Intronic Splicing Regulatory Elements Genomewide comparative genomics Arraybased
Intronic Splicing Regulatory Elements Genome-wide comparative genomics Array-based discovery from neural differentiation of human ES cells C. Carson J. Simon Eric Van Nostrand, Tiffany Liang Gene Yeo, Salk Institute Nicole Coufal, Christian Carson, Alysson Muotri, Xiangdong Xu, Tiffany Liang, Rusty Gage
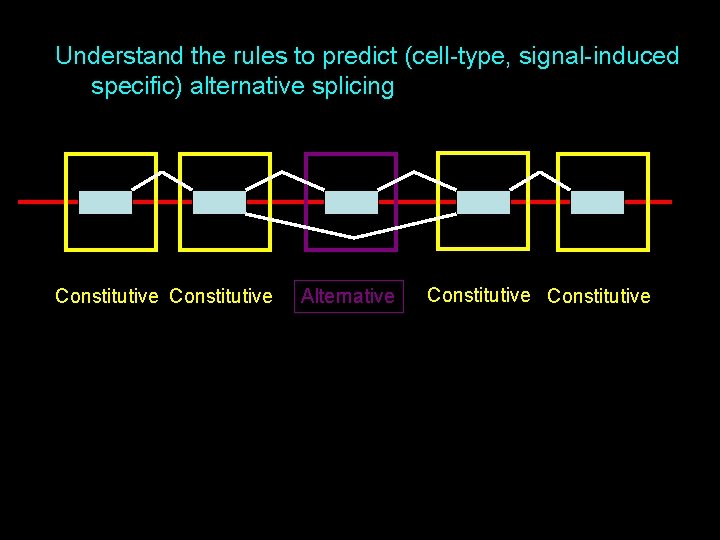
Understand the rules to predict (cell-type, signal-induced specific) alternative splicing Constitutive Alternative Constitutive
![ACEScan[+] exons on UCSC browser Yeo, PNAS 05 ACEScan[+] exons on UCSC browser Yeo, PNAS 05](http://slidetodoc.com/presentation_image_h/307a891a42181006f590a05e39ccc917/image-3.jpg)
ACEScan[+] exons on UCSC browser Yeo, PNAS 05
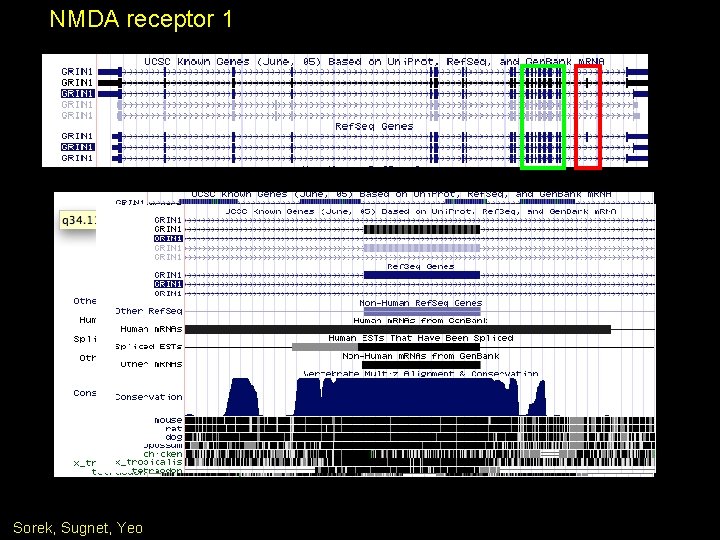
NMDA receptor 1 Sorek, Sugnet, Yeo
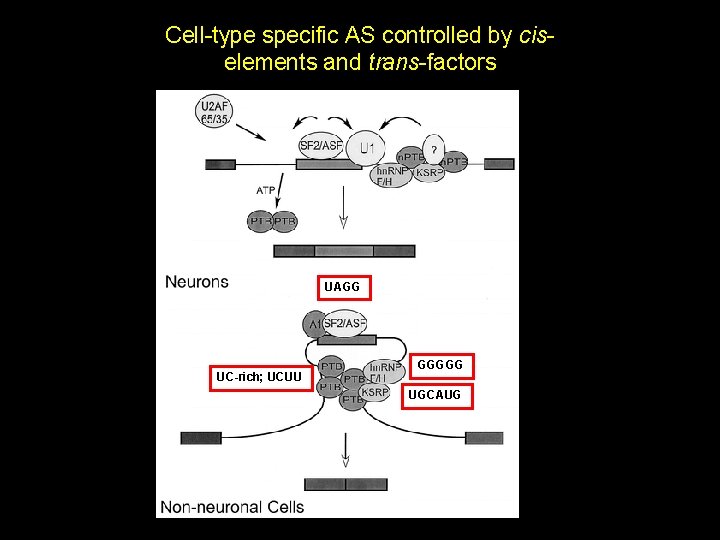
Cell-type specific AS controlled by ciselements and trans-factors UAGG UC-rich; UCUU GGGGG UGCAUG
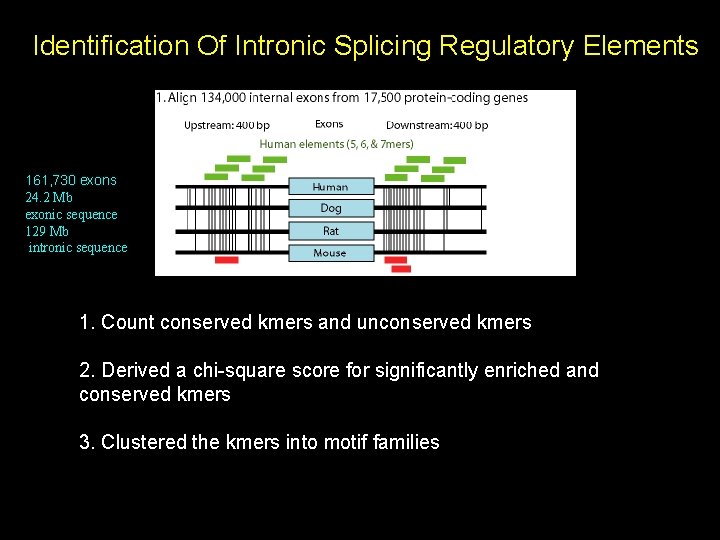
Identification Of Intronic Splicing Regulatory Elements 161, 730 exons 24. 2 Mb exonic sequence 129 Mb intronic sequence 1. Count conserved kmers and unconserved kmers 2. Derived a chi-square score for significantly enriched and conserved kmers 3. Clustered the kmers into motif families

156 upstream motif clusters 158 downstream motif clusters Example of a downstream motif cluster: TGCATG , TGCATGA, ATGCATG, CTGCATG, TGCATGC, TGCATGT, TGCATGG, GTGCATG What are the properties of these motifs? • Positional biases? • Near alternatively spliced exons? • Expression biases? • Overlap known elements?
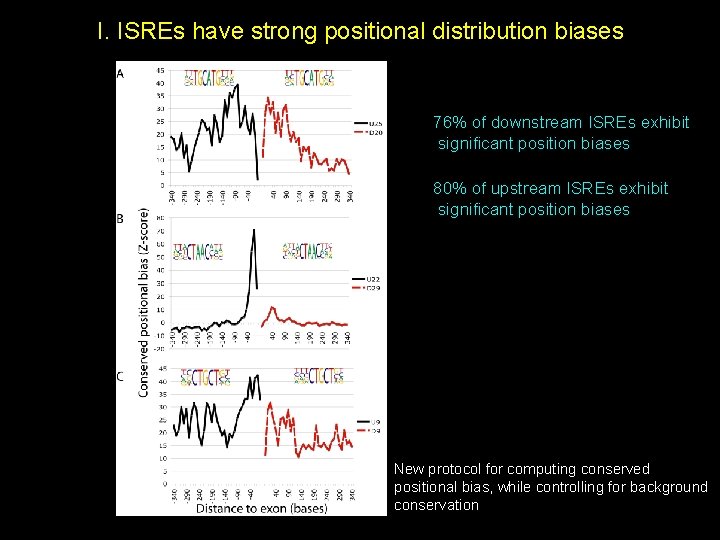
I. ISREs have strong positional distribution biases 76% of downstream ISREs exhibit significant position biases 80% of upstream ISREs exhibit significant position biases New protocol for computing conserved positional bias, while controlling for background conservation
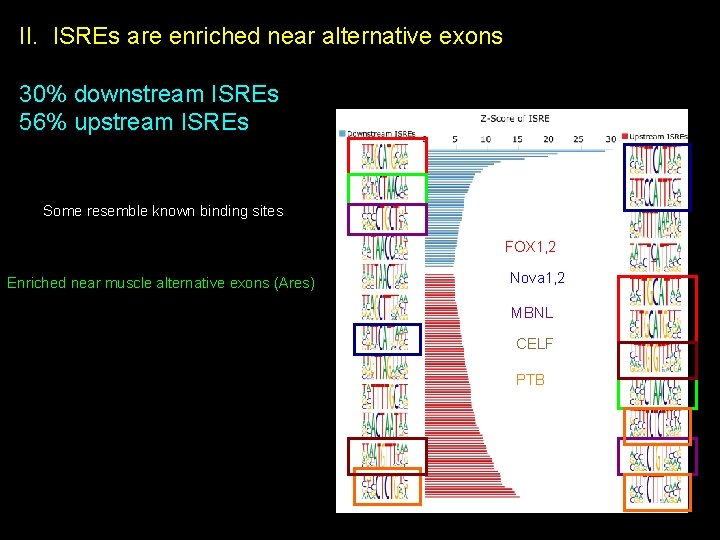
II. ISREs are enriched near alternative exons 30% downstream ISREs 56% upstream ISREs Some resemble known binding sites FOX 1, 2 Enriched near muscle alternative exons (Ares) Nova 1, 2 MBNL CELF PTB
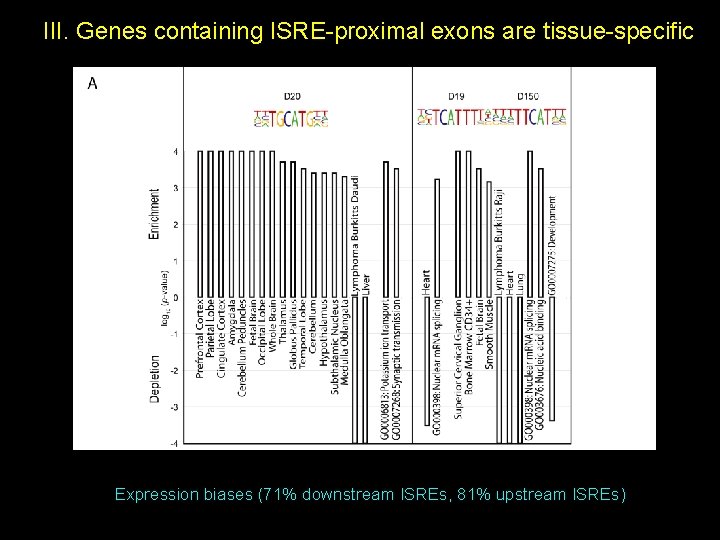
III. Genes containing ISRE-proximal exons are tissue-specific Expression biases (71% downstream ISREs, 81% upstream ISREs)
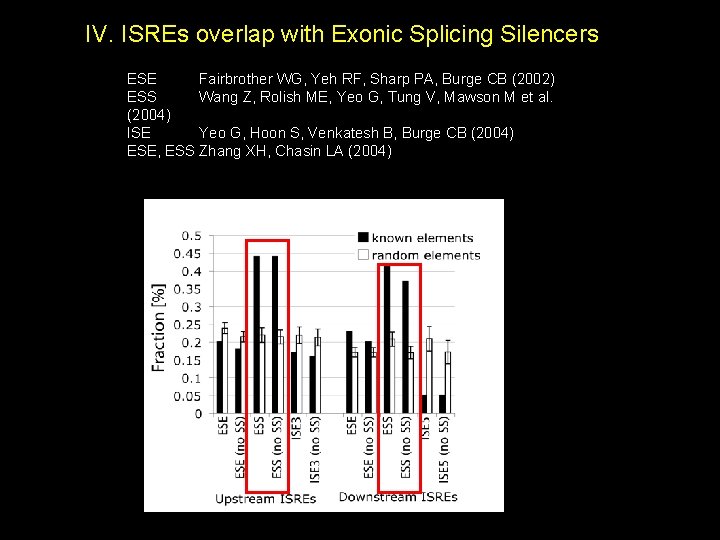
IV. ISREs overlap with Exonic Splicing Silencers ESE Fairbrother WG, Yeh RF, Sharp PA, Burge CB (2002) ESS Wang Z, Rolish ME, Yeo G, Tung V, Mawson M et al. (2004) ISE Yeo G, Hoon S, Venkatesh B, Burge CB (2004) ESE, ESS Zhang XH, Chasin LA (2004)

Like ESS, do ISREs affect Splice Site Choice ? Competing 5’ss and 3’ss reporter (Wang et al, Mol Cell, July, 2006)
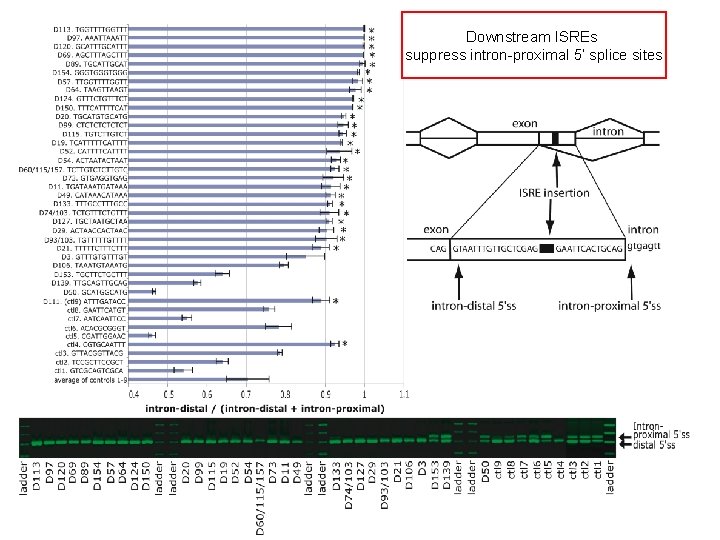
Downstream ISREs suppress intron-proximal 5’ splice sites
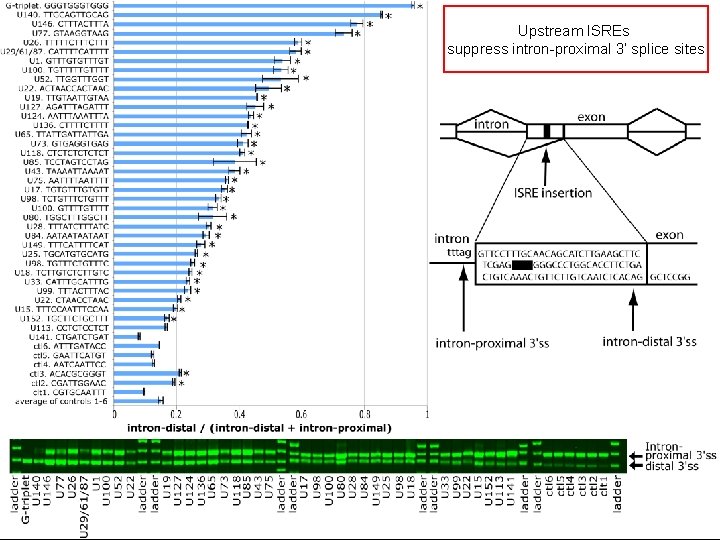
Upstream ISREs suppress intron-proximal 3’ splice sites

Applications of ISREs: (1) splicing arrays --tissue specific alternative splicing? included muscle skipped included skipped brain Enriched ISREs in downstream introns TGCATG ACTAAC TTGGTTT GCATG TCATTTT TTTCAT Enriched Depleted Sugnet et al. PLo. S Comput Biol, 2006

Applications of ISREs: (2) predicting RNA binding sites 1. Several proteins have been reported to affect their own alternative splicing (e. g. hn. RNP A 1, SRP 20, SC 35, TIA 1, TIAR 2, FOX 2, PTB) 2. Evolutionarily conserved AS exons have high intronic conservation flanking the exon, resulting in algorithms that perform genomic predictions of alternative conserved exons 3. Alternative conserved exons are enriched in genes encoding RNA binding proteins and splicing factors
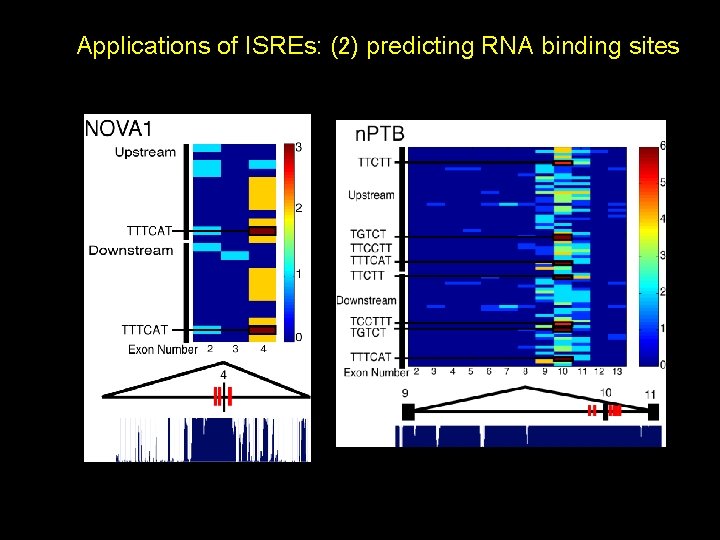
Applications of ISREs: (2) predicting RNA binding sites
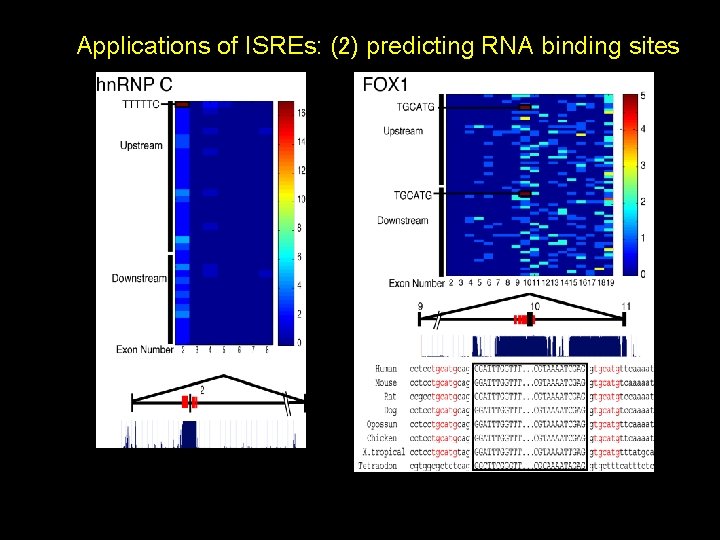
Applications of ISREs: (2) predicting RNA binding sites

ISREs are likely functional • ISREs identified in mammals via comparative genomics. • ISREs have positional biases, are enriched in tissue-specific genes, and overlap with ESS. • ISREs alter splice site choice in vitro. • Some ISREs resemble known sites of known alt splicing factors. • A fraction of ISREs are proximal to alternative exons. • ISREs can be utilized to analyze splicing-array data. • ISREs can be utilized to identify autoregulated exons, and has other implications.
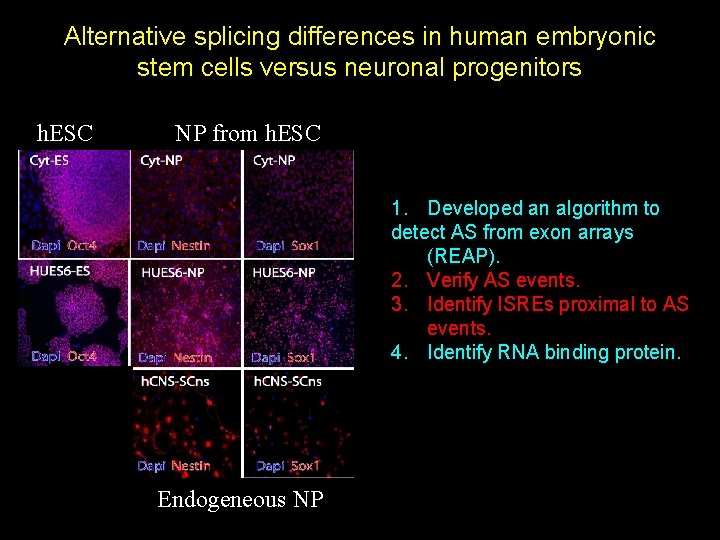
Alternative splicing differences in human embryonic stem cells versus neuronal progenitors h. ESC NP from h. ESC 1. Developed an algorithm to detect AS from exon arrays (REAP). 2. Verify AS events. 3. Identify ISREs proximal to AS events. 4. Identify RNA binding protein. Endogeneous NP

Exon arrays have probesets in every exon Simple representation Exon array, alternative splicing, gene expression Tiling arrays
![REAP predictions agree with A) EST-verified alternative splicing B) ACEScan[+] exons REAP predictions agree with A) EST-verified alternative splicing B) ACEScan[+] exons](http://slidetodoc.com/presentation_image_h/307a891a42181006f590a05e39ccc917/image-22.jpg)
REAP predictions agree with A) EST-verified alternative splicing B) ACEScan[+] exons
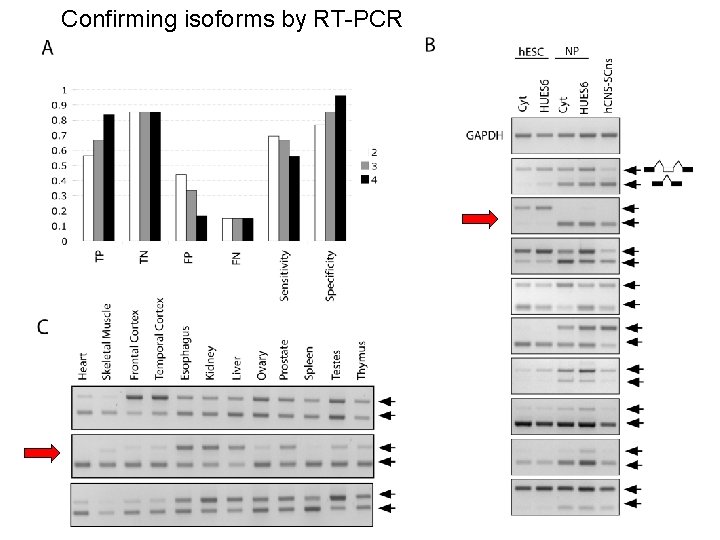
Confirming isoforms by RT-PCR
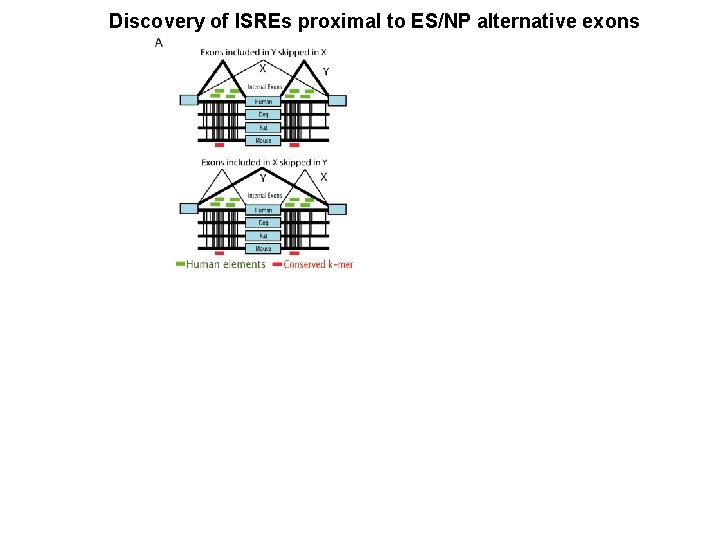
Discovery of ISREs proximal to ES/NP alternative exons

ISREs in ES/NP ongoing… • REAP algorithm designed for exon array based detection of AS events • REAP[+] exons correlate with EST-based and ACEScan[+] exons • ISREs identified specific for ES/NP AS events • FOX 1/2 may regulate ES/NP-specific AS events

Alternative splicing at the Crick-Jacobs Center, Salk Institute Computational Modeling, Integration Constitutive Alternative Constitutive Stem cells, early neuronal differentiation Cis-elements Association of RNA binding proteins to elements
- Slides: 26