1990, 1997) – demanding of time and memory resources – Speed up searches by an order of magnitude compared to Smith. Waterman – Good statistics – Extremely fast • One order of magnitude faster than FASTA • Two orders of magnitude faster than Smith-Waterman – Almost as sensitive as FASTA

Things to keep in mind when working with alignments • Pairwise alignment programs always find the optimal alignment of two sequences – They do so even if it does not make any sense at all to align the two sequences – ”Optimal” means optimal according to the substitution matrix and gap penalties you choose – also if you choose the wrong ones • Generally the underlying assumptions are wrong – The frequency of substitution is not the same at all positions – Nor is the frequencies of insertions and deletions the same – Affine gap penalties do not properly model indel events Center for Biological Sequence Analysis © 2001
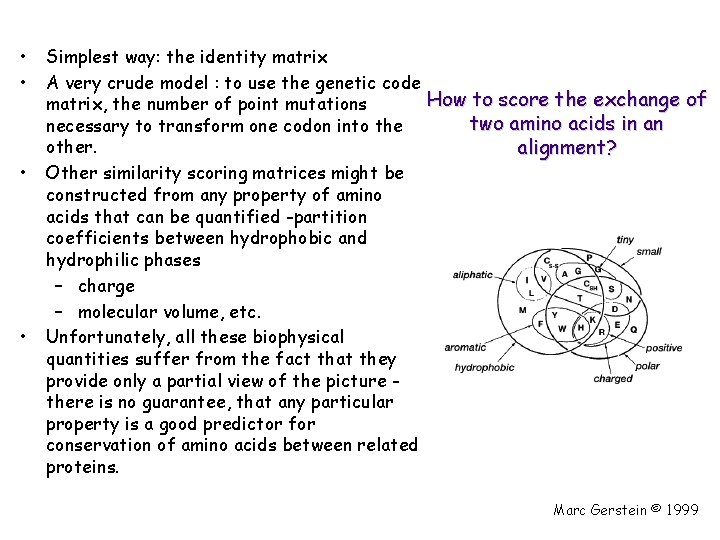
• • Simplest way: the identity matrix A very crude model : to use the genetic code How to score the exchange of matrix, the number of point mutations two amino acids in an necessary to transform one codon into the other. alignment? Other similarity scoring matrices might be constructed from any property of amino acids that can be quantified -partition coefficients between hydrophobic and hydrophilic phases – charge – molecular volume, etc. Unfortunately, all these biophysical quantities suffer from the fact that they provide only a partial view of the picture there is no guarantee, that any particular property is a good predictor for conservation of amino acids between related proteins. Marc Gerstein © 1999

Pairwise alignment of hemoglobin alpha chain and myoglobin 24. 7% identity; Global alignment score: 130 10 20 30 40 50 HBA_HU VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHFDLSHGSAQVKG--: : : . . . : : : . : . MYG_PH VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKTEAEMKASED 10 20 30 40 50 60 60 70 80 90 100 110 HBA_HU ---HGKKVADALTNAVAHVDDMPNALSALSDLHAHKLRVDPVNF-KLLSHCLLVTLAAHL : : : . . MYG_PH LKKHGVTVLTALGAILKKKGHHEAELKPLAQSHATKHKI-PIKYLEFISEAIIHVLHSRH 70 80 90 100 110 120 130 140 HBA_HU PAEFTPAVHASLDKFLASVSTVLTSKYR-----: . . . . : : . MYG_PH PGDFGADAQGAMNKALELFRKDIAAKYKELGYQG 120 130 140 150 Center for Biological Sequence Analysis © 2001

Important things to remember when using alignment to search databases • When searching in databases, size does matter! • Doing things differently can lead to different conclusions • Think before and after you search – Searching large databases take very long time – The significance of matches drops when the database is expanded – Nucleotide comparison vs. protein comparison – The obvious thing to do is not always the right thing to do – Conclusions based on matches should be drawn with greater care Marc Gerstein © 1999

Why multiple alignment is better • More sequences contain more information • Multiple sequence alignment allows us to compare all related proteins simultaneously • It allows us to identify features that are conserved among the sequences • Using a multiple sequence alignment (a profile) one can find more related sequences than by simple pairwise comparison Center for Biological Sequence Analysis © 2001

A multiple sequence alignment of globins HBB_HUMAN HBB_HORSE HBA_HUMAN HBA_HORSE MYG_PHYCA GLB 5_PETMA LGB 2_LUPLU ----VHLTPEEKSAVTALWGKVN--VDEVGGEALGRLLVVYPWTQRFFESFGDLST ----VQLSGEEKAAVLALWDKVN--EEEVGGEALGRLLVVYPWTQRFFDSFGDLSN -----VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLS-----VLSAADKTNVKAAWSKVGGHAGEYGAEALERMFLGFPTTKTYFPHF-DLS-----VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKT PIVDTGSVAPLSAAEKTKIRSAWAPVYSTYETSGVDILVKFFTSTPAAQEFFPKFKGLTT ----GALTESQAALVKSSWEEFNANIPKHTHRFFILVLEIAPAAKDLFSFLKGTSE *: : : * : . HBB_HUMAN HBB_HORSE HBA_HUMAN HBA_HORSE MYG_PHYCA GLB 5_PETMA LGB 2_LUPLU PDAVMGNPKVKA HGKKVLGAFSDGLAHLDN-----LKGTFAT LSELHCDKLHVDPENFRL PGAVMGNPKVKA HGKKVLHSFGEGVHHLDN-----LKGTFAA LSELHCDKLHVDPENFRL ----HGSAQVKG HGKKVADALTNAVAHVDD-----MPNALSA LSDLHAHKLRVDPVNFKL ----HGSAQVKA HGKKVGDALTLAVGHLDD-----LPGALSN LSDLHAHKLRVDPVNFKL EAEMKASEDLKK HGVTVLTALGAILKKKGH-----HEAELKP LAQSHATKHKIPIKYLEF ADQLKKSADVRW HAERIINAVNDAVASMDDT--EKMSMKLRD LSGKHAKSFQVDPQYFKV VP--QNNPELQA HAGKVFKLVYEAAIQLQVTGVVVTDATLKN LGSVHVSKGVAD-AHFPV. . : : *. *. : . Center for Biological Sequence Analysis © 2001
- Slides: 37